Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
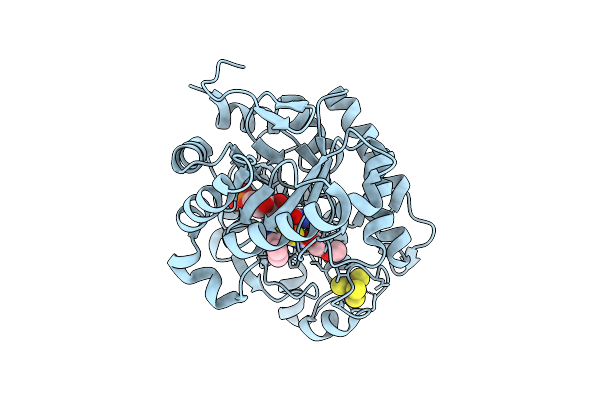 |
Crystal Structure Of Dehydrogenase/Isomerase Fabx From Helicobacter Pylori In Complex With Inhibitor 1
Organism: Helicobacter pylori
Method: X-RAY DIFFRACTION Release Date: 2025-12-24 Classification: BIOSYNTHETIC PROTEIN Ligands: SF4, A1ELE, FMN |
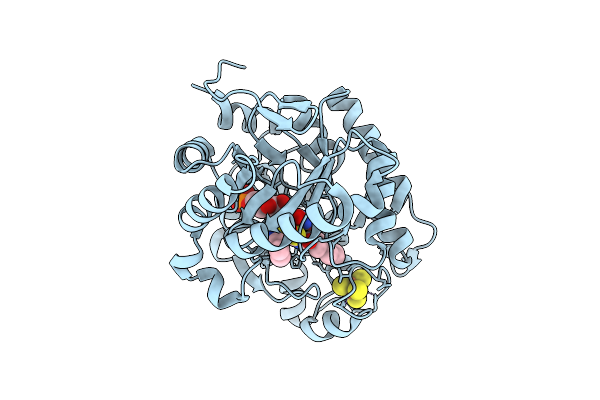 |
Crystal Structure Of Dehydrogenase/Isomerase Fabx From Helicobacter Pylori In Complex With Inhibitor 21
Organism: Helicobacter pylori
Method: X-RAY DIFFRACTION Release Date: 2025-12-24 Classification: BIOSYNTHETIC PROTEIN Ligands: FMN, SF4, A1ELF |
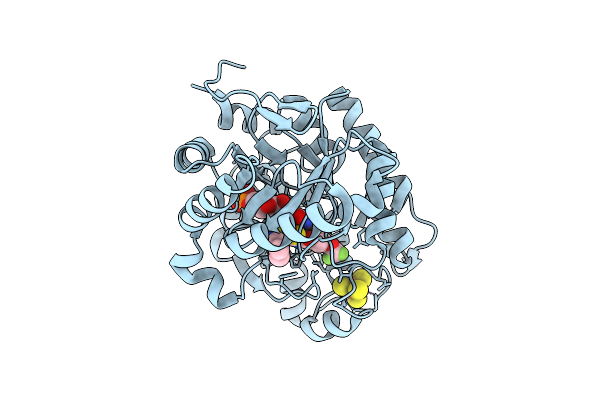 |
Crystal Structure Of Dehydrogenase/Isomerase Fabx From Helicobacter Pylori In Complex With Inhibitor 11
Organism: Helicobacter pylori
Method: X-RAY DIFFRACTION Release Date: 2025-12-24 Classification: BIOSYNTHETIC PROTEIN Ligands: FMN, SF4, A1ELG |
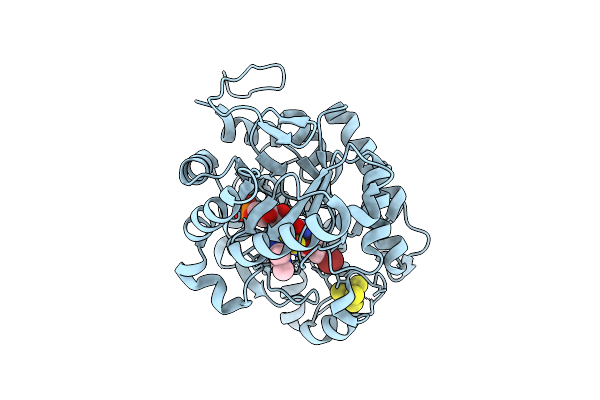 |
Crystal Structure Of Dehydrogenase/Isomerase Fabx From Helicobacter Pylori In Complex With Inhibitor 35
Organism: Helicobacter pylori
Method: X-RAY DIFFRACTION Release Date: 2025-12-24 Classification: BIOSYNTHETIC PROTEIN Ligands: FMN, SF4, A1ELH |
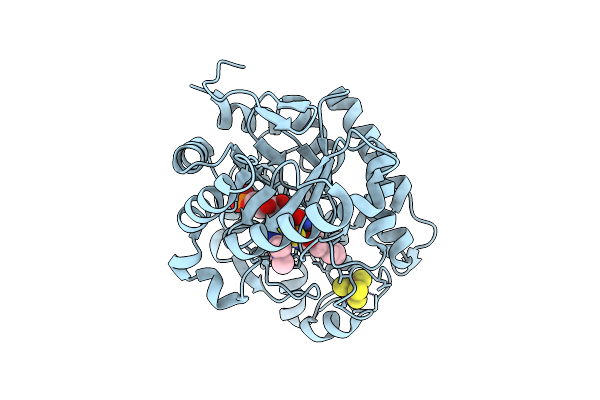 |
Crystal Structure Of Dehydrogenase/Isomerase Fabx From Helicobacter Pylori In Complex With Inhibitor 47
Organism: Helicobacter pylori
Method: X-RAY DIFFRACTION Release Date: 2025-12-24 Classification: BIOSYNTHETIC PROTEIN Ligands: FMN, SF4, A1EMD |
 |
Organism: Helicobacter pylori g27
Method: ELECTRON MICROSCOPY Release Date: 2025-12-03 Classification: HYDROLASE |
 |
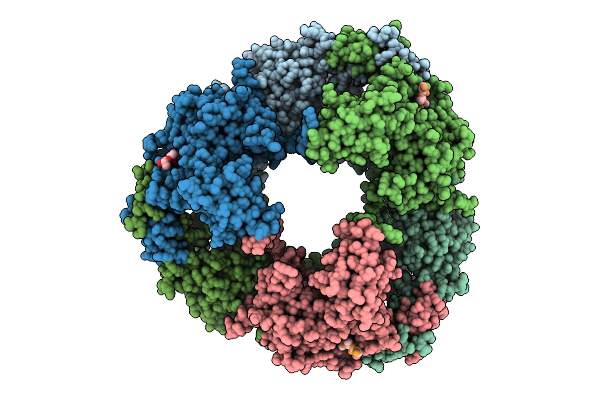
Organism: Helicobacter pylori g27
Method: X-RAY DIFFRACTION Release Date: 2025-12-03 Classification: HYDROLASE |
 |
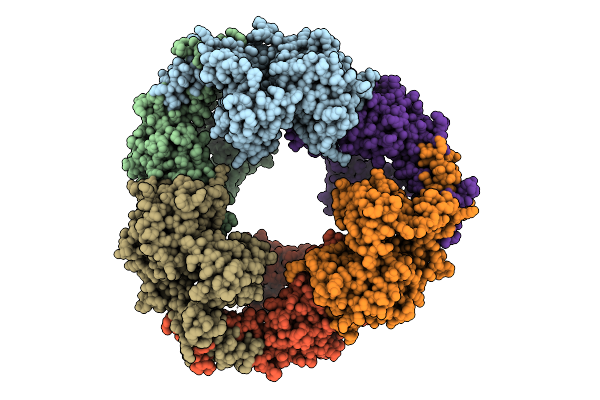
Cryo-Em Structure Of Ctpa S300A/K325A/Q329A Mutant From Helicobacter Pylori
Organism: Helicobacter pylori g27, Escherichia coli bl21(de3)
Method: ELECTRON MICROSCOPY Release Date: 2025-12-03 Classification: HYDROLASE |
 |
Organism: Helicobacter pylori g27
Method: ELECTRON MICROSCOPY Release Date: 2025-12-03 Classification: HYDROLASE |
 |
Organism: Helicobacter pylori g27
Method: ELECTRON MICROSCOPY Release Date: 2025-12-03 Classification: HYDROLASE |
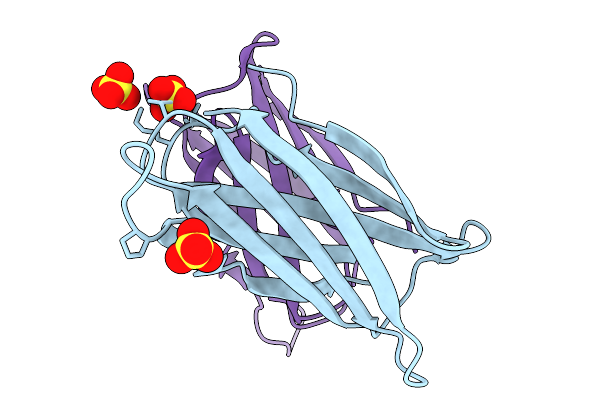 |
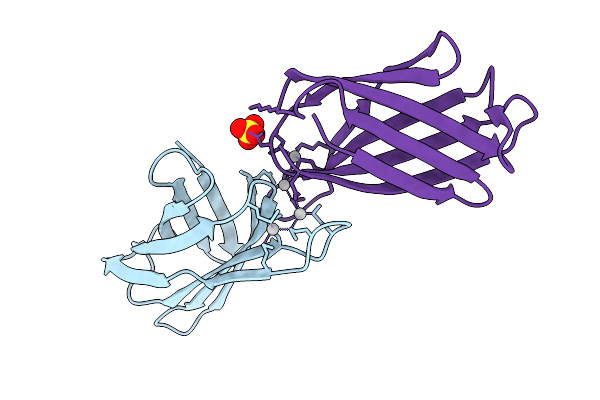
Crystal Structure Of The Helicobacter Pylori Copper Resistance Determinant Crda In The Apo Form
Organism: Helicobacter pylori
Method: X-RAY DIFFRACTION Release Date: 2025-11-19 Classification: METAL BINDING PROTEIN Ligands: SO4 |
 |
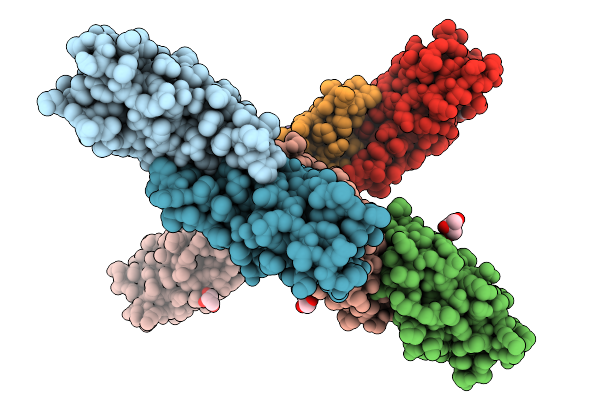
Crystal Structure Of The Helicobacter Pylori Copper Resistance Determinant Crda In Complex With Silver Ions In Space Group P21
Organism: Helicobacter pylori
Method: X-RAY DIFFRACTION Release Date: 2025-11-19 Classification: METAL BINDING PROTEIN Ligands: AG, SO4 |
 |
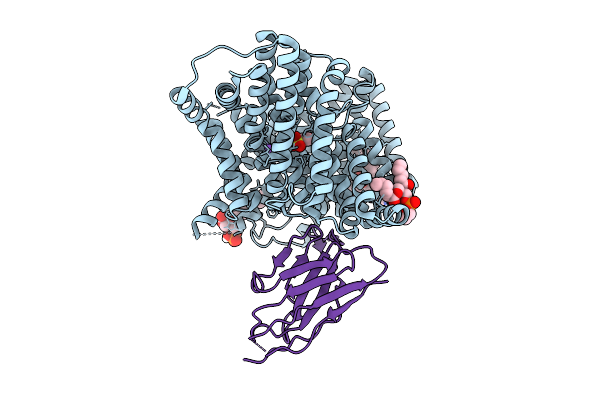
Crystal Structure Of The Helicobacter Pylori Copper Resistance Determinant Crda In Complex With Silver Ions In Space Group P1
Organism: Helicobacter pylori
Method: X-RAY DIFFRACTION Release Date: 2025-11-19 Classification: METAL BINDING PROTEIN Ligands: AG, GOL |
 |
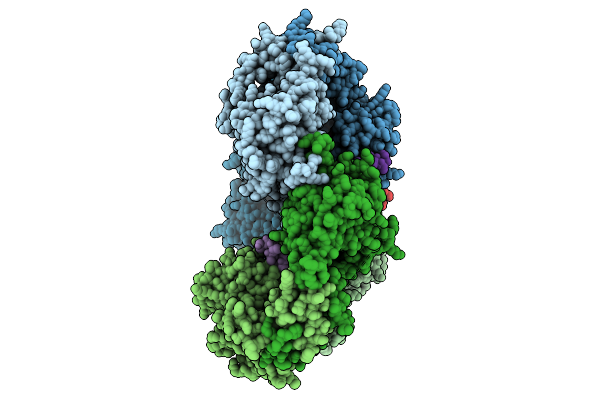
Cryo-Em Structure Of The Isethionate Trap Transporter Iseqm From Oleidesulfovibrio Alaskensis With Bound Isethionate
Organism: Oleidesulfovibrio alaskensis g20, Helicobacter pylori
Method: ELECTRON MICROSCOPY Release Date: 2025-11-12 Classification: TRANSPORT PROTEIN |
 |
Crystal Structure Of Helicobacter Pylori Chemoreceptor Tlpa Ligand-Binding Domain In Complex With Indole
Organism: Helicobacter pylori pmss1
Method: X-RAY DIFFRACTION Release Date: 2025-10-22 Classification: SIGNALING PROTEIN Ligands: IND, EDO |
 |
Organism: Helicobacter pylori g27, Escherichia coli bl21(de3)
Method: X-RAY DIFFRACTION Release Date: 2025-10-01 Classification: HYDROLASE |
 |
Crystal Structure Of Dehydrogenase/Isomerase Fabx From Helicobacter Pylori In Complex With Inhibitor 1991
Organism: Helicobacter pylori
Method: X-RAY DIFFRACTION Release Date: 2025-08-20 Classification: BIOSYNTHETIC PROTEIN Ligands: A1ECY, FMN, SF4, MG |
 |
Crystal Structure Of Dehydrogenase/Isomerase Fabx From Helicobacter Pylori In Complex With Inhibitor 1872
Organism: Helicobacter pylori
Method: X-RAY DIFFRACTION Release Date: 2025-08-20 Classification: BIOSYNTHETIC PROTEIN Ligands: FMN, SF4, A1EEQ |
 |
Organism: Helicobacter pylori ss1
Method: X-RAY DIFFRACTION Release Date: 2025-07-23 Classification: OXIDOREDUCTASE Ligands: HEM, GOL, NA, CL |
 |
Organism: Helicobacter pylori 26695
Method: X-RAY DIFFRACTION Release Date: 2025-07-16 Classification: BIOSYNTHETIC PROTEIN Ligands: IPA, CL |