Search Count: 301
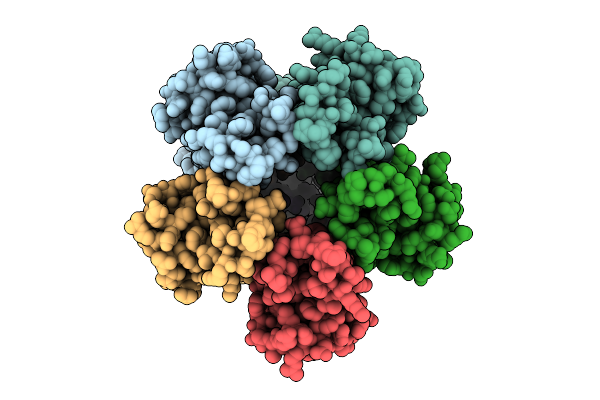 |
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: MEMBRANE PROTEIN |
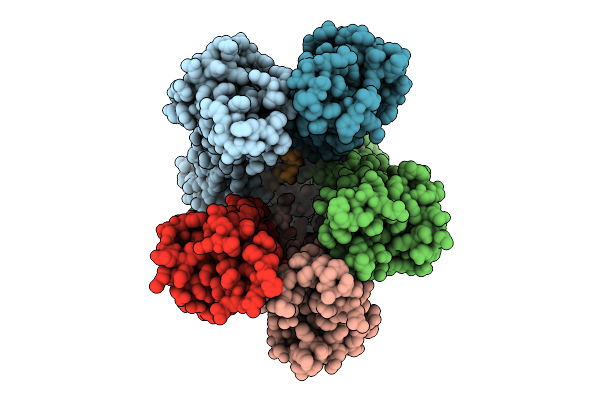 |
Organism: Escherichia coli str. k-12 substr. mg1655
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: MEMBRANE PROTEIN |
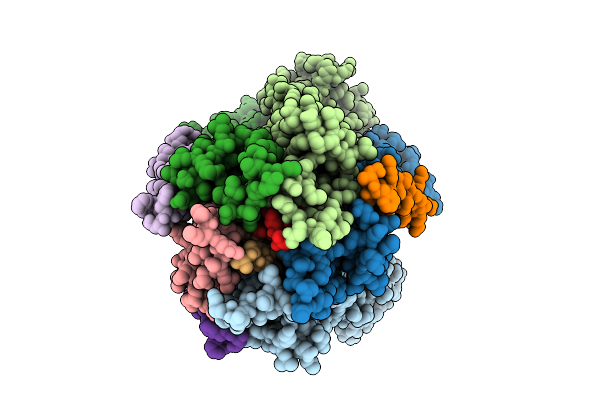 |
Organism: Escherichia coli str. k-12 substr. mg1655
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: MEMBRANE PROTEIN |
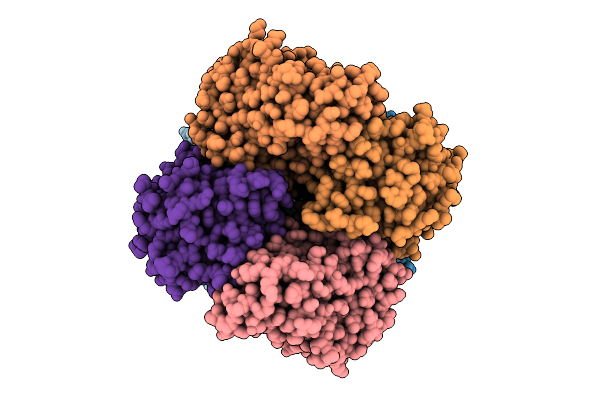 |
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2025-12-10 Classification: IMMUNE SYSTEM |
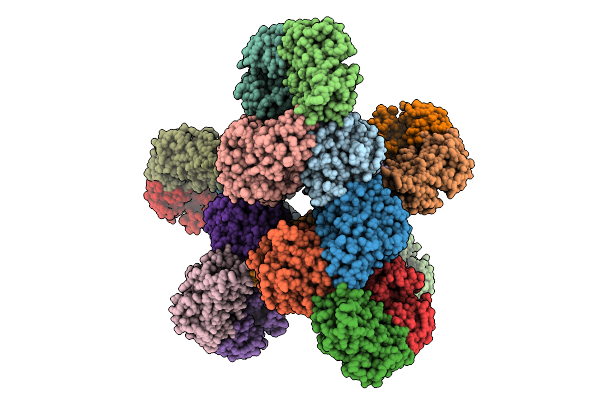 |
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2025-12-10 Classification: IMMUNE SYSTEM |
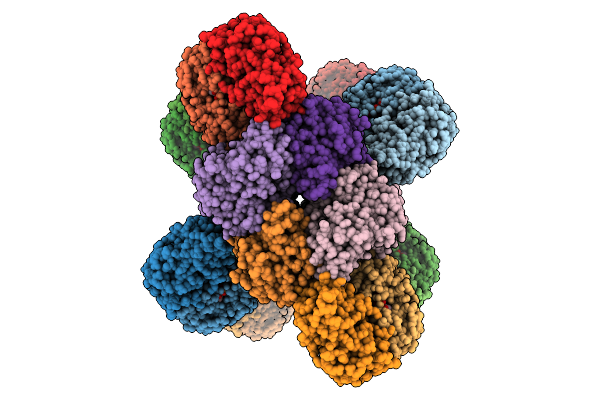 |
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2025-12-10 Classification: IMMUNE SYSTEM Ligands: Y43, MG |
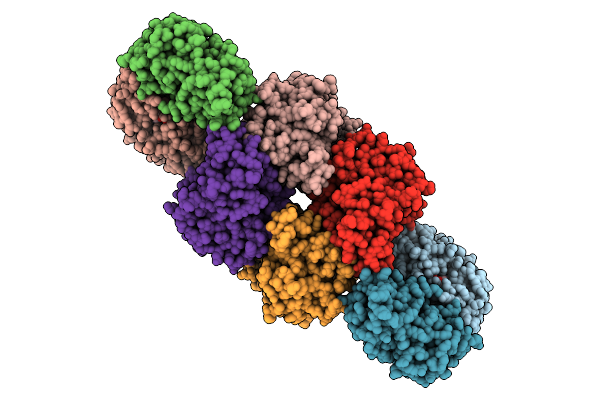 |
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2025-12-10 Classification: IMMUNE SYSTEM Ligands: Y43, NAD |
 |
Omicron-Specific Ultra-Potent Sars-Cov-2 Neutralizing Antibodies Targeting The N1/N2 Loop Of Spike N-Terminal Domain
Organism: Homo sapiens, Severe acute respiratory syndrome coronavirus 2
Method: ELECTRON MICROSCOPY Release Date: 2025-12-03 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
 |
Structure Of Cargo Complex (Btpea-Btaeb-Btapc) Bound To The Vgrg Spike From The Type Vi Secretion System
Organism: Bacteroides fragilis
Method: ELECTRON MICROSCOPY Release Date: 2025-09-10 Classification: ANTIMICROBIAL PROTEIN Ligands: ZN |
 |
Organism: Bacteroides fragilis
Method: X-RAY DIFFRACTION Release Date: 2025-09-03 Classification: ANTIMICROBIAL PROTEIN Ligands: PEG |
 |
Crystal Structure Of The Type Vi Secretion System Effector-Immunity Complex Btpea Ctd-Btpia From Bacteroides Fragilis.
Organism: Bacteroides fragilis
Method: X-RAY DIFFRACTION Release Date: 2025-09-03 Classification: ANTIMICROBIAL PROTEIN Ligands: GOL |
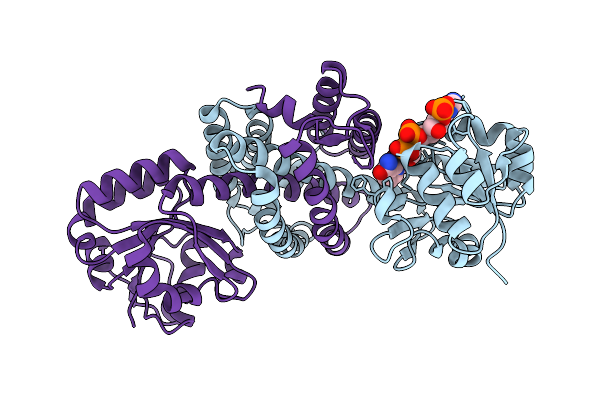 |
Structure Of Imine Reductase 361 From Micromonospora Sp. Mutant M125W/I127F/L179V/H250L
Organism: Micromonospora
Method: X-RAY DIFFRACTION Release Date: 2025-08-27 Classification: OXIDOREDUCTASE Ligands: NAP |
 |
Organism: Alternaria alternata
Method: ELECTRON MICROSCOPY Release Date: 2025-06-18 Classification: TOXIN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Release Date: 2025-05-07 Classification: IMMUNE SYSTEM Ligands: CDP, A1L1P, ZN |
 |
Organism: Homo sapiens, Streptococcus pneumoniae, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2025-04-02 Classification: SIGNALING PROTEIN/IMMUNE SYSTEM Ligands: SA0 |
 |
Organism: Homo sapiens, Streptococcus pneumoniae
Method: ELECTRON MICROSCOPY Release Date: 2025-04-02 Classification: SIGNALING PROTEIN Ligands: SA0 |
 |
Organism: Homo sapiens, Streptococcus pneumoniae
Method: ELECTRON MICROSCOPY Release Date: 2025-04-02 Classification: SIGNALING PROTEIN Ligands: SA0 |
 |
Organism: Homo sapiens, Streptococcus pneumoniae
Method: ELECTRON MICROSCOPY Release Date: 2025-04-02 Classification: SIGNALING PROTEIN Ligands: SA0 |
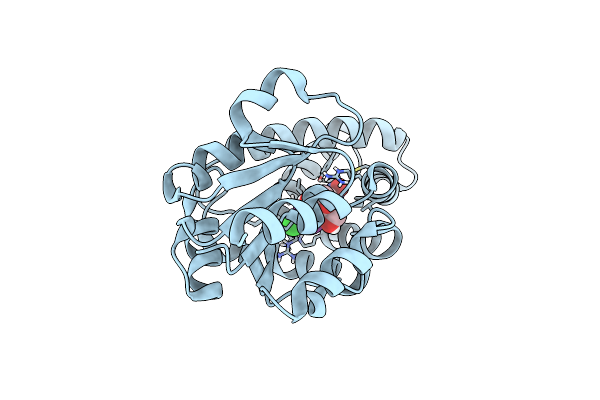 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.71 Å Release Date: 2025-01-29 Classification: BIOSYNTHETIC PROTEIN Ligands: PEG, A1IGD, CL |
 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.88 Å Release Date: 2025-01-29 Classification: BIOSYNTHETIC PROTEIN Ligands: IOD, PG4 |

