Search Count: 23
 |
Grouped 150-240 Ms Dark Structure Of Sensory Rhodopsin-Ii Solved By Serial Millisecond Crystallography
Organism: Natronomonas pharaonis
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2025-04-30 Classification: SIGNALING PROTEIN Ligands: RET, CL, BOG, MPG |
 |
Continuously Illuminated Structure Of Sensory Rhodopsin Ii Solved By Serial Millisecond Crystallography
Organism: Natronomonas pharaonis
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2025-04-30 Classification: SIGNALING PROTEIN Ligands: RET, CL |
 |
Continuous Dark State Structure Of Sensory Rhodopsin Ii Solved By Serial Millisecond Crystallography
Organism: Natronomonas pharaonis
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2025-04-30 Classification: SIGNALING PROTEIN Ligands: RET, CL, BOG, MPG |
 |
Light Structure Of Sensory Rhodopsin-Ii Solved By Serial Millisecond Crystallography 90-120 Milliseconds Time-Bin
Organism: Natronomonas pharaonis
Method: X-RAY DIFFRACTION Resolution:1.94 Å Release Date: 2025-02-05 Classification: SIGNALING PROTEIN Ligands: RET, CL |
 |
Light Structure Of Sensory Rhodopsin-Ii Solved By Serial Millisecond Crystallography 60-90 Milliseconds Time-Bin
Organism: Natronomonas pharaonis
Method: X-RAY DIFFRACTION Resolution:1.96 Å Release Date: 2025-02-05 Classification: SIGNALING PROTEIN Ligands: RET, CL |
 |
Light Structure Of Sensory Rhodopsin-Ii Solved By Serial Millisecond Crystallography. 30-60 Milliseconds Time-Bin
Organism: Natronomonas pharaonis
Method: X-RAY DIFFRACTION Resolution:2.14 Å Release Date: 2025-02-05 Classification: SIGNALING PROTEIN Ligands: RET, CL |
 |
Light Structure Of Sensory Rhodopsin-Ii Solved By Serial Millisecond Crystallography 0-30 Milliseconds Time-Bin
Organism: Natronomonas pharaonis
Method: X-RAY DIFFRACTION Resolution:2.08 Å Release Date: 2025-02-05 Classification: SIGNALING PROTEIN Ligands: RET, CL |
 |
Light Structure Of Sensory Rhodopsin-Ii Solved By Serial Millisecond Crystallography 120-150 Milliseconds Time-Bin
Organism: Natronomonas pharaonis
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2025-02-05 Classification: SIGNALING PROTEIN Ligands: RET, CL |
 |
Serial Femtosecond Crystallography Structure Of Co Bound Ba3- Type Cytochrome C Oxidase Without Pump Laser Irradiation
Organism: Thermus thermophilus hb8
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2023-11-15 Classification: ELECTRON TRANSPORT Ligands: CU, HEM, HAS, OLC, CMO, CUA |
 |
Serial Femtosecond Crystallography Structure Of Photo Dissociated Co From Ba3- Type Cytochrome C Oxidase Determined By Extrapolation Method
Organism: Thermus thermophilus hb8
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2023-11-15 Classification: ELECTRON TRANSPORT Ligands: CU, HEM, HAS, OLC, CMO, CUA |
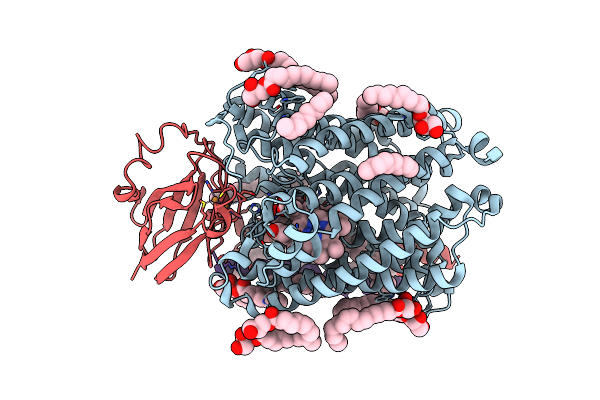 |
Serial Femtosecond Crystallography Structure Of Co Bound Ba3- Type Cytochrome C Oxidase At 2 Milliseconds After Irradiation By A 532 Nm Laser
Organism: Thermus thermophilus
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2023-08-16 Classification: ELECTRON TRANSPORT Ligands: CU, HEM, HAS, OLC, CMO, CUA |
 |
Lipidic Cubic Phase Serial Femtosecond Crystallography Structure Of A Photosynthetic Reaction Centre
Organism: Blastochloris viridis
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2022-06-22 Classification: MEMBRANE PROTEIN Ligands: HEC, DGA, HTO, OLC, SO4, BCB, BPB, UQ1, LDA, FE2, MQ7, NS5 |
 |
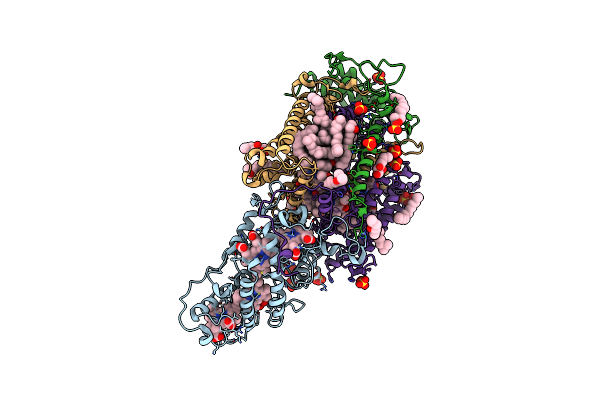
Lipidic Cubic Phase Serial Femtosecond Crystallography Structure Of A Photosynthetic Reaction Centre
Organism: Blastochloris viridis
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2022-06-22 Classification: MEMBRANE PROTEIN Ligands: HEC, DGA, LDA, SO4, BCB, BPB, HTO, UQ2, FE2, MQ9, NS5, OLC |
 |
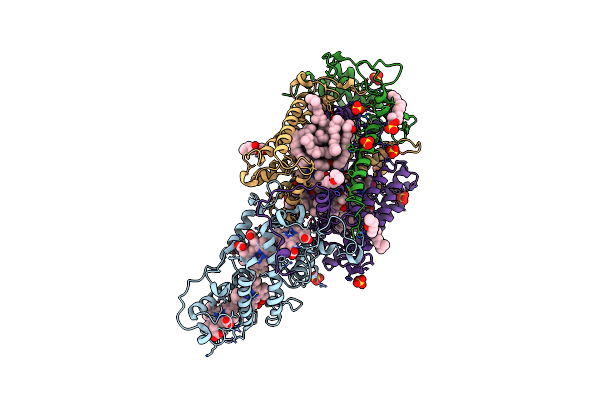
Ultrafast Structural Response To Charge Redistribution Within A Photosynthetic Reaction Centre - 1 Ps Structure
Organism: Blastochloris viridis
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2020-12-09 Classification: ELECTRON TRANSPORT Ligands: HEC, DGA, SO4, LDA, HTO, BCB, BPB, FE, MQ7, NS5 |
 |
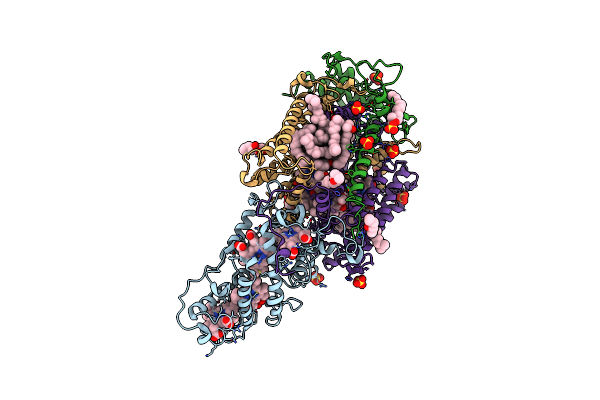
Ultrafast Structural Response To Charge Redistribution Within A Photosynthetic Reaction Centre - 5 Ps (A) Structure
Organism: Blastochloris viridis
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2020-12-09 Classification: ELECTRON TRANSPORT Ligands: HEC, DGA, SO4, LDA, HTO, BCB, BPB, FE, MQ7, NS5 |
 |
Ultrafast Structural Response To Charge Redistribution Within A Photosynthetic Reaction Centre - 300 Ps (A) Structure
Organism: Blastochloris viridis
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2020-12-09 Classification: ELECTRON TRANSPORT Ligands: HEC, DGA, SO4, LDA, HTO, BCB, BPB, FE, MQ7, NS5 |
 |
Ultrafast Structural Response To Charge Redistribution Within A Photosynthetic Reaction Centre - 20 Ps Structure
Organism: Blastochloris viridis
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2020-12-09 Classification: ELECTRON TRANSPORT Ligands: HEC, DGA, SO4, LDA, HTO, BCB, BPB, FE, MQ7, NS5 |
 |
Ultrafast Structural Response To Charge Redistribution Within A Photosynthetic Reaction Centre - 300 Ps (B) Structure
Organism: Blastochloris viridis
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2020-12-09 Classification: ELECTRON TRANSPORT Ligands: HEC, DGA, SO4, LDA, HTO, BCB, BPB, FE, MQ7, NS5 |
 |
Ultrafast Structural Response To Charge Redistribution Within A Photosynthetic Reaction Centre - 8 Us Structure
Organism: Blastochloris viridis
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2020-12-09 Classification: ELECTRON TRANSPORT Ligands: HEC, DGA, SO4, LDA, HTO, BCB, BPB, FE, MQ7, NS5 |
 |
Ultrafast Structural Response To Charge Redistribution Within A Photosynthetic Reaction Centre - 5 Ps (B) Structure
Organism: Blastochloris viridis
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2020-12-09 Classification: ELECTRON TRANSPORT Ligands: HEC, DGA, SO4, LDA, HTO, BCB, BPB, FE, MQ7, NS5 |

