Search Count: 6
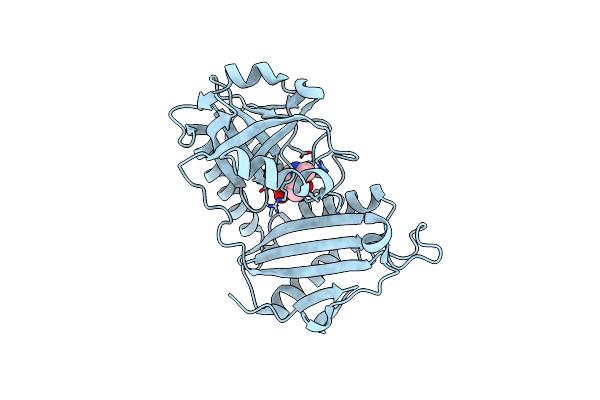 |
Crystal Structure Of D-Amino Acid Transaminase From Haliscomenobacter Hydrossis In The Holo Form Obtained At Ph 7.0
Organism: Haliscomenobacter hydrossis dsm 1100
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2024-04-17 Classification: TRANSFERASE Ligands: PLP |
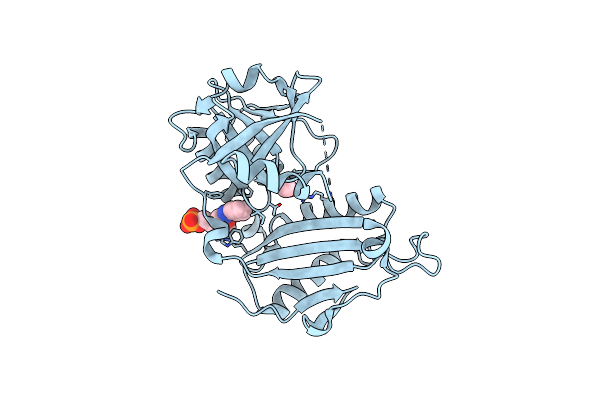 |
Crystal Structure Of D-Amino Acid Transaminase From Haliscomenobacter Hydrossis (Apo Form) After 15 Sec Of Soaking With Phenylhydrazine
Organism: Haliscomenobacter hydrossis dsm 1100
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2024-04-17 Classification: TRANSFERASE Ligands: ACT, ZXN |
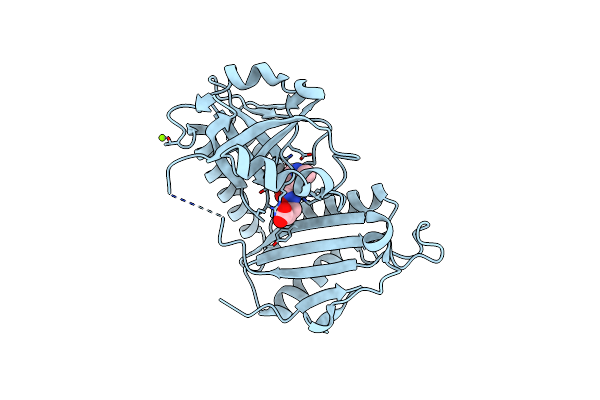 |
Crystal Structure Of D-Amino Acid Transaminase From Haliscomenobacter Hydrossis Complexed With 3-Aminooxypropionic Acid
Organism: Haliscomenobacter hydrossis dsm 1100
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2024-04-17 Classification: TRANSFERASE Ligands: OCF, MG |
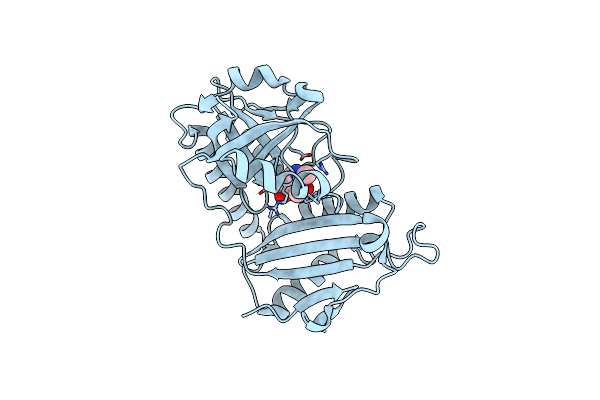 |
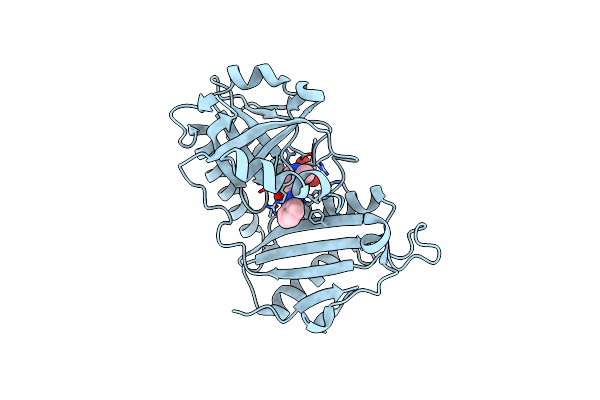
Crystal Structure Of D-Amino Acid Transaminase From Haliscomenobacter Hydrossis Point Mutant R90I (Holo Form)
Organism: Haliscomenobacter hydrossis dsm 1100
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2023-12-27 Classification: TRANSFERASE Ligands: PLP |
 |
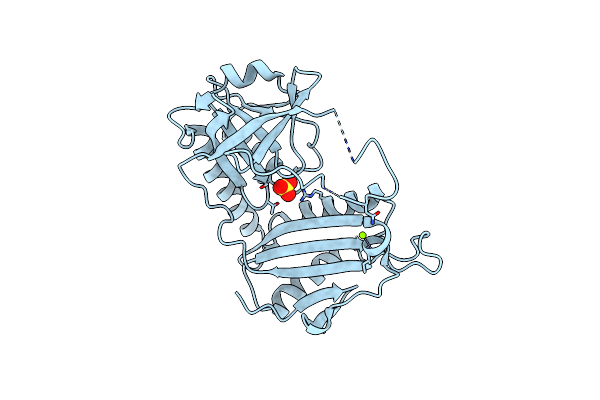
Crystal Structure Of D-Amino Acid Transaminase From Haliscomenobacter Hydrossis Point Mutant R90I Complexed With Phenylhydrazine
Organism: Haliscomenobacter hydrossis dsm 1100
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2023-12-27 Classification: TRANSFERASE Ligands: ZXN, GOL |
 |
Crystal Structure Of D-Aminoacid Transaminase From Haliscomenobacter Hydrossis In Its Intermediate Form
Organism: Haliscomenobacter hydrossis dsm 1100
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2022-08-03 Classification: TRANSFERASE Ligands: SO4, MG |

