Search Count: 15
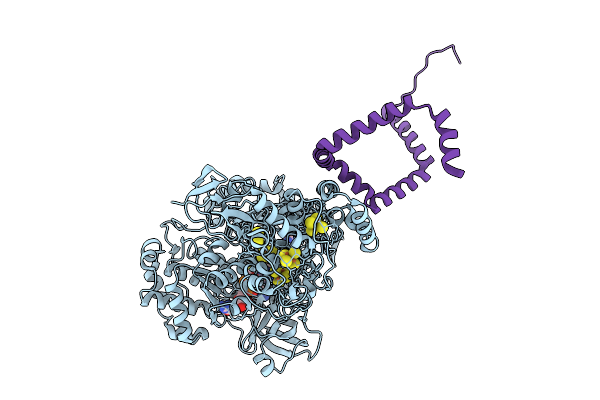 |
Organism: Bacillus subtilis
Method: X-RAY DIFFRACTION Release Date: 2025-09-10 Classification: ELECTRON TRANSPORT Ligands: FES, SF4, MGD, 4MO |
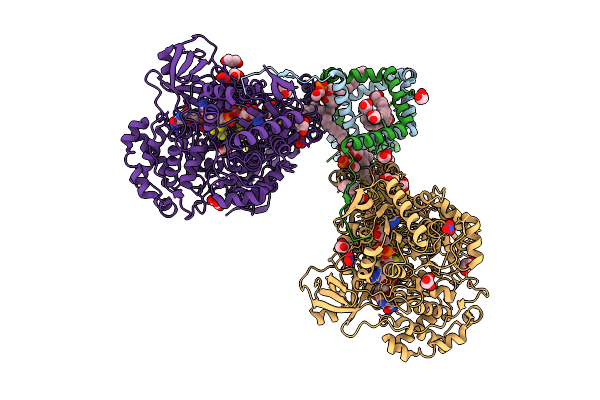 |
Crystal Structure Of Molybdenum Bispyranopterin Guanine Dinucleotide Formate Dehydrogenases Force1 From Bacillus Subtilis
Organism: Bacillus subtilis subsp. subtilis str. 168
Method: X-RAY DIFFRACTION Release Date: 2025-08-06 Classification: OXIDOREDUCTASE Ligands: MYR, GOL, LHG, FES, SF4, MGD, 4MO, MQ7, H2S, MLI, NO3, NA |
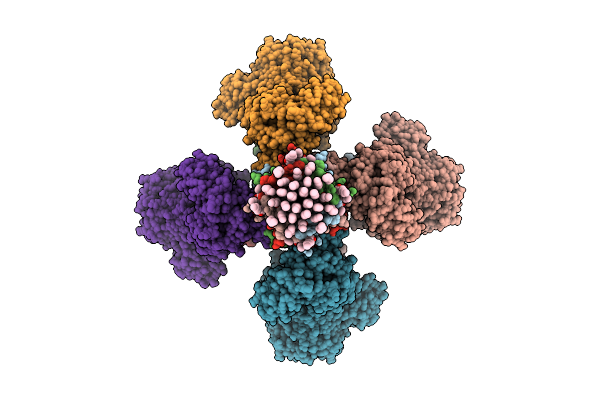 |
Cryoem Structure Of Molybdenum Bispyranopterin Guanine Dinucleotide Formate Dehydrogenases Force1 From Bacillus Subtilis
Organism: Bacillus subtilis subsp. subtilis str. 168
Method: ELECTRON MICROSCOPY Release Date: 2025-08-06 Classification: OXIDOREDUCTASE Ligands: 11A, A1H2V, SHV, KNA, MYR, FES, SF4, MGD, 4MO, H2S, MQ7 |
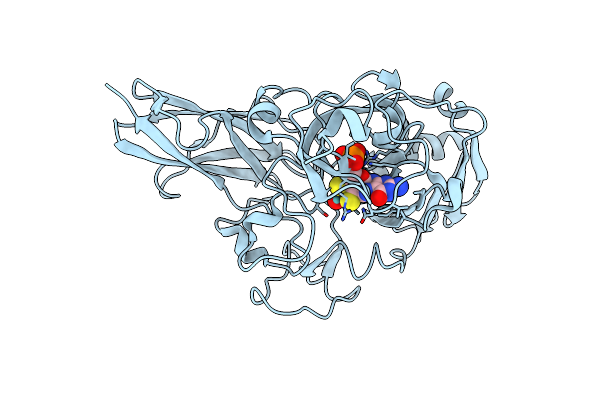 |
Organism: Thermus thermophilus hb8
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2024-11-27 Classification: OXIDOREDUCTASE Ligands: MSS, CL |
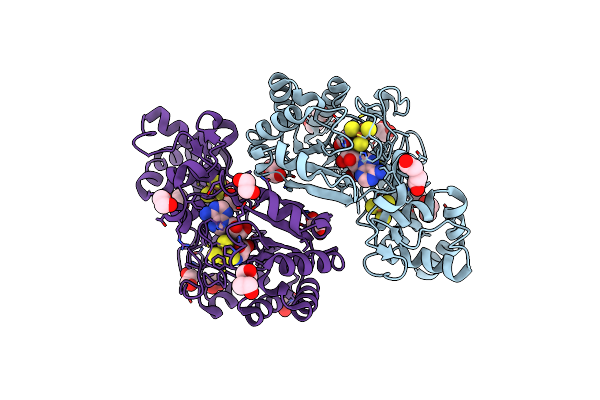 |
Crystal Structure Of Radical Sam Epimerase Epee From Bacillus Subtilis With [4Fe-4S] Clusters And S-Adenosyl-L-Homocysteine Bound.
Organism: Bacillus subtilis
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2024-01-10 Classification: METAL BINDING PROTEIN Ligands: SF4, SAH, EDO, PEG |
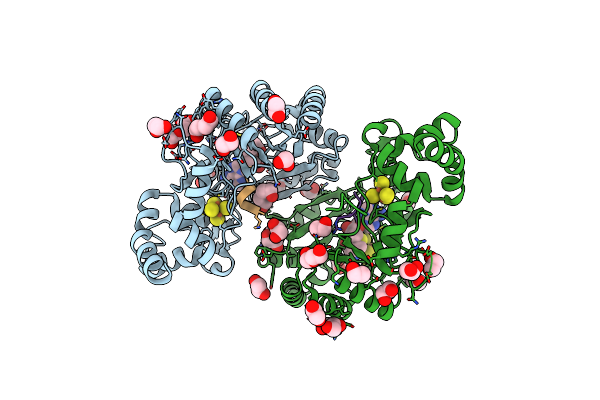 |
Crystal Structure Of Radical Sam Epimerase Epee From Bacillus Subtilis With [4Fe-4S] Clusters, S-Adenosyl-L-Homocysteine And Ripp Peptide 5 Bound
Organism: Bacillus subtilis
Method: X-RAY DIFFRACTION Resolution:2.39 Å Release Date: 2024-01-10 Classification: METAL BINDING PROTEIN Ligands: SF4, SAH, EDO, PEG |
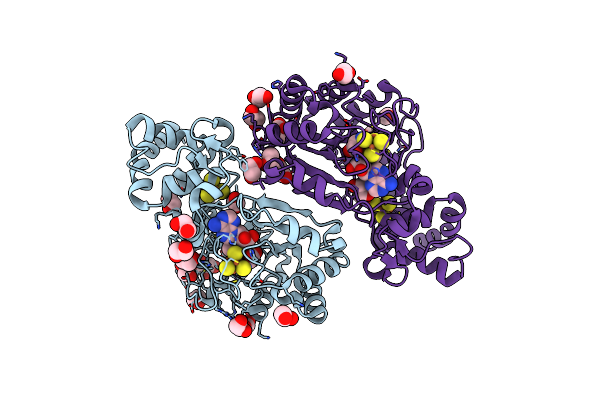 |
Crystal Structure Of Radical Sam Epimerase Epee C223A Mutant From Bacillus Subtilis With [4Fe-4S] Clusters And S-Adenosyl-L-Methionine Bound
Organism: Bacillus subtilis
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2024-01-10 Classification: METAL BINDING PROTEIN Ligands: SF4, SAM, EDO, GOL |
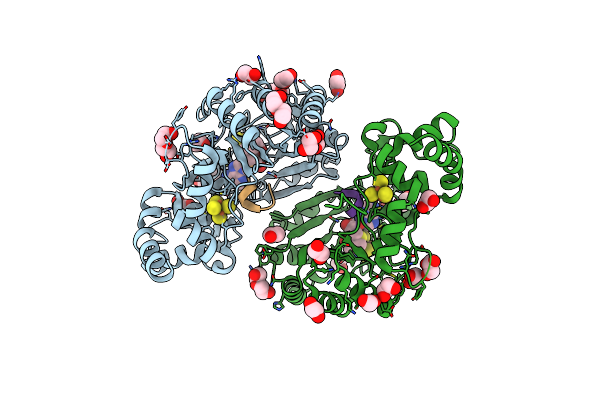 |
Crystal Structure Of Radical Sam Epimerase Epee C223A Mutant From Bacillus Subtilis With [4Fe-4S] Clusters, S-Adenosyl-L-Homocysteine And Ripp Peptide 5 Bound
Organism: Bacillus subtilis
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2024-01-10 Classification: METAL BINDING PROTEIN Ligands: SF4, SAH, EDO, PEG, GOL |
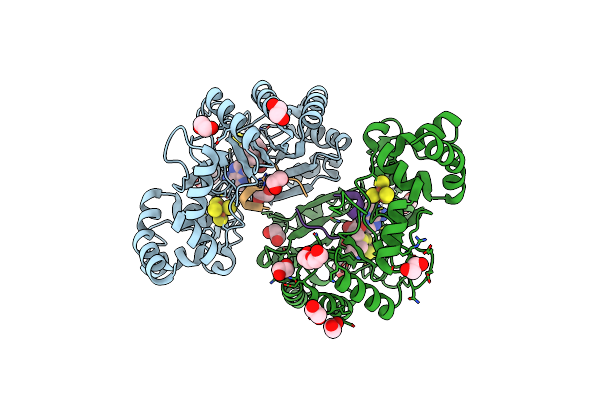 |
Crystal Structure Of Radical Sam Epimerase Epee C223A Mutant From Bacillus Subtilis With [4Fe-4S] Clusters, S-Adenosyl-L-Homocysteine And Ripp Peptide 6 Bound
Organism: Bacillus subtilis
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2024-01-10 Classification: METAL BINDING PROTEIN Ligands: SF4, SAH, EDO, PEG |
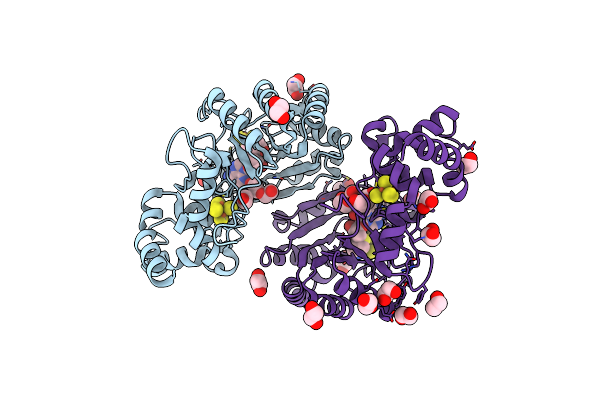 |
Crystal Structure Of Radical Sam Epimerase Epee D210A Mutant From Bacillus Subtilis With [4Fe-4S] Clusters, S-Adenosyl-L-Homocysteine And Persulfurated Cysteine Bound
Organism: Bacillus subtilis
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2024-01-10 Classification: METAL BINDING PROTEIN Ligands: SF4, SAH, EDO, PEG |
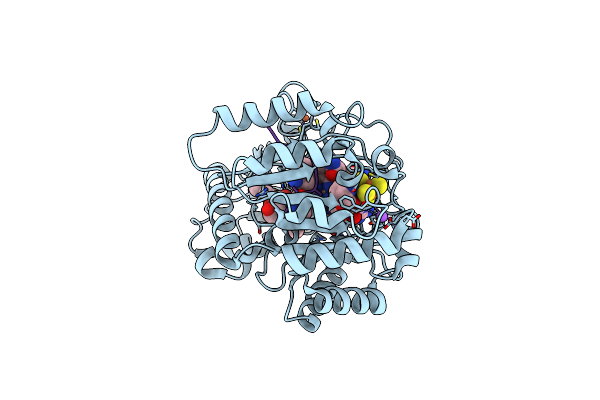 |
B12-Dependent Radical Sam Methyltransferase, Mmp10 With [4Fe-4S] Cluster, Cobalamin, S-Adenosyl-L-Methionine, And Peptide Bound.
Organism: Methanosarcina acetivorans
Method: X-RAY DIFFRACTION Resolution:2.33 Å Release Date: 2022-02-02 Classification: METAL BINDING PROTEIN Ligands: SF4, FE, SAM, COB, NA |
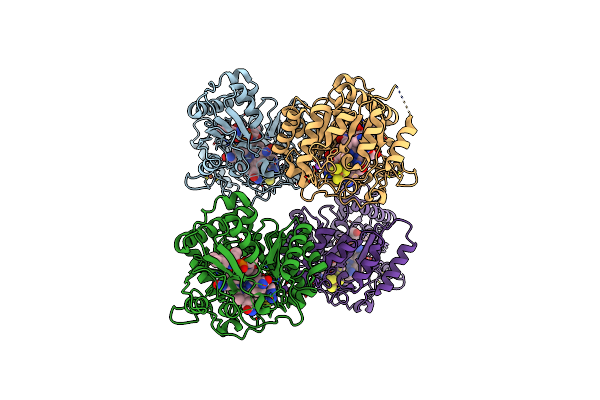 |
B12-Dependent Radical Sam Methyltransferase, Mmp10 With [4Fe-4S] Cluster, Cobalamin, And S-Methyl-5'-Thioadenosine Bound.
Organism: Methanosarcina acetivorans
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2022-02-02 Classification: METAL BINDING PROTEIN Ligands: SF4, FE, MTA, COB, NA |
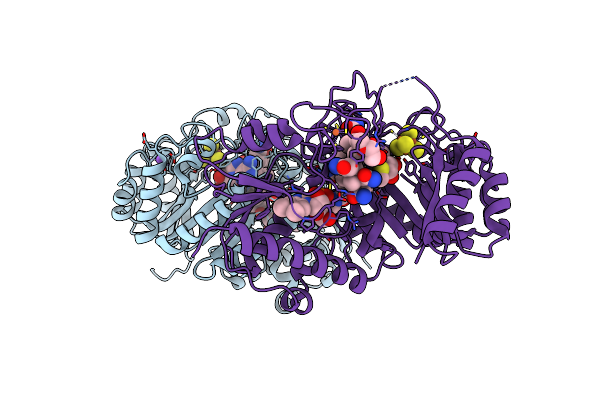 |
B12-Dependent Radical Sam Methyltransferase, Mmp10 With [4Fe-4S] Cluster, Cobalamin, And S-Methyl-5'-Thioadenosine Bound.
Organism: Methanosarcina acetivorans
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2022-02-02 Classification: METAL BINDING PROTEIN Ligands: SF4, FE, MTA, COB, NA, PEG |
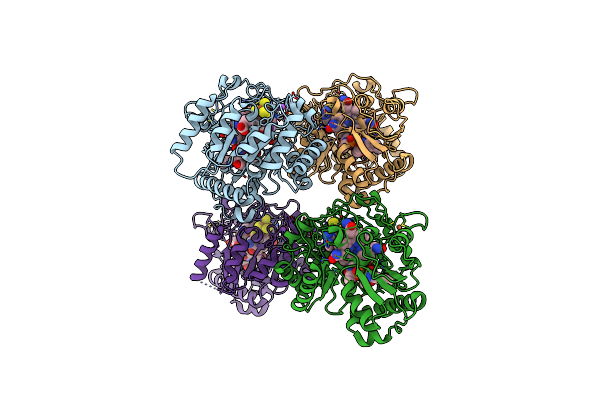 |
B12-Dependent Radical Sam Methyltransferase, Mmp10 With [4Fe-4S] Cluster, Cobalamin, And S-Adenosyl-L-Homocysteine Bound.
Organism: Methanosarcina acetivorans
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2022-02-02 Classification: METAL BINDING PROTEIN Ligands: SF4, FE, SAH, COB, NA |
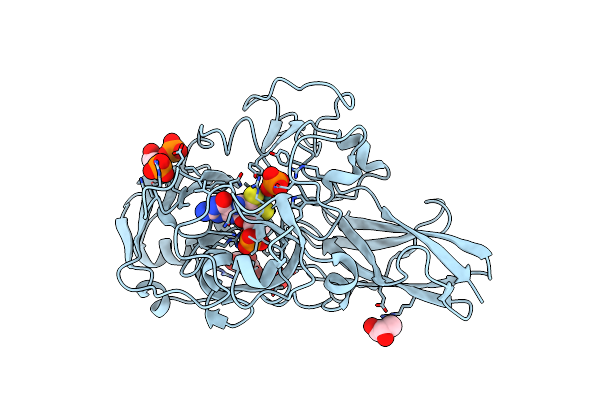 |
Organism: Thermus thermophilus (strain hb8 / atcc 27634 / dsm 579)
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2020-08-05 Classification: OXIDOREDUCTASE Ligands: MSS, GOL, PO4 |

