Search Count: 350
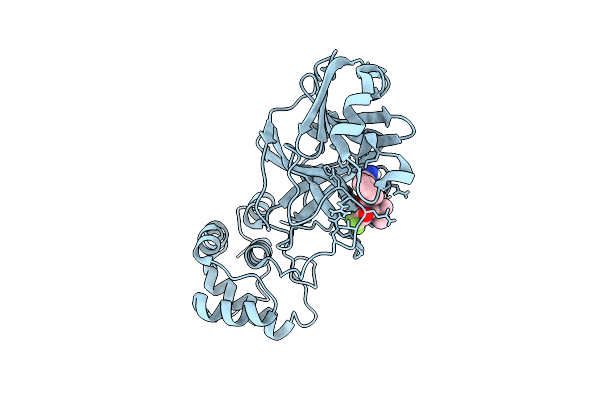 |
X-Ray Structure Of Sars-Cov-2 Main Protease V186G Covalently Bound To Compound Grl-051-22 At 1.3 A
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Release Date: 2025-11-26 Classification: VIRAL PROTEIN, HYDROLASE Ligands: A1BFE |
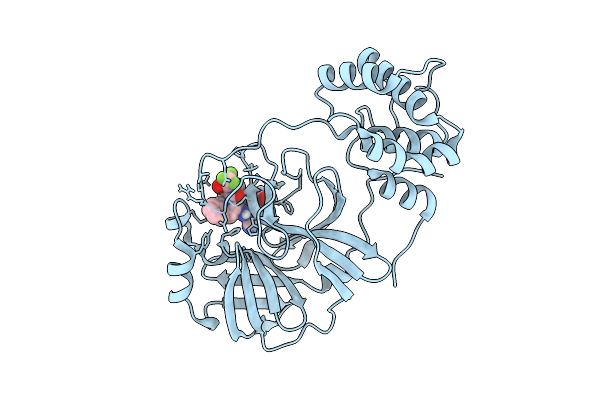 |
X-Ray Structure Of Sars-Cov-2 Main Protease Covalently Bound To Compound Grl-051-22 At 1.75 A.
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Release Date: 2025-11-26 Classification: VIRAL PROTEIN, HYDROLASE Ligands: A1BFE |
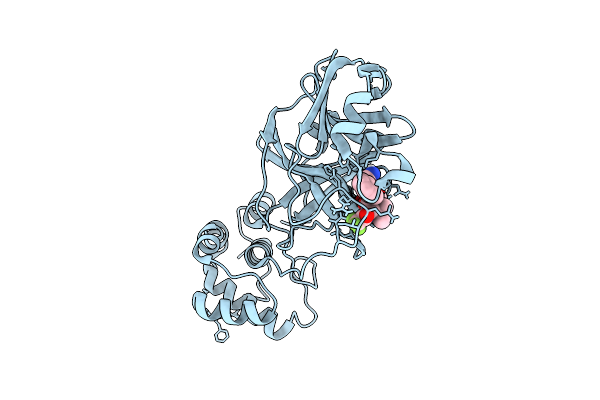 |
X-Ray Structure Of Sars-Cov-2 Main Protease T190I Covalently Bound To Compound Grl-051-22 At 1.5 A
Organism: Severe acute respiratory syndrome coronavirus
Method: X-RAY DIFFRACTION Release Date: 2025-11-26 Classification: VIRAL PROTEIN, HYDROLASE Ligands: A1BFE, NA |
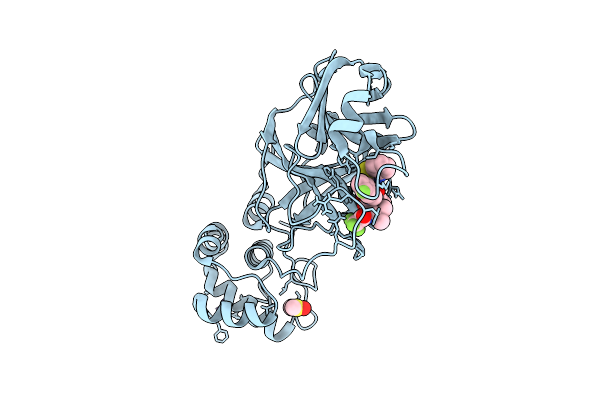 |
X-Ray Structure Of Sars-Cov-2 Main Protease Covalently Bound To Compound Grl-050-23 At 1.6 A
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Release Date: 2025-11-26 Classification: VIRAL PROTEIN Ligands: DMS, A1BMU, NA |
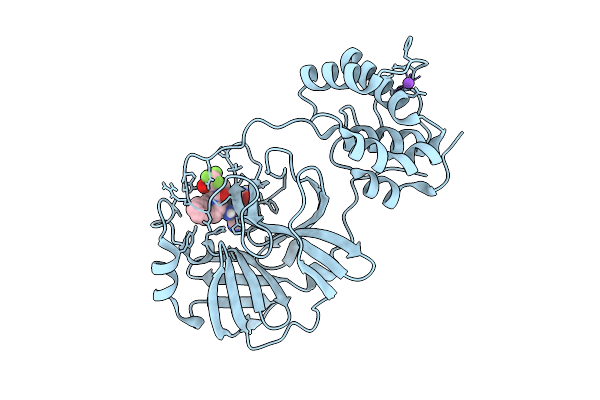 |
X-Ray Structure Of Sars-Cov-2 Main Protease M49I Covalently Bound To Inhibitor Grl-051-22 At 1.50 A
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Release Date: 2025-11-26 Classification: VIRAL PROTEIN Ligands: A1BFE, NA |
 |
X-Ray Structure Of Sars-Cov-2 Main Protease M165I Covalently Bound To Inhibitor Grl-051-22 At 1.90 A
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Release Date: 2025-11-26 Classification: VIRAL PROTEIN Ligands: A1BFE, NA |
 |
X-Ray Structure Of Sars-Cov-2 Main Protease V186F Covalently Bound To Inhibitor Grl-051-22 At 1.60 A
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Release Date: 2025-11-26 Classification: VIRAL PROTEIN, HYDROLASE Ligands: A1BFE, DMS |
 |
X-Ray Structure Of Sars-Cov-2 Main Protease V186I Covalently Bound To Inhibitor Grl-051-22 At 1.90 A
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Release Date: 2025-11-26 Classification: VIRAL PROTEIN, HYDROLASE Ligands: A1BFE, K |
 |
X-Ray Structure Of Sars-Cov-2 Main Protease V186F Covalently Bound To Inhibitor Grl-050-23 At 1.55 A
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Release Date: 2025-11-26 Classification: VIRAL PROTEIN, HYDROLASE Ligands: A1BMU |
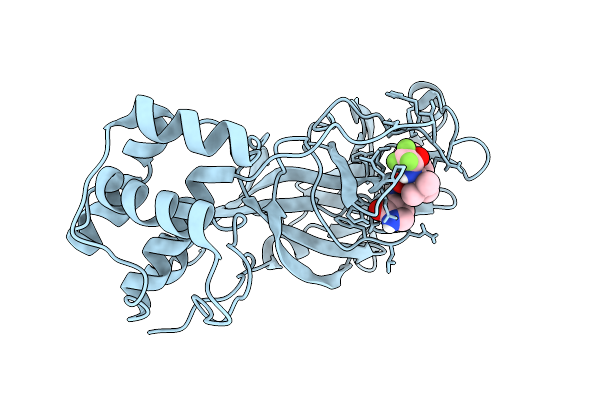 |
X-Ray Structure Of Sars-Cov-2 Main Protease V186G Covalently Bound To Inhibitor Nirmatrelvir At 1.81 A
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Release Date: 2025-11-26 Classification: VIRAL PROTEIN, HYDROLASE/INHIBITOR Ligands: 4WI |
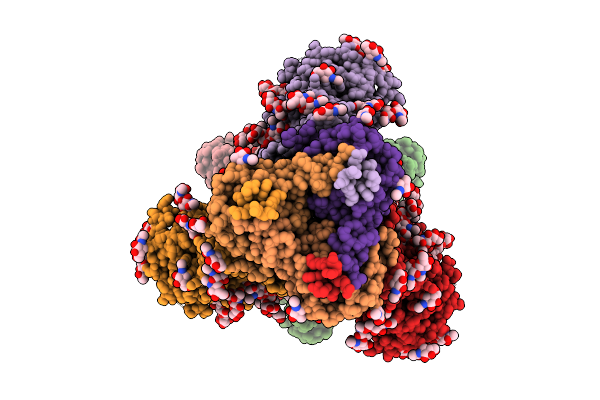 |
Cryo-Em Structure Of Vaccine-Elicited Antibody T6_P_H03 In Complex With Hiv Env Trimer Q23-Apex-Gt1
Organism: Mus musculus, Human immunodeficiency virus type 1
Method: ELECTRON MICROSCOPY Release Date: 2025-11-12 Classification: VIRAL PROTEIN Ligands: NAG |
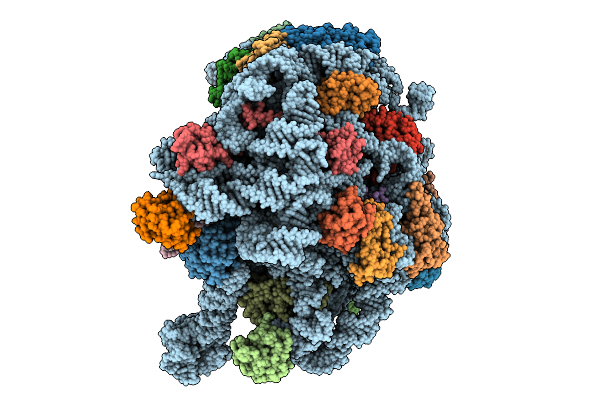 |
Organism: Methanosarcina acetivorans
Method: ELECTRON MICROSCOPY Release Date: 2025-09-24 Classification: RIBOSOME Ligands: MG, ZN |
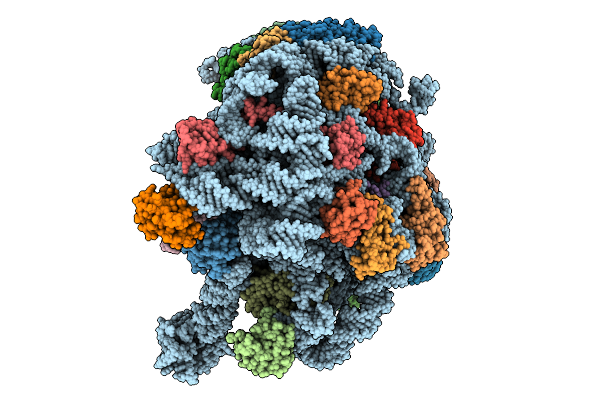 |
Organism: Methanosarcina acetivorans
Method: ELECTRON MICROSCOPY Release Date: 2025-09-24 Classification: RIBOSOME Ligands: MG, K, ZN |
 |
Organism: Methanosarcina acetivorans
Method: ELECTRON MICROSCOPY Release Date: 2025-09-24 Classification: RIBOSOME Ligands: MG, ZN |
 |
Organism: Methanosarcina acetivorans
Method: ELECTRON MICROSCOPY Release Date: 2025-09-24 Classification: RIBOSOME |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Release Date: 2025-09-10 Classification: TRANSFERASE Ligands: A1INI |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Release Date: 2025-09-10 Classification: TRANSFERASE Ligands: A1IOI |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Release Date: 2025-09-10 Classification: TRANSFERASE Ligands: A1IOP |
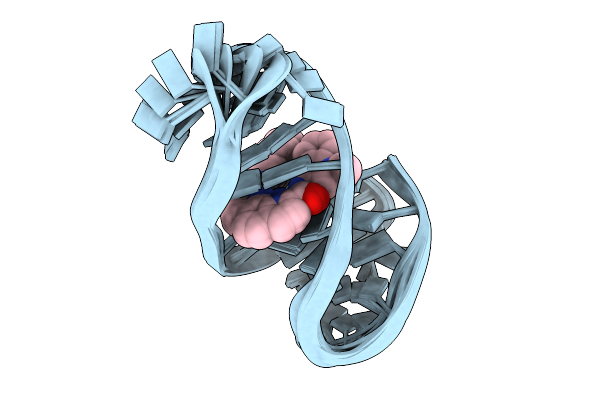 |
Organism: Homo sapiens
Method: SOLUTION NMR Release Date: 2025-08-27 Classification: DNA Ligands: PQ3 |
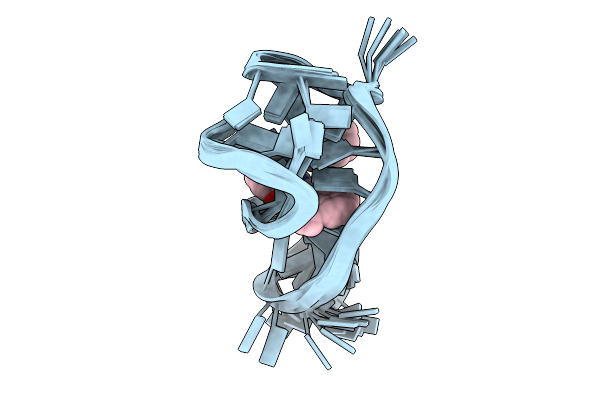 |
Organism: Homo sapiens
Method: SOLUTION NMR Release Date: 2025-08-27 Classification: DNA Ligands: LWQ |

