Search Count: 2,588
All
Selected
 |
Organism: Geobacillus stearothermophilus
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-02-25 Classification: DNA BINDING PROTEIN Ligands: EDO, PGE, CL |
 |
Organism: Geobacillus phage gbsv1
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: VIRAL PROTEIN |
 |
Crystal Structure Of Substrate Bound Gh5_22 Exo-Beta-Xylosidase From The Seaweed-Derived Thermophile Geobacillus Thermodenitrificans Os27
Organism: Geobacillus thermodenitrificans
Method: X-RAY DIFFRACTION Resolution:1.52 Å Release Date: 2026-02-04 Classification: HYDROLASE Ligands: GOL, FE |
 |
Crystal Structure Of Gh5_22 Exo-Beta-Xylosidase From The Seaweed-Derived Thermophile Geobacillus Thermodenitrificans Os27
Organism: Geobacillus thermodenitrificans
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2026-02-04 Classification: HYDROLASE Ligands: FE |
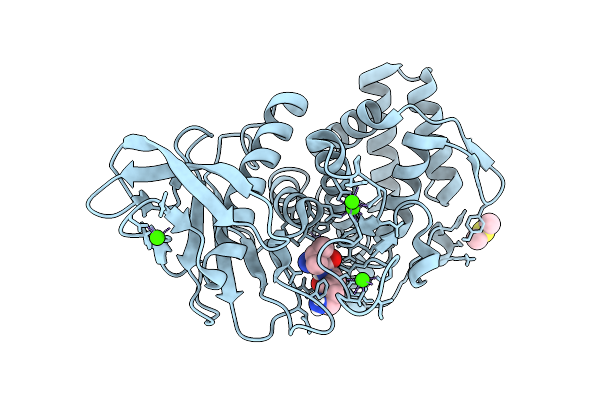 |
Gradient Equilibration Of Hexagonal Thermolysin To Low Salt Over 15 Minutes
Organism: Geobacillus stearothermophilus
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2025-10-15 Classification: HYDROLASE Ligands: VAL, LYS, CA, ZN, DMS |
 |
Biochemical And Structural Characterization Of A Novel 4-Hydroxyphenylacetate-3-Monooxygenase From Geobacillus Mahadii Geo-05
Organism: Geobacillus mahadia
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2025-09-10 Classification: HYDROLASE Ligands: PEU |
 |
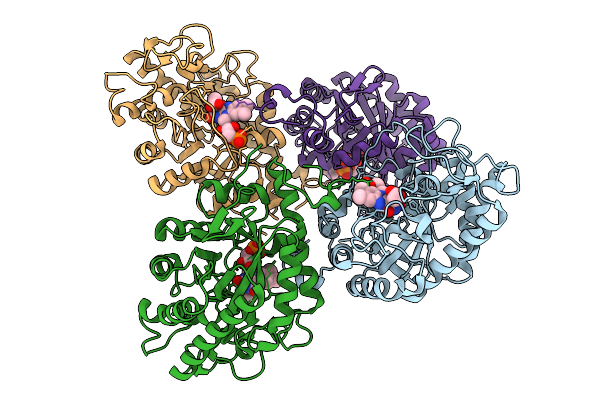
Organism: Geobacillus stearothermophilus
Method: X-RAY DIFFRACTION Resolution:1.49 Å Release Date: 2025-08-27 Classification: RNA BINDING PROTEIN |
 |
Organism: Parageobacillus thermantarcticus
Method: X-RAY DIFFRACTION Resolution:1.74 Å Release Date: 2025-08-27 Classification: OXIDOREDUCTASE Ligands: ACT, FMN |
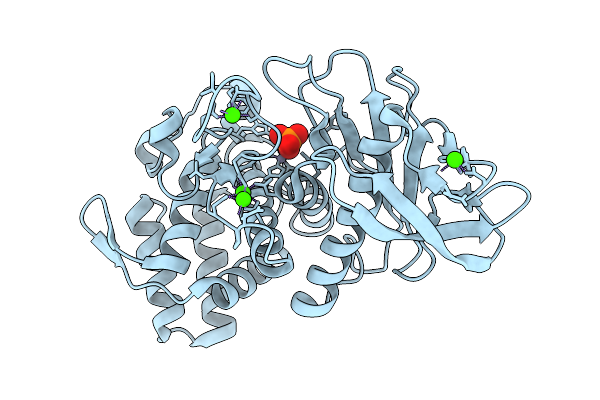 |
Organism: Geobacillus stearothermophilus
Method: X-RAY DIFFRACTION Resolution:1.73 Å Release Date: 2025-08-27 Classification: HYDROLASE Ligands: ZN, CA, PO4 |
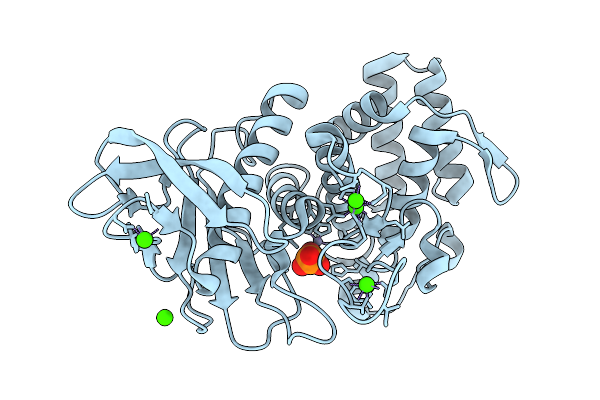 |
Organism: Geobacillus stearothermophilus
Method: X-RAY DIFFRACTION Resolution:1.66 Å Release Date: 2025-08-27 Classification: HYDROLASE Ligands: ZN, CA, PO4 |
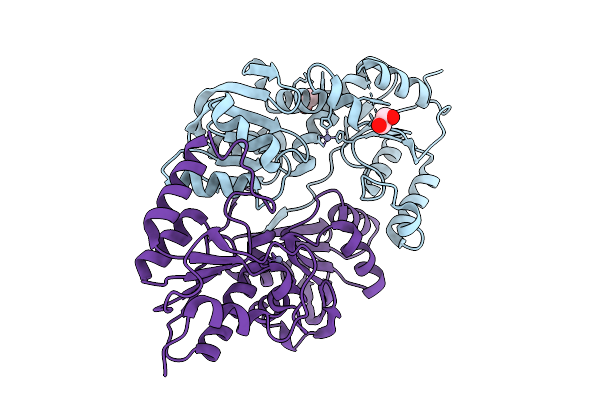 |
The Structure Of Xynx, A Nif3 Family Protein From Geobacillus Proteiniphilus T-6
Organism: Geobacillus proteiniphilus
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2025-08-06 Classification: UNKNOWN FUNCTION Ligands: EDO, ZN |
 |
Structure Of The Pro-Pro Endopeptidase (Ppep-3) E153A Y189F In Complex With Substrate Peptide Ac-Eplpppp-Nh2 From Geobacillus Thermodenitrificans
Organism: Geobacillus thermodenitrificans ng80-2, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-07-30 Classification: HYDROLASE Ligands: ZN, PG4, PGE |
 |
Structure Of The Pro-Pro Endopeptidase (Ppep-3) E153A Y189F From Geobacillus Thermodenitrificans
Organism: Geobacillus thermodenitrificans ng80-2
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2025-07-23 Classification: HYDROLASE Ligands: ZN, BCN, PG4, PEG |
 |
Structure Of The Pro-Pro Endopeptidase (Ppep-3) From Geobacillus Thermodenitrificans
Organism: Geobacillus thermodenitrificans ng80-2
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2025-07-16 Classification: HYDROLASE Ligands: ZN, BCN, PEG, EDO |
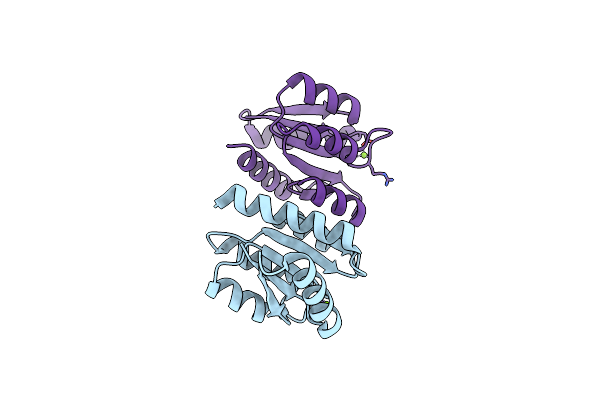 |
The Rec Domain (In The Non-Phosphorylated State) Of Xync, A Response Regulator From G.Proteiniphilus T-6
Organism: Geobacillus proteiniphilus
Method: X-RAY DIFFRACTION Resolution:2.35 Å Release Date: 2025-06-25 Classification: DNA BINDING PROTEIN Ligands: MG |
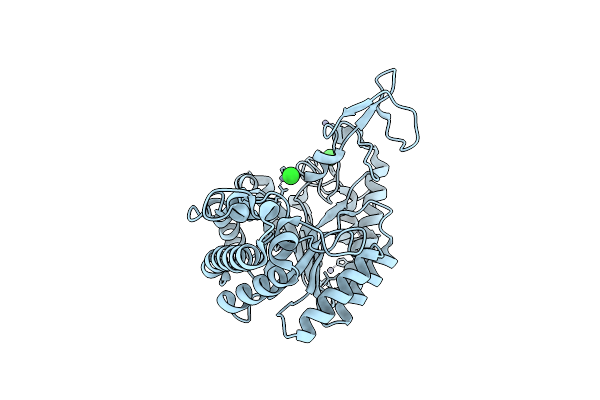 |
Organism: Geobacillus proteiniphilus
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2025-06-18 Classification: HYDROLASE Ligands: ZN, CL |
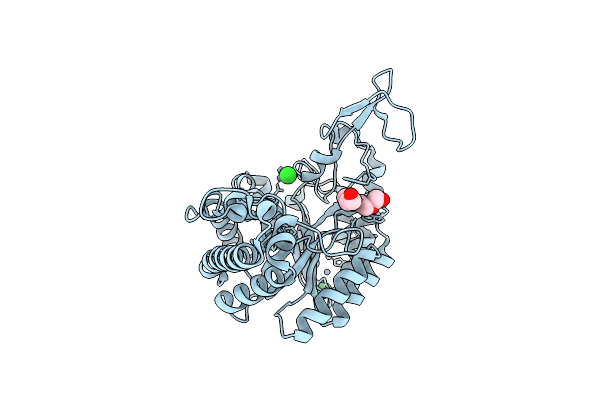 |
Organism: Geobacillus proteiniphilus
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2025-06-18 Classification: HYDROLASE Ligands: PG4, CL, ZN |
 |
Organism: Geobacillus proteiniphilus
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2025-06-18 Classification: HYDROLASE Ligands: ZN, CL |
 |
The Structure Of Xt6 From G.Proteiniphilus T-6: The E265G/Q238A/W241A Mutant
Organism: Geobacillus proteiniphilus
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2025-06-18 Classification: HYDROLASE Ligands: CL, ZN |
 |
The Structure Of Glycosynthase Ixt6 (E241G Mutant), The Intracellular Xylanase Of G.Proteiniphilus T-6
Organism: Geobacillus proteiniphilus
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2025-06-18 Classification: HYDROLASE |

