Search Count: 1,963
All
Selected
 |
Organism: Bacterium
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-03-04 Classification: HYDROLASE Ligands: CIT, EDO, CA |
 |
Organism: Catenovulum maritimum
Method: X-RAY DIFFRACTION Resolution:1.64 Å Release Date: 2026-03-04 Classification: HYDROLASE Ligands: EDO, O4B, MG, SO4, NA, CL, EPE, PEG |
 |
Crystal Structure Of The Complex Of The Gh16 Carrageenase Sfgh16 From Saccharicrinis Fermentans With An Oligotetrasaccharide Of Iota-Carrageenan
Organism: Saccharicrinis fermentans dsm 9555 = jcm 21142
Method: X-RAY DIFFRACTION Resolution:1.44 Å Release Date: 2026-03-04 Classification: HYDROLASE |
 |
Crystal Structure Of Substrate Bound Gh5_22 Exo-Beta-Xylosidase From The Seaweed-Derived Thermophile Geobacillus Thermodenitrificans Os27
Organism: Geobacillus thermodenitrificans
Method: X-RAY DIFFRACTION Resolution:1.52 Å Release Date: 2026-02-04 Classification: HYDROLASE Ligands: GOL, FE |
 |
Crystal Structure Of Gh5_22 Exo-Beta-Xylosidase From The Seaweed-Derived Thermophile Geobacillus Thermodenitrificans Os27
Organism: Geobacillus thermodenitrificans
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2026-02-04 Classification: HYDROLASE Ligands: FE |
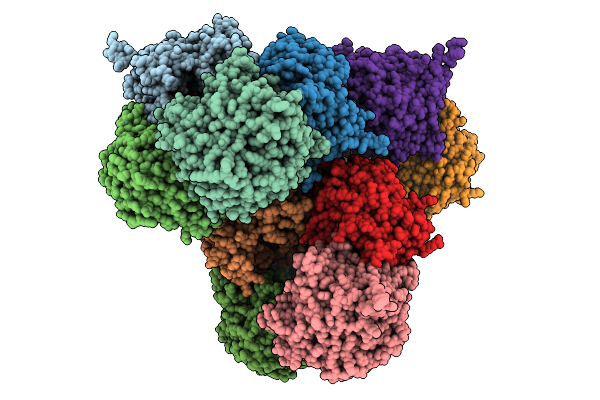 |
Organism: Prevotella
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-01-28 Classification: HYDROLASE Ligands: GOL |
 |
Sfx Crystal Structure Of Hen Egg-White Lysozyme (Hewl) Using A Guanosine Derivative Hydrogel As Crystal Matrix
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2026-01-28 Classification: HYDROLASE |
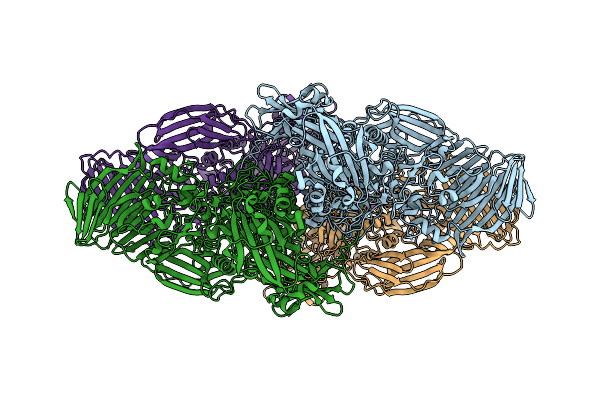 |
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: HYDROLASE Ligands: MG, NA |
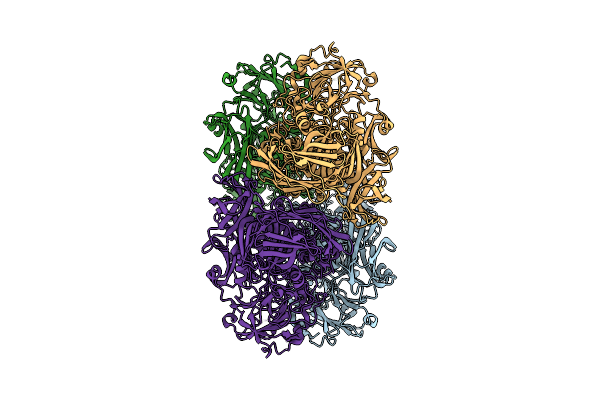 |
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:3.70 Å Release Date: 2025-12-17 Classification: HYDROLASE |
 |
Organism: Microbacterium oxydans
Method: X-RAY DIFFRACTION Resolution:1.92 Å Release Date: 2025-12-17 Classification: HYDROLASE Ligands: EDO |
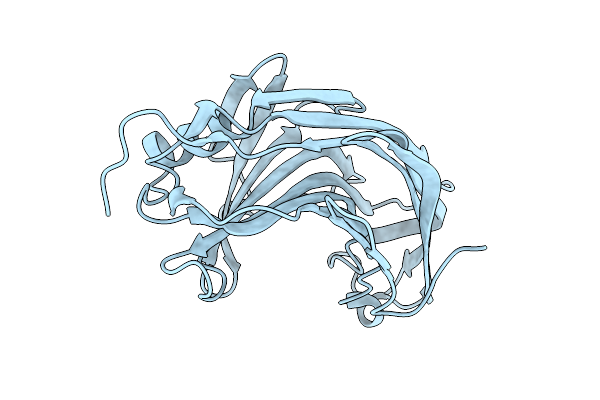 |
Crystal Structure Of The Pathogen-Secreted Apoplastic Gh12 Xyloglucan-Specific Endoglucanase Xeg1
Organism: Phytophthora sojae (strain p6497)
Method: X-RAY DIFFRACTION Resolution:2.07 Å Release Date: 2025-12-10 Classification: HYDROLASE |
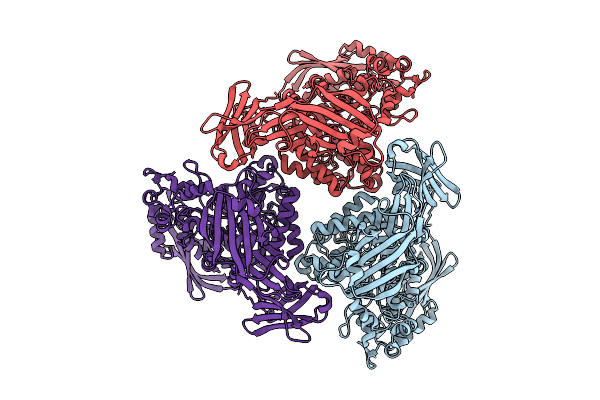 |
Active Conformation Of A Redox-Regulated Glycoside Hydrolase (Capgh2B) From The Gh2 Family
Organism: Metagenome
Method: ELECTRON MICROSCOPY Resolution:2.62 Å Release Date: 2025-11-12 Classification: HYDROLASE |
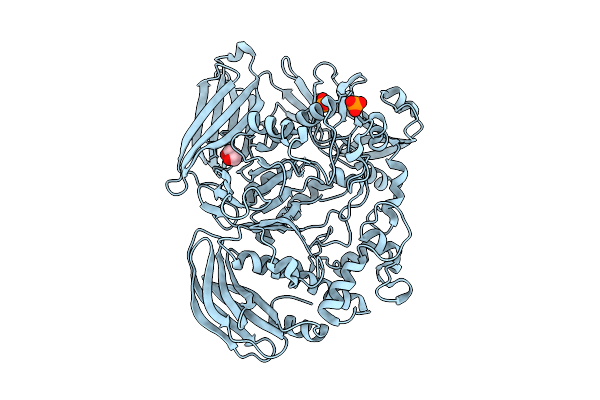 |
Crystal Structure Of The Inactive Conformation Of A Glycoside Hydrolase (Capgh2B) From The Gh2 Family In The Space Group I213 At 2.05 A
Organism: Metagenome
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-11-12 Classification: HYDROLASE Ligands: PO4, GOL |
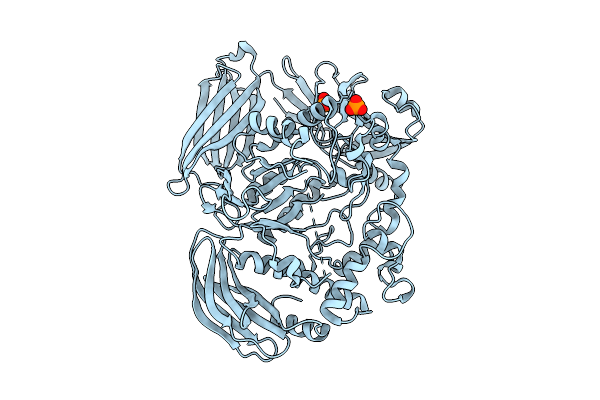 |
Crystal Structure Of The Inactive Conformation Of A Glycoside Hydrolase (Capgh2B - E553Q Mutant) From The Gh2 Family In The Space Group I213 At 2.6 A
Organism: Metagenome
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2025-11-12 Classification: HYDROLASE Ligands: PO4 |
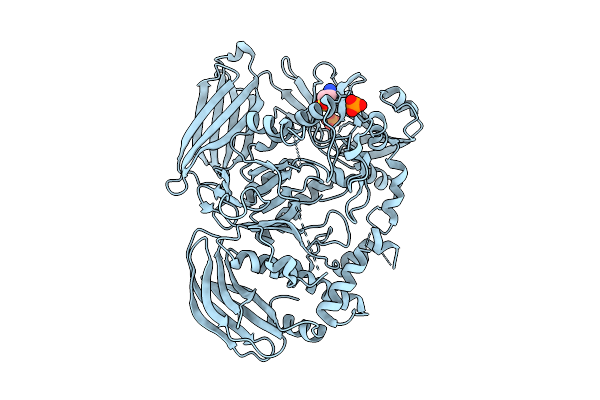 |
Crystal Structure Of The Inactive Conformation Of A Glycoside Hydrolase (Capgh2B) From The Gh2 Family In The Space Group I213 At 2.75 A
Organism: Metagenome
Method: X-RAY DIFFRACTION Resolution:2.75 Å Release Date: 2025-11-12 Classification: HYDROLASE Ligands: PO4, TAU |
 |
Crystal Structure Of The Inactive Conformation Of A Glycoside Hydrolase (Capgh2B) From The Gh2 Family In The Space Group R3 At 2.45 A
Organism: Metagenome
Method: X-RAY DIFFRACTION Resolution:2.45 Å Release Date: 2025-11-12 Classification: HYDROLASE |
 |
Crystal Structure Of The Inactive Conformation Of A Glycoside Hydrolase (Capgh2B) From The Gh2 Family In The Space Group I212121 At 2.65 A
Organism: Metagenome
Method: X-RAY DIFFRACTION Resolution:2.65 Å Release Date: 2025-11-12 Classification: HYDROLASE Ligands: PO4, PEG, PG4 |
 |
Crystal Structure Of The Inactive Conformation Of A Glycoside Hydrolase (Capgh2B - E553Q Mutant) From The Gh2 Family In The Space Group P3121 At 3.05 A
Organism: Metagenome
Method: X-RAY DIFFRACTION Resolution:3.05 Å Release Date: 2025-11-12 Classification: HYDROLASE Ligands: PO4, ACT, MLI |
 |
Crystal Structure Of The Inactive Conformation Of A Glycoside Hydrolase (Capgh2B) From The Gh2 Family In The Space Group P1 At 2.40 A
Organism: Metagenome
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2025-11-12 Classification: HYDROLASE Ligands: PO4, GOL |
 |
Crystal Structure Of The Inactive Conformation Of A Glycoside Hydrolase (Capgh2B) From The Gh2 Family In The Space Group P1 At 2.15 A
Organism: Metagenome
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2025-11-12 Classification: HYDROLASE Ligands: PO4, GOL, ACT |

