Search Count: 35
All
Selected
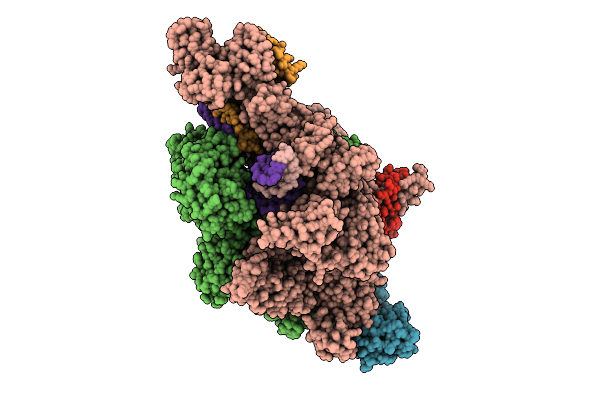 |
Gtp Bound In De Novo Transcription Initiation T. Thermophilus Rna Polymerase Complex With Tc Dna Template
Organism: Thermus thermophilus hb8, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:2.76 Å Release Date: 2025-12-17 Classification: TRANSCRIPTION Ligands: MG, ZN, GTP, 2TM |
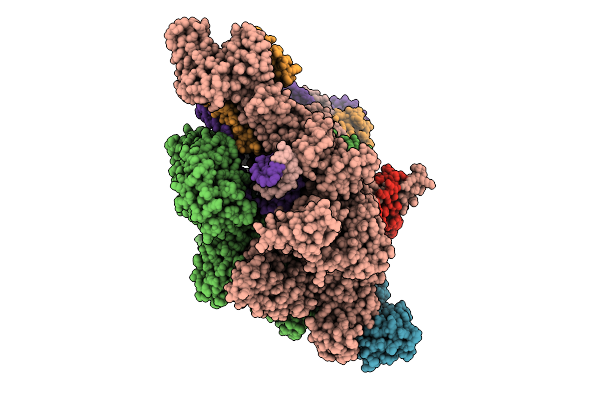 |
Ap3G Bound In De Novo Transcription Initiation T. Thermophilus Rna Polymerase Complex With Tc Dna Template
Organism: Thermus thermophilus hb8, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:2.51 Å Release Date: 2025-12-17 Classification: TRANSCRIPTION Ligands: MG, ZN, 2TM, G3A |
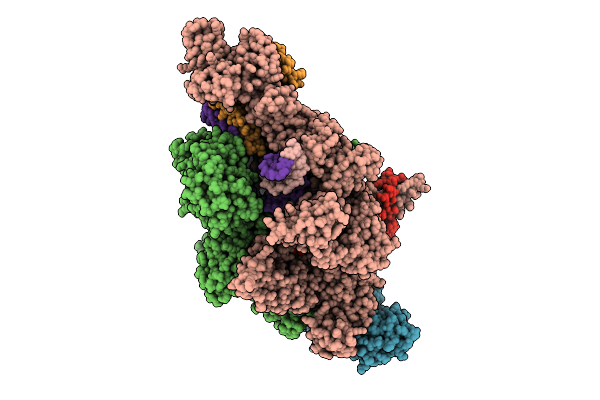 |
Ap4G Bound In De Novo Transcription Initiation T. Thermophilus Rna Polymerase Complex With Tc Dna Template
Organism: Thermus thermophilus hb8, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.00 Å Release Date: 2025-12-17 Classification: TRANSCRIPTION Ligands: MG, ZN, 2TM, A1IF9 |
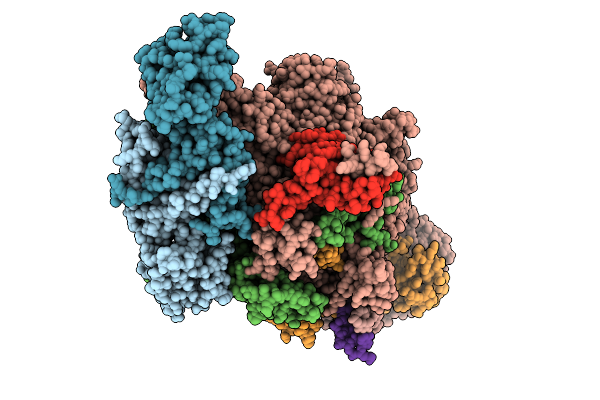 |
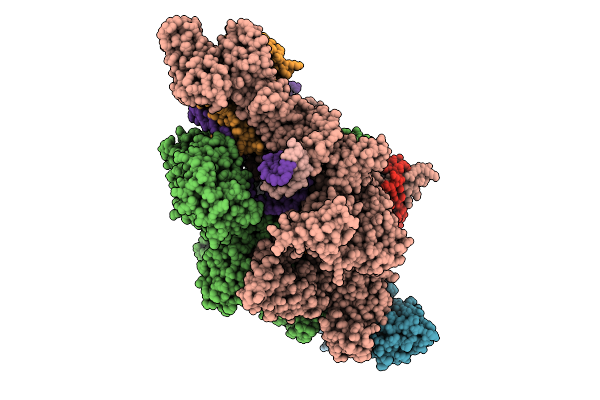
Ap4A Bound In De Novo Transcription Initiation T. Thermophilus Rna Polymerase Complex With Tc Dna Template
Organism: Thermus thermophilus hb8, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:2.42 Å Release Date: 2025-12-17 Classification: TRANSCRIPTION Ligands: MG, B4P, ZN, G2P |
 |
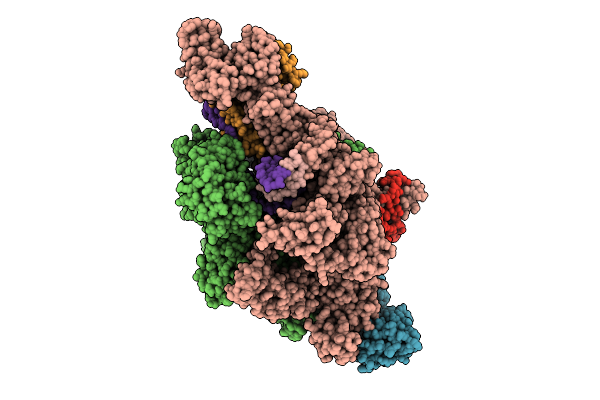
Ap4A Bound In De Novo Transcription Initiation T. Thermophilus Rna Polymerase Complex With Att Dna Template
Organism: Thermus thermophilus hb8, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:2.66 Å Release Date: 2025-12-17 Classification: TRANSCRIPTION Ligands: MG, ZN, 2TM, B4P |
 |
Transcription Initiation T. Thermophilus Rna Polymerase Complex With Anti-Scrunched Tc Dna Template
Organism: Thermus thermophilus hb8, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.80 Å Release Date: 2025-12-17 Classification: TRANSCRIPTION Ligands: ZN, MG |
 |
Organism: Mycolicibacterium smegmatis
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2025-08-27 Classification: OXIDOREDUCTASE Ligands: IMP |
 |
Crystal Structure Of M. Smegmatis Gmp Reductase With Xmp* Intermidiate In Complex With Nadp+ And Imp.
Organism: Mycolicibacterium smegmatis
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2025-08-27 Classification: OXIDOREDUCTASE Ligands: IMP, NAP |
 |
Crystal Structure Of M. Smegmatis Gmp Reductase In Complex With Gmp And Atp.
Organism: Mycolicibacterium smegmatis
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2025-08-27 Classification: OXIDOREDUCTASE Ligands: ATP, 5GP |
 |
Crystal Structure Of M. Smegmatis Gmp Reductase In Complex With Imp And Atp.
Organism: Mycolicibacterium smegmatis
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2025-08-27 Classification: OXIDOREDUCTASE Ligands: IMP, ATP |
 |
Crystal Structure Of M. Smegmatis Gmp Reductase In Complex With Gmp And Gtp.
Organism: Mycolicibacterium smegmatis
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2025-08-27 Classification: OXIDOREDUCTASE Ligands: 5GP, GTP |
 |
Cryoem Structure Of M. Smegmatis Gmp Reductase In Complex With Gmp At Ph 6.6, Extended Conformation.
Organism: Mycolicibacterium smegmatis
Method: ELECTRON MICROSCOPY Release Date: 2025-08-20 Classification: OXIDOREDUCTASE Ligands: 5GP |
 |
Cryoem Structure Of M. Smegmatis Gmp Reductase In Complex With Gmp At Ph 6.6, Compressed Conformation.
Organism: Mycolicibacterium smegmatis
Method: ELECTRON MICROSCOPY Release Date: 2025-08-20 Classification: OXIDOREDUCTASE Ligands: 5GP |
 |
Cryoem Structure Of M. Smegmatis Gmp Reductase In Complex With Gmp And Gtp At Ph 6.6, Extended Conformation I.
Organism: Mycolicibacterium smegmatis
Method: ELECTRON MICROSCOPY Release Date: 2025-08-20 Classification: OXIDOREDUCTASE Ligands: 5GP, GTP |
 |
Cryoem Structure Of M. Smegmatis Gmp Reductase In Complex With Gmp And Gtp At Ph 6.6, Extended Conformation Ii.
Organism: Mycolicibacterium smegmatis
Method: ELECTRON MICROSCOPY Release Date: 2025-08-20 Classification: OXIDOREDUCTASE Ligands: 5GP, GTP |
 |
Cryoem Structure Of M. Smegmatis Gmp Reductase In Complex With Gmp And Atp At Ph 6.6, Compressed Conformation.
Organism: Mycolicibacterium smegmatis
Method: ELECTRON MICROSCOPY Release Date: 2025-08-20 Classification: OXIDOREDUCTASE Ligands: 5GP, ATP |
 |
Cryoem Structure Of M. Smegmatis Gmp Reductase Apoform At Ph 6.6, Extended Conformation I.
Organism: Mycolicibacterium smegmatis
Method: ELECTRON MICROSCOPY Release Date: 2025-08-20 Classification: OXIDOREDUCTASE |
 |
Cryoem Structure Of M. Smegmatis Gmp Reductase Apoform At Ph 6.6, Extended Conformation Ii.
Organism: Mycolicibacterium smegmatis
Method: ELECTRON MICROSCOPY Release Date: 2025-08-20 Classification: OXIDOREDUCTASE |
 |
Cryoem Structure Of M. Smegmatis Gmp Reductase Apoform At Ph 7.8, Extended Conformation I.
Organism: Mycolicibacterium smegmatis
Method: ELECTRON MICROSCOPY Release Date: 2025-08-20 Classification: OXIDOREDUCTASE |
 |
Cryoem Structure Of M. Smegmatis Gmp Reductase Apoform At Ph 7.8, Extended Conformation Ii.
Organism: Mycolicibacterium smegmatis
Method: ELECTRON MICROSCOPY Release Date: 2025-08-20 Classification: OXIDOREDUCTASE |

