Search Count: 3,177
 |
Organism: Synechococcus sp. pcc 7335
Method: ELECTRON MICROSCOPY Release Date: 2026-01-14 Classification: OXIDOREDUCTASE Ligands: HEM, FAD, FMN, NAP |
 |
Organism: Synechococcus sp. pcc 7335
Method: ELECTRON MICROSCOPY Release Date: 2026-01-14 Classification: OXIDOREDUCTASE Ligands: HEM, FAD, FMN |
 |
Organism: Synechococcus sp. pcc 7335
Method: ELECTRON MICROSCOPY Release Date: 2026-01-14 Classification: OXIDOREDUCTASE Ligands: HEM, FAD, FMN |
 |
Organism: Synechococcus sp. pcc 7335
Method: ELECTRON MICROSCOPY Release Date: 2026-01-14 Classification: OXIDOREDUCTASE Ligands: HEM, FAD, FMN |
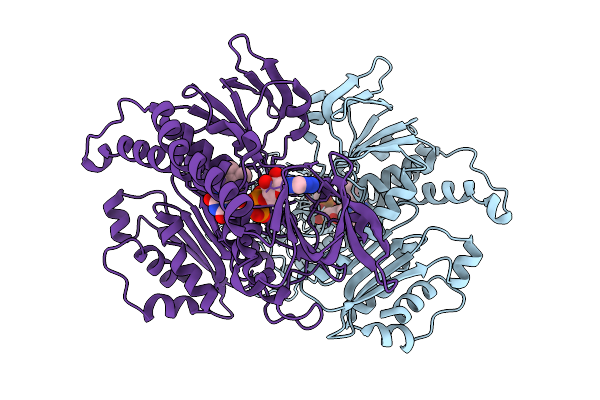 |
Organism: Mycobacteroides abscessus
Method: X-RAY DIFFRACTION Release Date: 2026-01-07 Classification: OXIDOREDUCTASE Ligands: FAD |
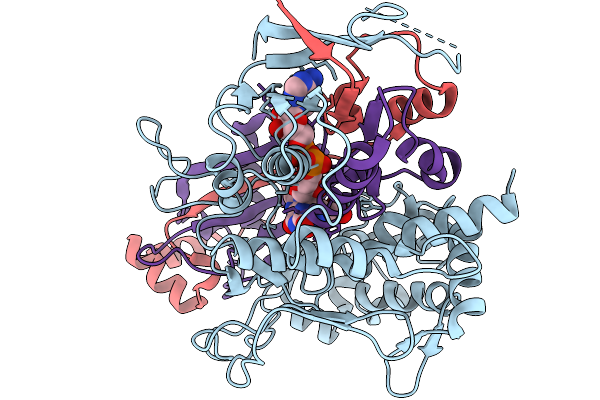 |
Organism: Streptomyces sp. x-119-6
Method: X-RAY DIFFRACTION Release Date: 2026-01-07 Classification: OXIDOREDUCTASE Ligands: FAD, GLU |
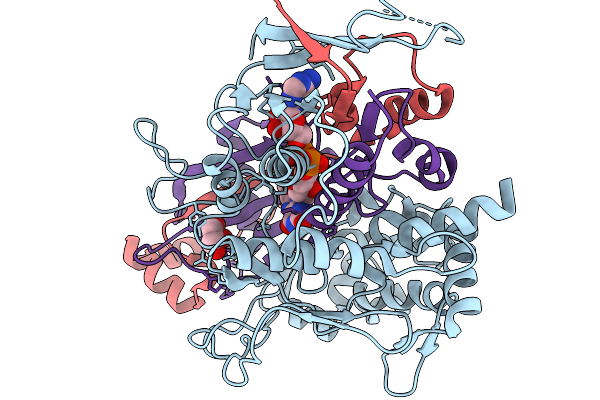 |
Organism: Streptomyces sp. x-119-6
Method: X-RAY DIFFRACTION Release Date: 2026-01-07 Classification: OXIDOREDUCTASE Ligands: FAD, GOL |
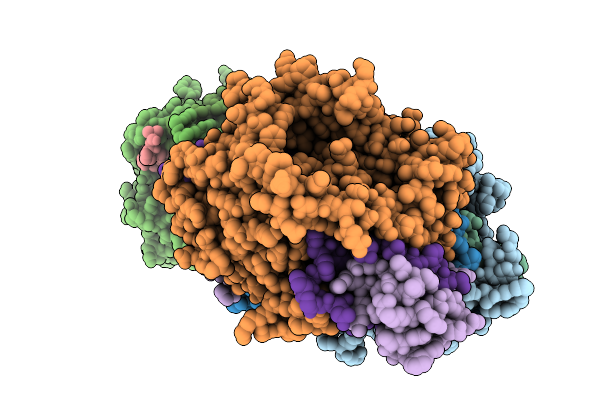 |
Organism: Streptomyces sp. x-119-6
Method: X-RAY DIFFRACTION Release Date: 2026-01-07 Classification: OXIDOREDUCTASE Ligands: FAD, GLU |
 |
Organism: Streptomyces sp. x-119-6
Method: X-RAY DIFFRACTION Release Date: 2026-01-07 Classification: OXIDOREDUCTASE Ligands: FAD |
 |
Organism: Streptomyces sp. x-119-6
Method: X-RAY DIFFRACTION Release Date: 2026-01-07 Classification: OXIDOREDUCTASE Ligands: FAD |
 |
Xfel Structure Of Hnqo1 Mixed With Nadh In An Extended Orientation At 0.3 S
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Release Date: 2025-12-10 Classification: OXIDOREDUCTASE Ligands: NAD, FAD, ACT |
 |
Organism: Parastagonospora nodorum sn15
Method: X-RAY DIFFRACTION Release Date: 2025-12-10 Classification: OXIDOREDUCTASE Ligands: FAD, GOL |
 |
Organism: Acinetobacter baumannii
Method: X-RAY DIFFRACTION Release Date: 2025-12-10 Classification: OXIDOREDUCTASE Ligands: FAD |
 |
Organism: Streptomyces lavendulae
Method: X-RAY DIFFRACTION Release Date: 2025-12-10 Classification: OXIDOREDUCTASE Ligands: FAD, EPE, PEG, ACT |
 |
Organism: Aetokthonos hydrillicola thurmond2011
Method: X-RAY DIFFRACTION Release Date: 2025-12-10 Classification: FLAVOPROTEIN Ligands: FAD, TRP |
 |
Organism: Aetokthonos hydrillicola thurmond2011
Method: X-RAY DIFFRACTION Release Date: 2025-12-10 Classification: FLAVOPROTEIN Ligands: FAD, TRP |
 |
Organism: Aetokthonos hydrillicola thurmond2011
Method: X-RAY DIFFRACTION Release Date: 2025-12-10 Classification: FLAVOPROTEIN Ligands: FAD, TRP |
 |
Crystal Structure Of Aetf-L183F/V220I/S523A In Complex With Fad And L-Tryptophan
Organism: Aetokthonos hydrillicola thurmond2011
Method: X-RAY DIFFRACTION Release Date: 2025-12-10 Classification: FLAVOPROTEIN Ligands: TRP, FAD |
 |
Organism: Porcellio dilatatus
Method: X-RAY DIFFRACTION Release Date: 2025-12-03 Classification: OXIDOREDUCTASE Ligands: FAD, FE |
 |
Crystal Structure Of A D-Lactate Dehydrogenase In Complex With D-Lactate From Porcellio Dilatatus
Organism: Porcellio dilatatus
Method: X-RAY DIFFRACTION Release Date: 2025-12-03 Classification: OXIDOREDUCTASE Ligands: GOL, LAC, FAD, FE |

