Search Count: 41
All
Selected
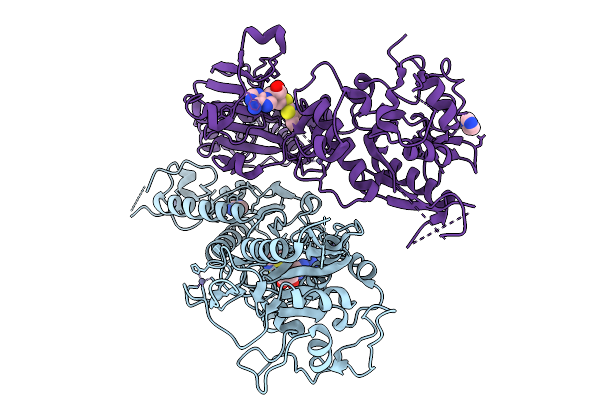 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:2.85 Å Release Date: 2026-01-21 Classification: VIRAL PROTEIN Ligands: ZN, IMD, A1JKS |
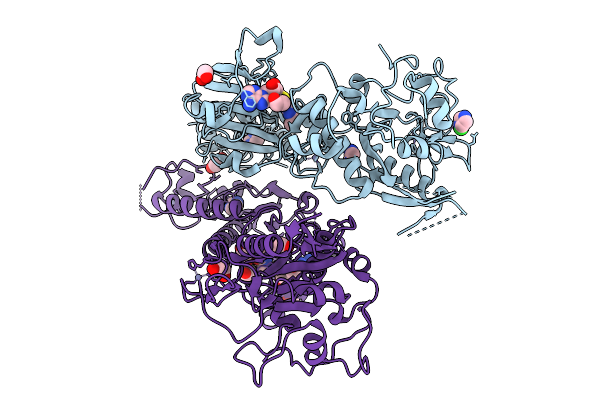 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:2.48 Å Release Date: 2026-01-21 Classification: VIRAL PROTEIN Ligands: ZN, IMD, EDO, CL, A1JMY |
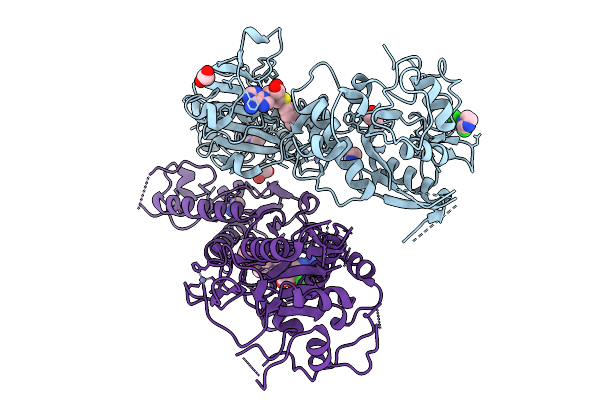 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2026-01-21 Classification: VIRAL PROTEIN Ligands: A1JMZ, IMD, EDO, CL, ZN |
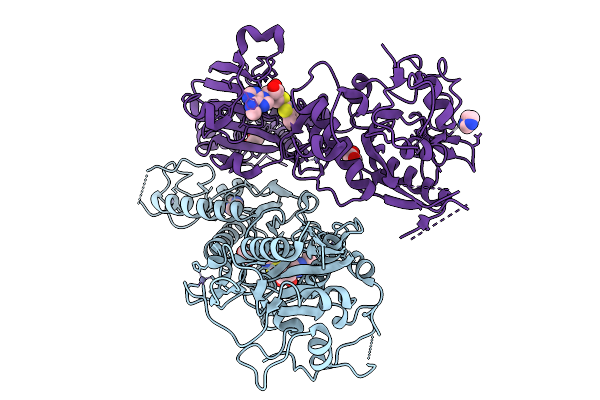 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:2.29 Å Release Date: 2026-01-21 Classification: VIRAL PROTEIN Ligands: A1JM0, IMD, EDO, ZN |
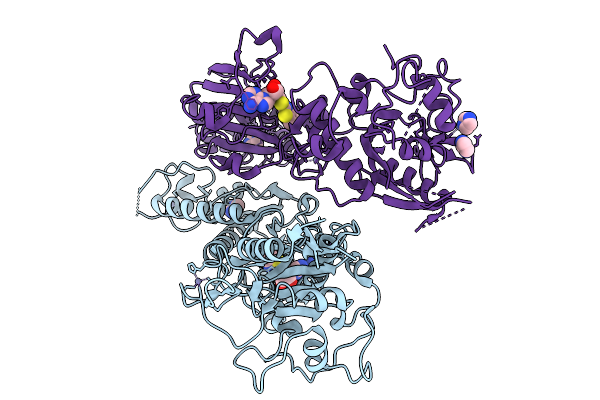 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:2.54 Å Release Date: 2026-01-21 Classification: VIRAL PROTEIN Ligands: ZN, IMD, A1JM1 |
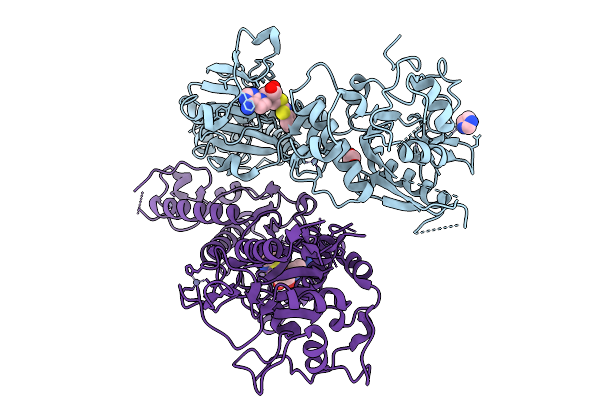 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:2.73 Å Release Date: 2026-01-21 Classification: VIRAL PROTEIN Ligands: A1JM2, IMD, EDO, ZN |
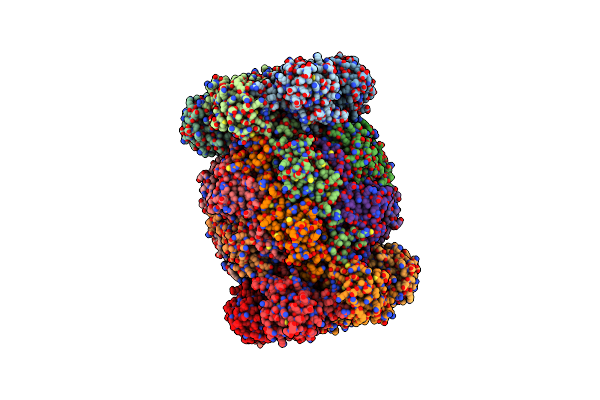 |
Cryoem Structure Of Recombinant Trypanosoma Cruzi Apo Proteasome 20S Subunit
Organism: Trypanosoma cruzi
Method: ELECTRON MICROSCOPY Resolution:2.25 Å Release Date: 2024-12-25 Classification: UNKNOWN FUNCTION |
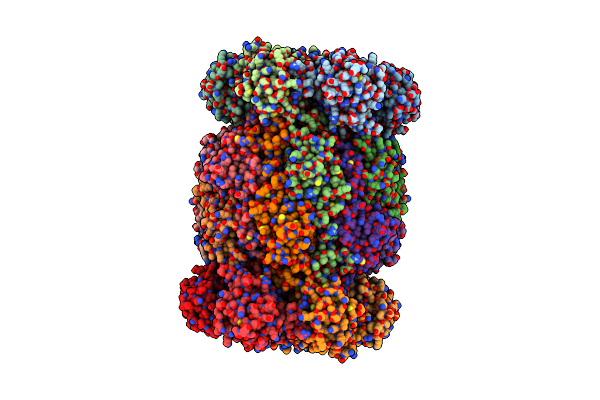 |
Organism: Trypanosoma cruzi
Method: ELECTRON MICROSCOPY Resolution:2.31 Å Release Date: 2024-12-25 Classification: UNKNOWN FUNCTION |
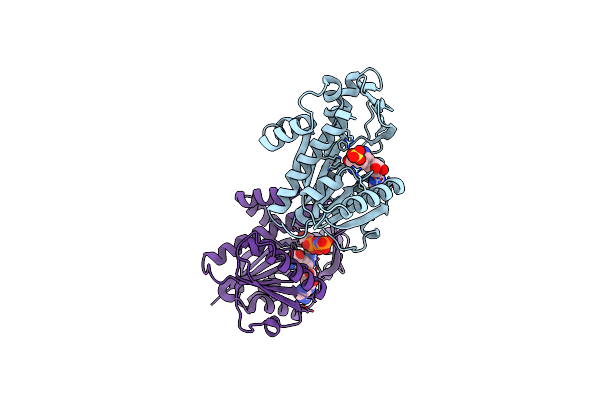 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2024-09-11 Classification: RNA BINDING PROTEIN Ligands: SFG, SO4, GNP |
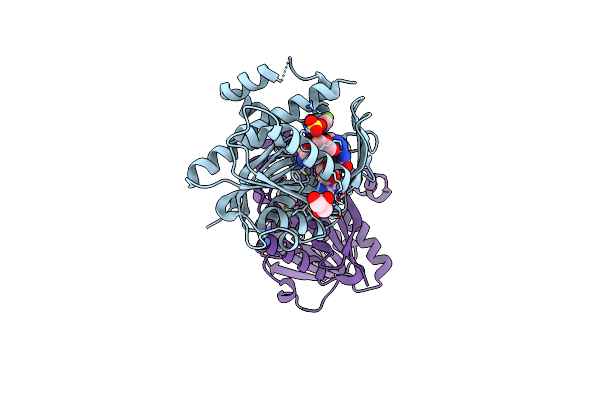 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2024-08-28 Classification: RNA BINDING PROTEIN Ligands: SAH, O94, SO4, GOL |
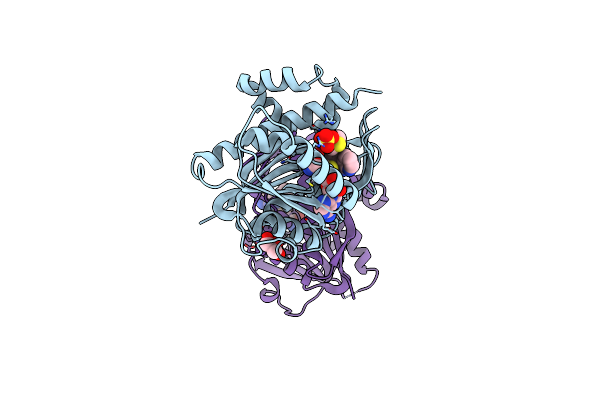 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.96 Å Release Date: 2024-08-21 Classification: RNA BINDING PROTEIN Ligands: SAH, K7R, SO4, GOL |
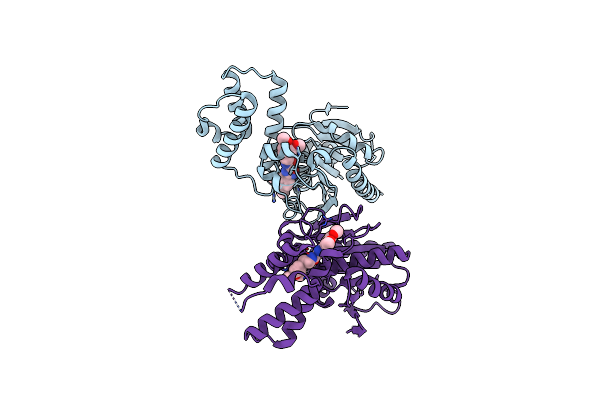 |
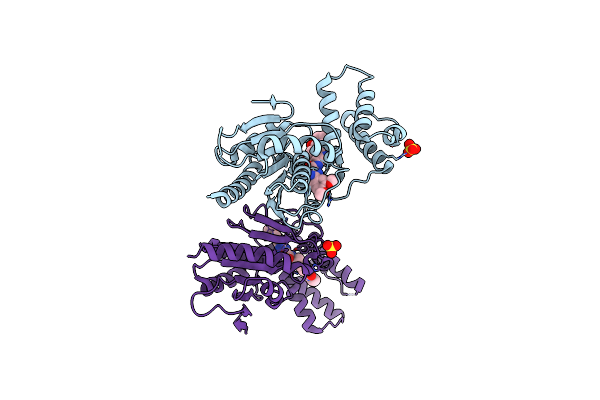
Identification And Optimisation Of Novel Inhibitors Of The Polyketide Synthetase 13 Thioesterase Domain With Antitubercular Activity
Organism: Mycobacterium tuberculosis
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2023-11-22 Classification: TRANSFERASE Ligands: IJJ |
 |
Identification And Optimisation Of Novel Inhibitors Of The Polyketide Synthetase 13 Thioesterase Domain With Antitubercular Activity
Organism: Mycobacterium tuberculosis
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2023-11-22 Classification: TRANSFERASE Ligands: SO4, IKG |
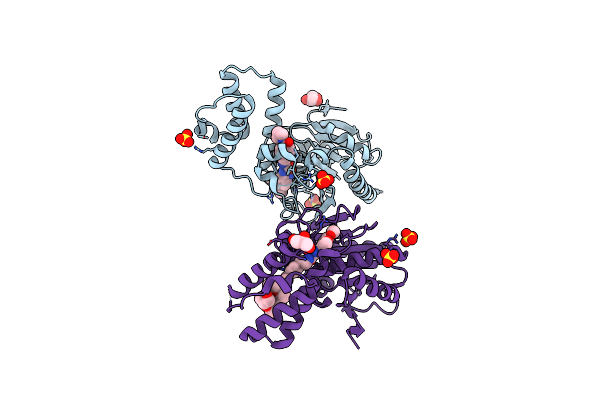 |
Identification And Optimisation Of Novel Inhibitors Of The Polyketide Synthetase 13 Thioesterase Domain With Antitubercular Activity
Organism: Mycobacterium tuberculosis
Method: X-RAY DIFFRACTION Resolution:1.71 Å Release Date: 2023-11-22 Classification: TRANSFERASE Ligands: SO4, IL8, GOL, 2Q5 |
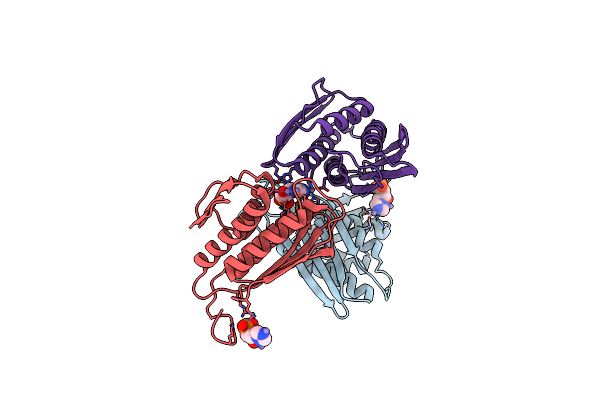 |
Crystal Structure Of Igpd From Pyrococcus Furiosus In Complex With (R)-C348
Organism: Pyrococcus furiosus
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2016-10-05 Classification: LYASE Ligands: MN, 5LD |
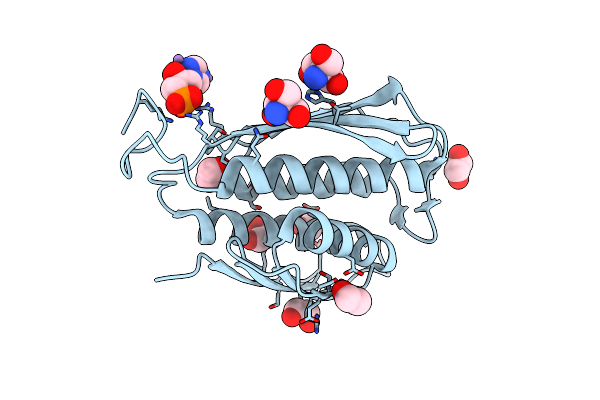 |
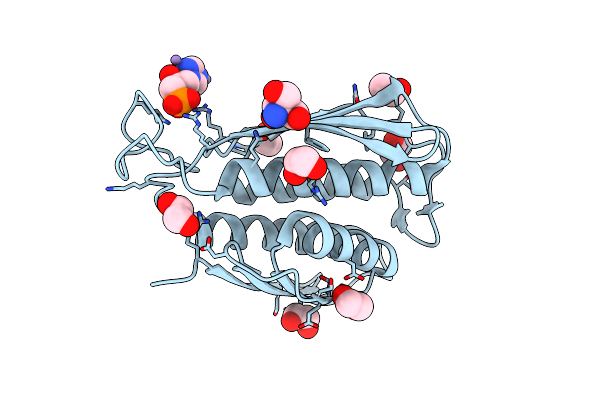
A. Thaliana Igpd2 In Complex With The Racemate Of The Triazole-Phosphonate Inhibitor, C348
Organism: Arabidopsis thaliana
Method: X-RAY DIFFRACTION Resolution:1.10 Å Release Date: 2016-10-05 Classification: LYASE Ligands: MN, 5DL, 5LD, EDO, TRS, CL |
 |
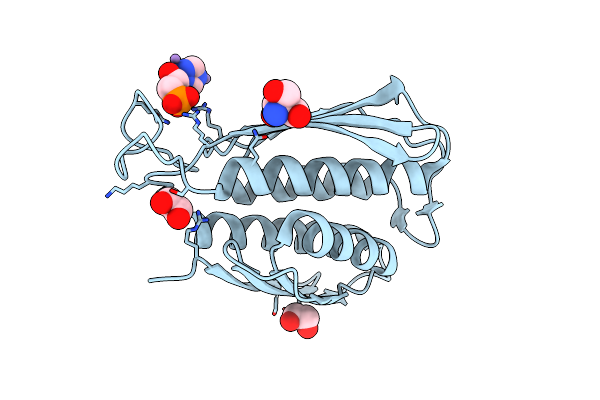
A. Thaliana Igpd2 In Complex With The Triazole-Phosphonate Inhibitor, (S)-C348, To 1.1A Resolution
Organism: Arabidopsis thaliana
Method: X-RAY DIFFRACTION Resolution:1.10 Å Release Date: 2016-10-05 Classification: LYASE Ligands: MN, 5DL, TRS, EDO |
 |
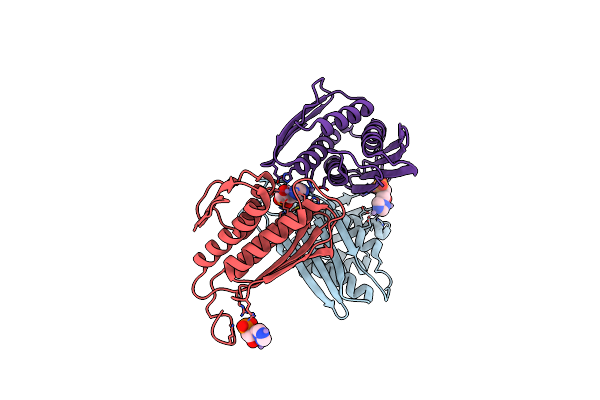
A. Thaliana Igpd2 In Complex With The Triazole-Phosphonate Inhibitor, (R)-C348, To 1.36A Resolution
Organism: Arabidopsis thaliana
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2016-10-05 Classification: LYASE Ligands: MN, 5LD, EDO, TRS, CL |
 |
Crystal Structure Of Igpd From Pyrococcus Furiosus In Complex With (S)-C348
Organism: Pyrococcus furiosus
Method: X-RAY DIFFRACTION Resolution:1.53 Å Release Date: 2016-09-28 Classification: LYASE Ligands: MN, 5DL |
 |
The Structure Of Wt A. Thaliana Igpd2 In Complex With Mn2+ And Formate At 1.3A Resolution
Organism: Arabidopsis thaliana
Method: X-RAY DIFFRACTION Resolution:1.30 Å Release Date: 2015-06-24 Classification: LYASE Ligands: MN, FMT, TRS, CL |

