Search Count: 34
 |
Organism: Solanum tuberosum
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2025-05-14 Classification: PLANT PROTEIN Ligands: GOL, ETE, MAN |
 |
Promiscuous Amino Acid Gamma Synthase From Caldicellulosiruptor Hydrothermalis In Closed Conformation
Organism: Caldicellulosiruptor hydrothermalis
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2023-12-27 Classification: TRANSFERASE Ligands: MES, NA, AE3, ETE, 1PE |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2023-02-08 Classification: IMMUNE SYSTEM Ligands: ETE |
 |
Organism: Escherichia coli k-12, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2021-05-19 Classification: TRANSPORT PROTEIN Ligands: LMT, HEX, GOL, ETE, 3YI, EDO, DDR, DDQ, OCT, D10, CL |
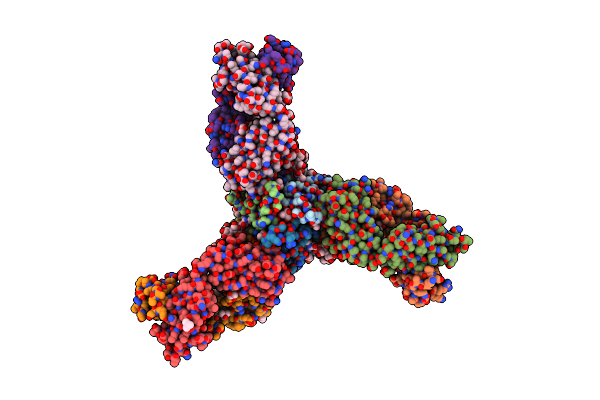 |
Crystal Structure Of The Fab Fragment Of Humanized 5C8 Antibody Containing The Fluorescent Non-Canonical Amino Acid L-(7-Hydroxycoumarin-4-Yl)Ethylglycine In Complex With Cd40L At Ph 6.8
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.82 Å Release Date: 2020-12-23 Classification: IMMUNE SYSTEM Ligands: PEG, TRS, TMO, ETE, 1PE |
 |
Mus Musculus Acetylcholinesterase In Complex With N-(2-(Diethylamino)Ethyl)-1-(4-(Trifluoromethyl)Phenyl)Methanesulfonamide
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2020-10-14 Classification: HYDROLASE Ligands: NAG, N2K, EDO, PG0, PE4, ETE |
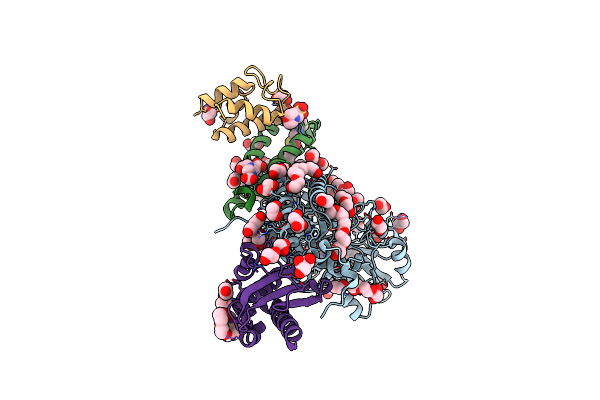 |
Structure Of Human Mitochondrial Complex Nfs1-Iscu2-Isd11 With E.Coli Acp1 At 1.95 A Resolution (Niau)2. N-Terminal Mutation Of Iscu2 (L35) Traps Nfs1 Cys Loop In The Active Site Of Iscu2 Without Metal Present.
Organism: Homo sapiens, Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2020-04-22 Classification: TRANSFERASE Ligands: PLP, EDO, GOL, PEG, PG4, ETE, PGE, MES, EDT, 8Q1, 1PE |
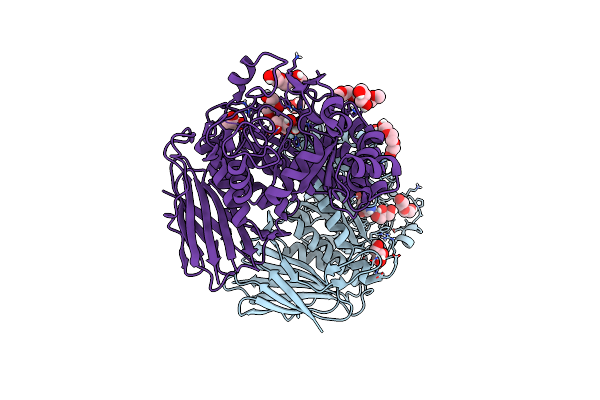 |
Organism: Geobacillus stearothermophilus
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2020-02-26 Classification: HYDROLASE Ligands: LXE, GOL, PEG, PGE, ETE |
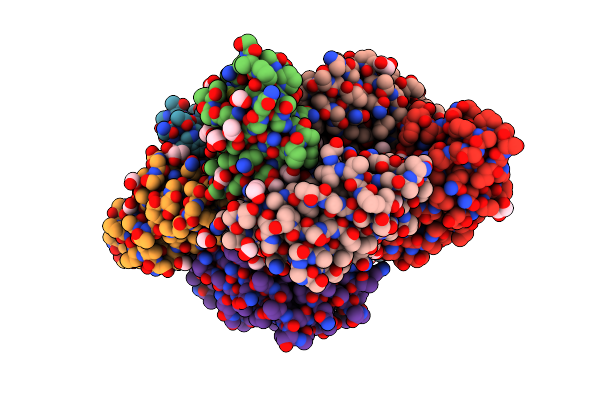 |
Organism: Cellulomonas fimi
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2019-12-18 Classification: SUGAR BINDING PROTEIN Ligands: ACT, EDO, FMT, 1PS, PEG, ETE, GOL |
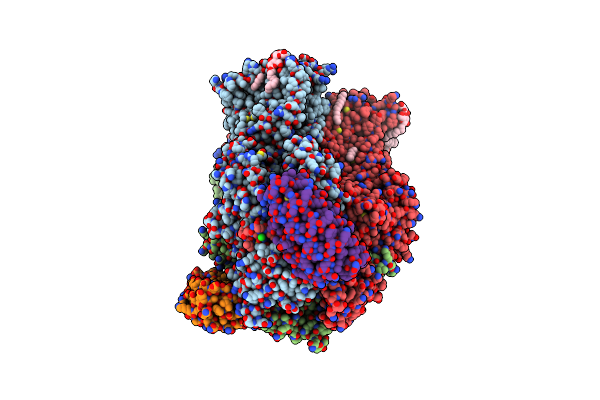 |
Organism: Escherichia coli k-12, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2019-11-13 Classification: TRANSPORT PROTEIN Ligands: LMT, D12, DDQ, GOL, HEX, FUA, PTY, DDR, OCT, CL, SO4, D10, C14, ETE |
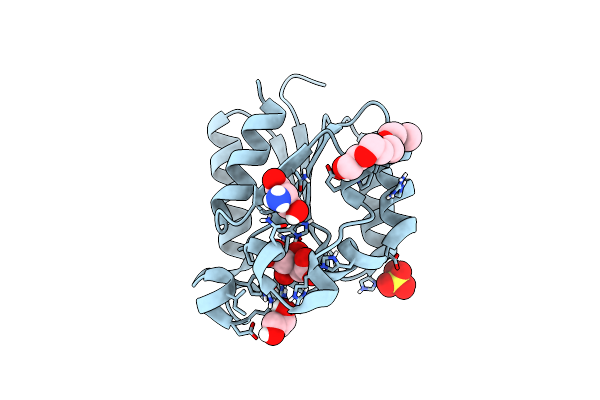 |
The Crystal Structure Of Type Ii Dehydroquinase From Butyrivibrio Crossotus Dsm 2876
Organism: Butyrivibrio crossotus dsm 2876
Method: X-RAY DIFFRACTION Resolution:0.92 Å Release Date: 2019-10-23 Classification: BIOSYNTHETIC PROTEIN Ligands: ETE, SER, SO4, FLC, GOL |
 |
Organism: Vaccinia virus (strain western reserve)
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2019-03-06 Classification: VIRAL PROTEIN Ligands: PG4, PG0, PEG, EDO, GOL, PDO, EOH, MOH, PGO, ETX, ETE, PGE, P33, P6G |
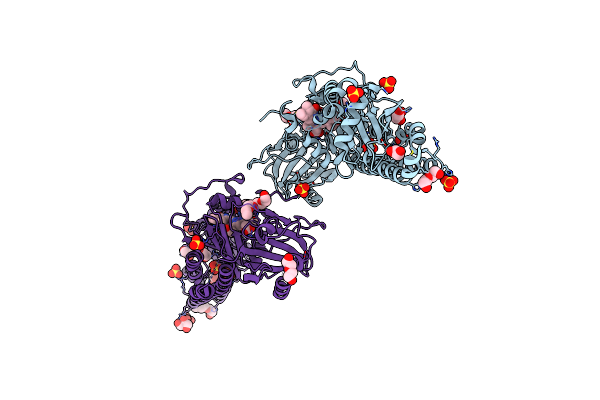 |
Crystal Structure Of A Fluorescence Optimized Bathy Phytochrome Pairfp2 Derived From Wild-Type Agp2 In Its Pfr State.
Organism: Agrobacterium fabrum (strain c58 / atcc 33970)
Method: X-RAY DIFFRACTION Resolution:2.03 Å Release Date: 2018-11-28 Classification: FLUORESCENT PROTEIN Ligands: EL5, SO4, GOL, PEG, ETE, PG0 |
 |
Crystal Structure Of A Fluorescence Optimized Bathy Phytochrome Pairfp2 Derived From Wild-Type Agp2 In Its Functional Meta-F Intermediate State.
Organism: Agrobacterium fabrum (strain c58 / atcc 33970)
Method: X-RAY DIFFRACTION Resolution:2.16 Å Release Date: 2018-11-28 Classification: FLUORESCENT PROTEIN Ligands: EL5, SO4, CL, GOL, PEG, ETE, PG0 |
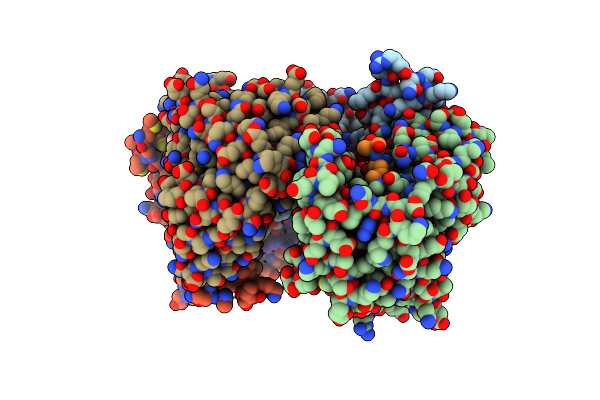 |
Crystal Structure Of Human 14-3-3 Beta In Complex With Cftr R-Domain Peptide Ps753-Ps768
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.86 Å Release Date: 2018-10-24 Classification: SIGNALING PROTEIN Ligands: ETE |
 |
Organism: Petromyzon marinus
Method: X-RAY DIFFRACTION Resolution:1.10 Å Release Date: 2018-05-09 Classification: IMMUNE SYSTEM Ligands: SO4, ETE |
 |
Crystal Structure Of Blac From Mycobacterium Tuberculosis Bound To Phosphate
Organism: Mycobacterium tuberculosis
Method: X-RAY DIFFRACTION Resolution:1.19 Å Release Date: 2017-11-15 Classification: HYDROLASE Ligands: PO4, ACT, ETE, AE4 |
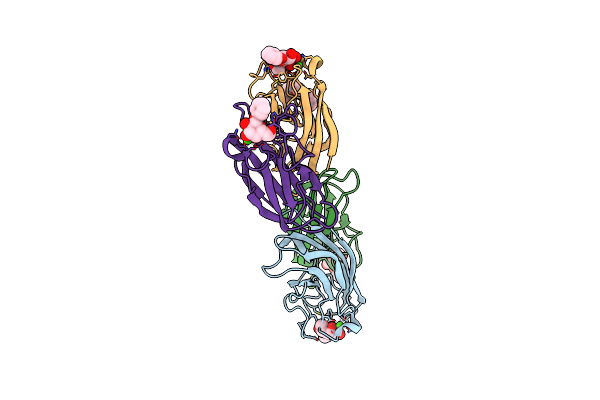 |
Crystal Structure Of The Lectin Leca From Pseudomonas Aeruginosa In Complex With A Phenyl-Epoxy-Galactopyranoside
Organism: Pseudomonas aeruginosa (strain atcc 15692 / dsm 22644 / cip 104116 / jcm 14847 / lmg 12228 / 1c / prs 101 / pao1)
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2017-10-11 Classification: SUGAR BINDING PROTEIN Ligands: CA, 7NU, ETE |
 |
Organism: Rattus norvegicus
Method: X-RAY DIFFRACTION Resolution:1.23 Å Release Date: 2017-02-22 Classification: TRANSPORT PROTEIN Ligands: FWD, PO4, PG4, PEG, TOE, PG0, ETE, EDO, NA |
 |
S1 Nuclease From Aspergillus Oryzae In Complex With Phosphate And Adenosine 5'-Monophosphate
Organism: Aspergillus oryzae rib40
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2016-12-28 Classification: HYDROLASE Ligands: ZN, NAG, PO4, AMP, NA, CA, BTB, ETE |

