Search Count: 32
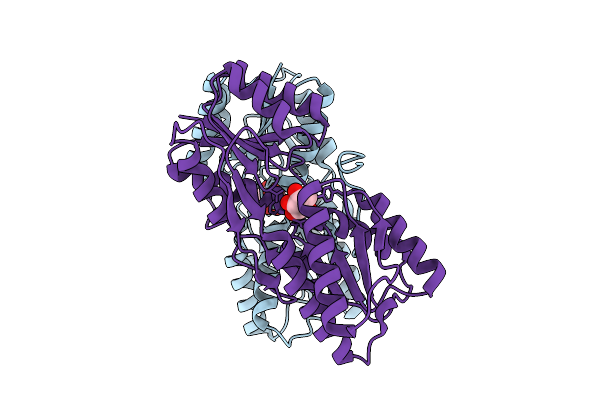 |
Crystal Structure Of A Allulose Transcriptional Regulator From Agrobacterium Fabrum
Organism: Agrobacterium fabrum str. c58
Method: X-RAY DIFFRACTION Release Date: 2025-11-26 Classification: TRANSCRIPTION Ligands: A1EGE |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Release Date: 2025-07-16 Classification: PEPTIDE BINDING PROTEIN Ligands: EDO, IMD, PEG |
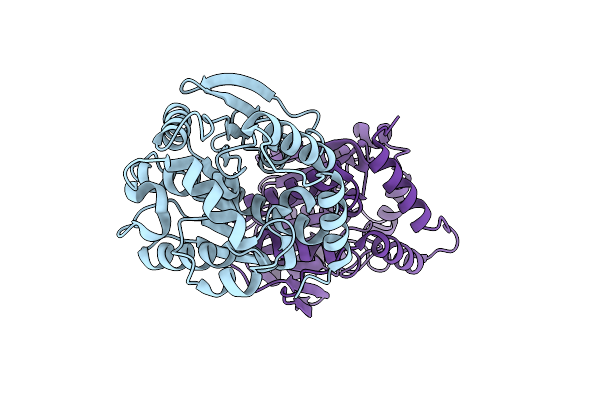 |
Organism: Paracoccus kondratievae
Method: X-RAY DIFFRACTION Resolution:1.69 Å Release Date: 2024-07-31 Classification: HYDROLASE |
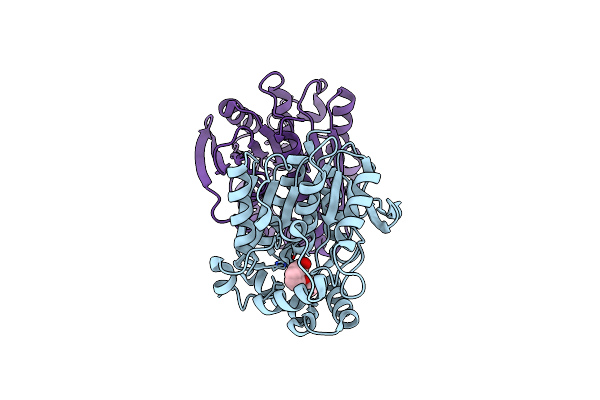 |
Organism: Paracoccus kondratievae
Method: X-RAY DIFFRACTION Resolution:1.76 Å Release Date: 2024-07-31 Classification: HYDROLASE Ligands: A1D6Y |
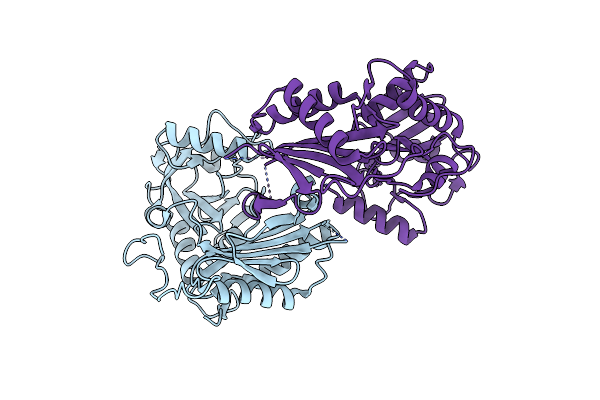 |
Organism: Mycolicibacterium vanbaalenii (strain dsm 7251 / jcm 13017 / bcrc 16820 / kctc 9966 / nrrl b-24157 / pyr-1)
Method: X-RAY DIFFRACTION Release Date: 2024-01-24 Classification: TRANSFERASE |
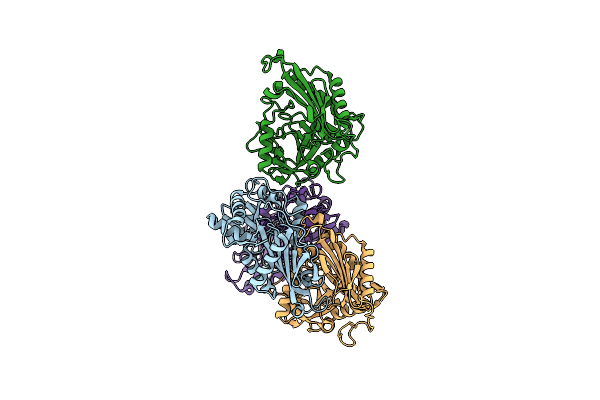 |
Organism: Mycolicibacterium vanbaalenii (strain dsm 7251 / jcm 13017 / bcrc 16820 / kctc 9966 / nrrl b-24157 / pyr-1)
Method: X-RAY DIFFRACTION Release Date: 2024-01-24 Classification: TRANSFERASE |
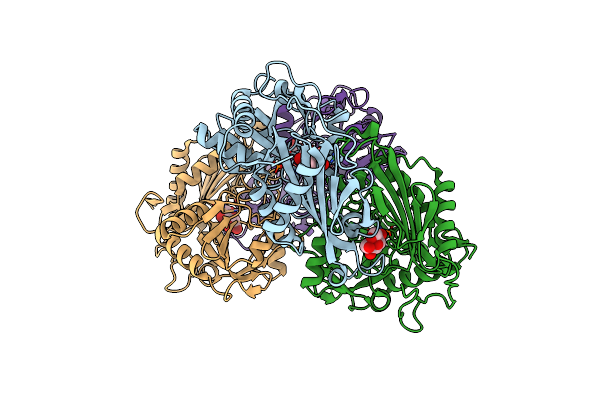 |
Crystal Structure Of Mv In Complex With Llp And Fru From Mycobacterium Vanbaalenii
Organism: Mycolicibacterium vanbaalenii (strain dsm 7251 / jcm 13017 / bcrc 16820 / kctc 9966 / nrrl b-24157 / pyr-1)
Method: X-RAY DIFFRACTION Release Date: 2024-01-24 Classification: TRANSFERASE Ligands: FUD |
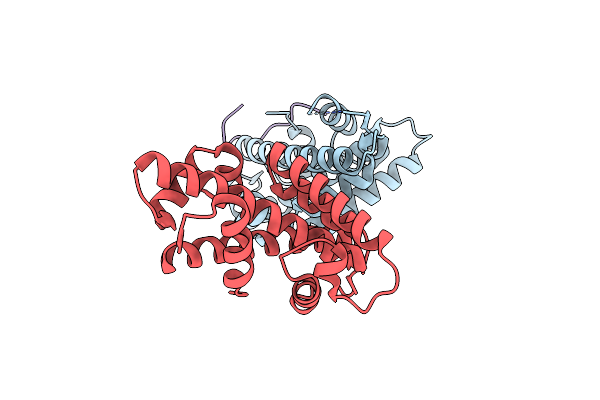 |
Crystal Structure Of Inner Membrane Protein Sad1 In Complex With Histone H2A-H2B
Organism: Schizosaccharomyces pombe
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2024-01-03 Classification: MEMBRANE PROTEIN |
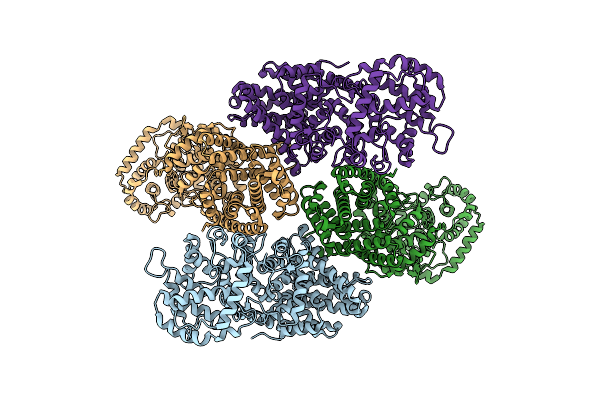 |
Organism: Oryza sativa japonica group
Method: ELECTRON MICROSCOPY Release Date: 2023-12-06 Classification: ISOMERASE |
 |
Organism: Oryza sativa japonica group
Method: ELECTRON MICROSCOPY Release Date: 2023-12-06 Classification: ISOMERASE |
 |
Organism: Oryza sativa japonica group
Method: ELECTRON MICROSCOPY Release Date: 2023-12-06 Classification: ISOMERASE |
 |
Organism: Oryza sativa japonica group
Method: ELECTRON MICROSCOPY Release Date: 2023-12-06 Classification: PLANT PROTEIN Ligands: GRG |
 |
The Crystal Structure Of Syn-Copalyl Diphosphate Synthase From Oryza Sativa
Organism: Oryza sativa japonica group
Method: X-RAY DIFFRACTION Resolution:3.49 Å Release Date: 2023-12-06 Classification: ISOMERASE |
 |
Organism: Schizosaccharomyces pombe (strain 972 / atcc 24843)
Method: X-RAY DIFFRACTION Resolution:3.38 Å Release Date: 2023-11-01 Classification: NUCLEAR PROTEIN |
 |
Organism: Desulfonema ishimotonii
Method: ELECTRON MICROSCOPY Release Date: 2023-02-01 Classification: RNA BINDING PROTEIN |
 |
Organism: Desulfonema ishimotonii
Method: ELECTRON MICROSCOPY Release Date: 2023-02-01 Classification: RNA BINDING PROTEIN |
 |
Organism: Desulfonema ishimotonii
Method: ELECTRON MICROSCOPY Release Date: 2023-02-01 Classification: RNA BINDING PROTEIN |
 |
Organism: Desulfonema ishimotonii
Method: ELECTRON MICROSCOPY Release Date: 2023-02-01 Classification: RNA BINDING PROTEIN |
 |
Organism: Desulfonema ishimotonii
Method: ELECTRON MICROSCOPY Release Date: 2023-02-01 Classification: RNA BINDING PROTEIN |
 |
Structure Of The Pore C-Terminal Domain, A Protein Of T9Ss From Porphyromonas Gingivalis
Organism: Porphyromonas gingivalis jcvi sc001
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2020-06-17 Classification: PROTEIN BINDING Ligands: MLD, NA |

