Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
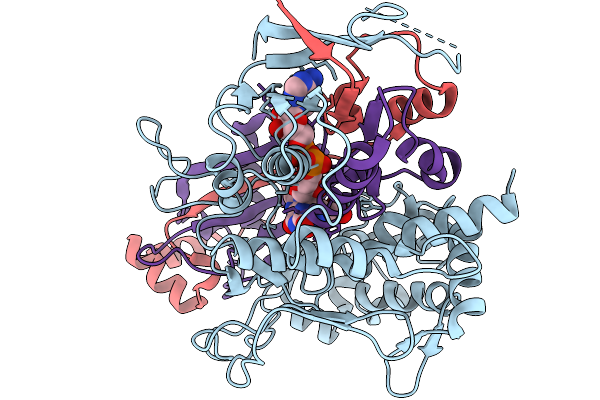 |
Organism: Streptomyces sp. x-119-6
Method: X-RAY DIFFRACTION Release Date: 2026-01-07 Classification: OXIDOREDUCTASE Ligands: FAD, GLU |
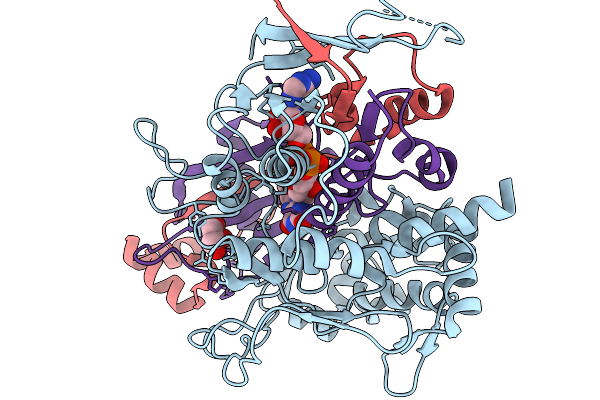 |
Organism: Streptomyces sp. x-119-6
Method: X-RAY DIFFRACTION Release Date: 2026-01-07 Classification: OXIDOREDUCTASE Ligands: FAD, GOL |
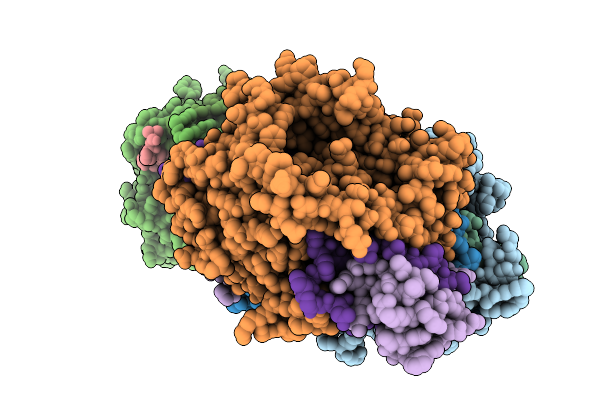 |
Organism: Streptomyces sp. x-119-6
Method: X-RAY DIFFRACTION Release Date: 2026-01-07 Classification: OXIDOREDUCTASE Ligands: FAD, GLU |
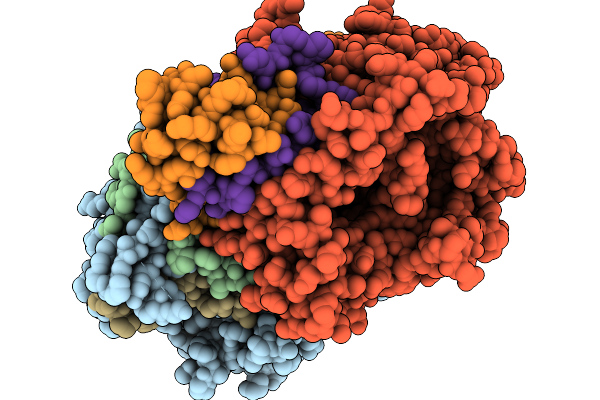 |
Organism: Streptomyces sp. x-119-6
Method: X-RAY DIFFRACTION Release Date: 2026-01-07 Classification: OXIDOREDUCTASE Ligands: FAD |
 |
Organism: Streptomyces sp. x-119-6
Method: X-RAY DIFFRACTION Release Date: 2026-01-07 Classification: OXIDOREDUCTASE Ligands: FAD |
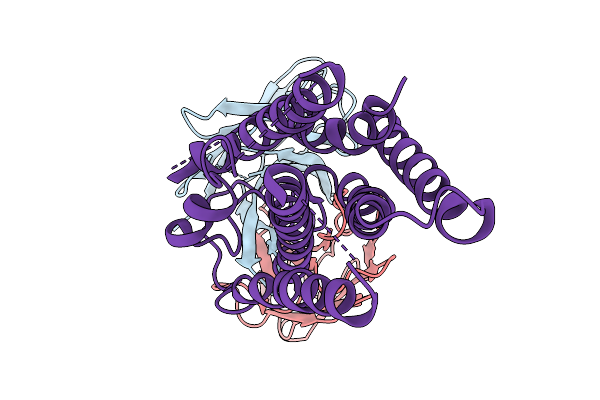 |
Cryo-Em Structure Of The Human A2A Adenosine Receptor In Complex With A Fab Antibody Fragment
Organism: Mus musculus, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-01-07 Classification: MEMBRANE PROTEIN |
 |
Organism: Streptomyces virginiae
Method: X-RAY DIFFRACTION Release Date: 2025-12-10 Classification: HYDROLASE Ligands: GOL, PEG |
 |
Organism: Streptomyces virginiae
Method: X-RAY DIFFRACTION Release Date: 2025-12-10 Classification: HYDROLASE Ligands: GOL |
 |
Organism: Streptomyces virginiae
Method: X-RAY DIFFRACTION Release Date: 2025-12-10 Classification: HYDROLASE Ligands: A1L6P, NA |
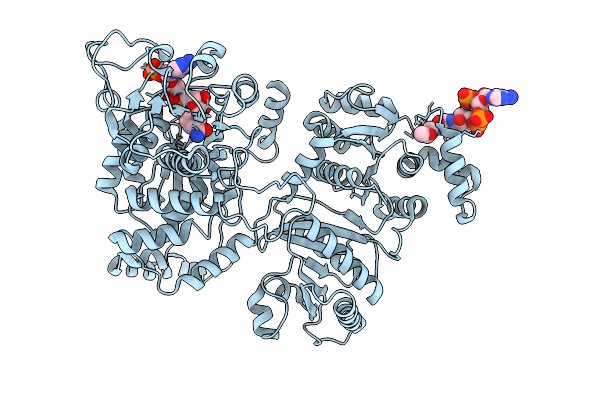 |
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2025-12-10 Classification: STRUCTURAL PROTEIN Ligands: NAP, ACO, MG |
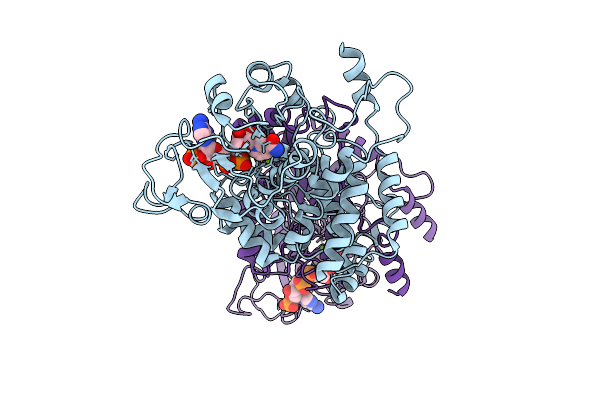 |
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2025-12-10 Classification: STRUCTURAL PROTEIN Ligands: NAP, MG |
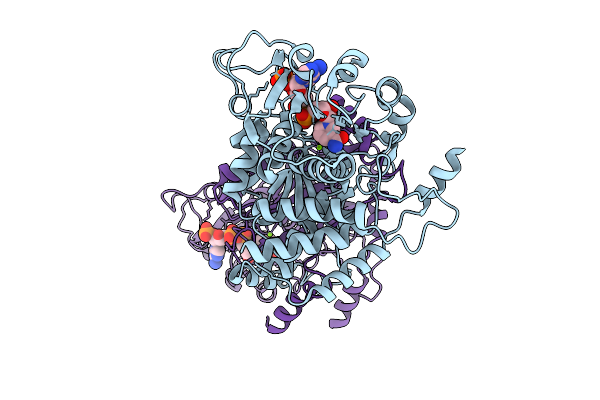 |
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2025-12-10 Classification: STRUCTURAL PROTEIN Ligands: NAP, MG |
 |
Organism: Caenorhabditis elegans
Method: X-RAY DIFFRACTION Release Date: 2025-12-10 Classification: PROTEIN BINDING |
 |
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2025-12-03 Classification: STRUCTURAL PROTEIN Ligands: NAP, MG |
 |
Organism: Bdellovibrio bacteriovorus
Method: ELECTRON MICROSCOPY Release Date: 2025-12-03 Classification: STRUCTURAL PROTEIN Ligands: NAP |
 |
Organism: Bdellovibrio bacteriovorus
Method: ELECTRON MICROSCOPY Release Date: 2025-12-03 Classification: STRUCTURAL PROTEIN Ligands: ACO, NAP |
 |
Organism: Escherichia coli k-12
Method: X-RAY DIFFRACTION Release Date: 2025-12-03 Classification: STRUCTURAL PROTEIN Ligands: MG |
 |
Organism: Escherichia coli k-12
Method: X-RAY DIFFRACTION Release Date: 2025-12-03 Classification: STRUCTURAL PROTEIN Ligands: SO4 |
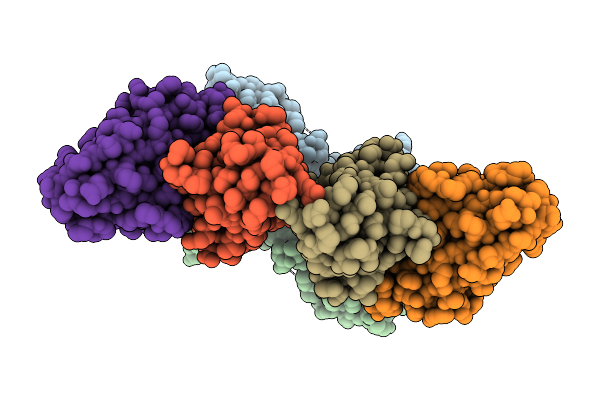 |
Cryo-Em Structure Of Human Complement C1S Cub Domain In Complex With Ray121
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-11-19 Classification: IMMUNE SYSTEM Ligands: CA |
 |
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2025-11-12 Classification: RNA BINDING PROTEIN Ligands: ADP, ZN, MG |