Search Count: 18
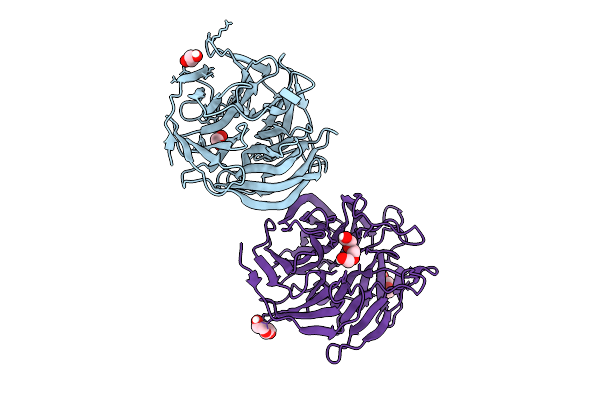 |
Crystal Structure Of Complement1.4 (Cent1.4), An Engineered Photoenzyme For Selective [2+2]-Cycloadditions
Organism: Loligo vulgaris
Method: X-RAY DIFFRACTION Release Date: 2025-07-16 Classification: BIOSYNTHETIC PROTEIN Ligands: EDO, PEG, PGE |
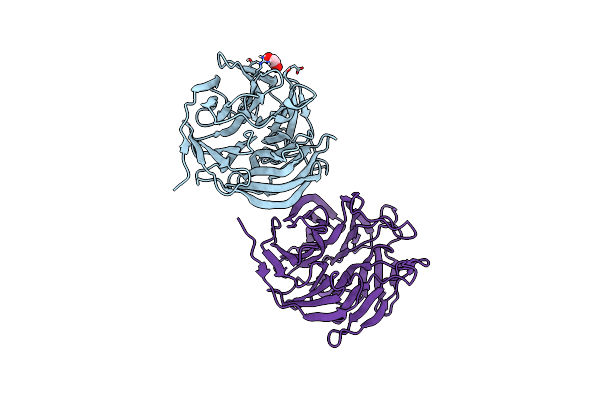 |
Organism: Loligo vulgaris
Method: X-RAY DIFFRACTION Release Date: 2025-05-14 Classification: BIOSYNTHETIC PROTEIN Ligands: EDO |
 |
Organism: Loligo vulgaris
Method: X-RAY DIFFRACTION Release Date: 2025-05-14 Classification: BIOSYNTHETIC PROTEIN Ligands: PEG, EDO |
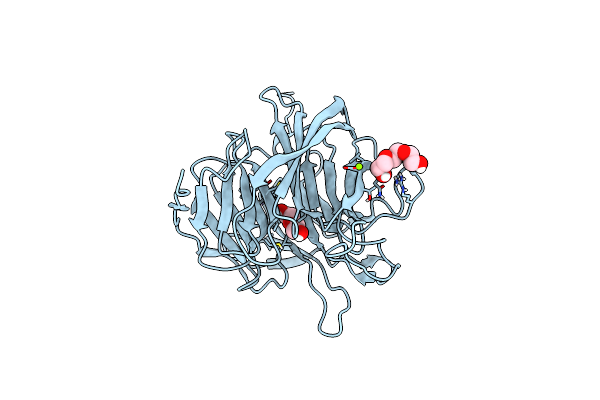 |
Organism: Loligo vulgaris
Method: X-RAY DIFFRACTION Release Date: 2025-05-14 Classification: BIOSYNTHETIC PROTEIN Ligands: PEG, PGE, MG |
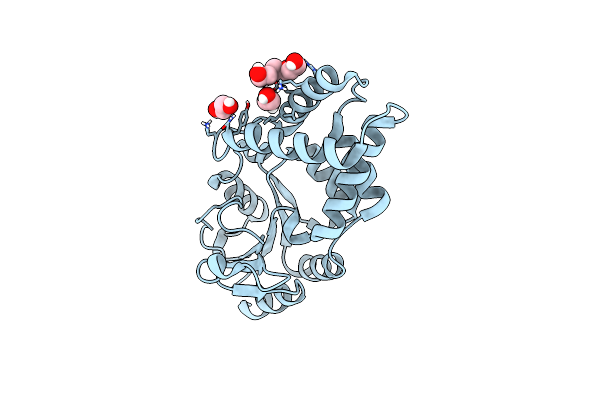 |
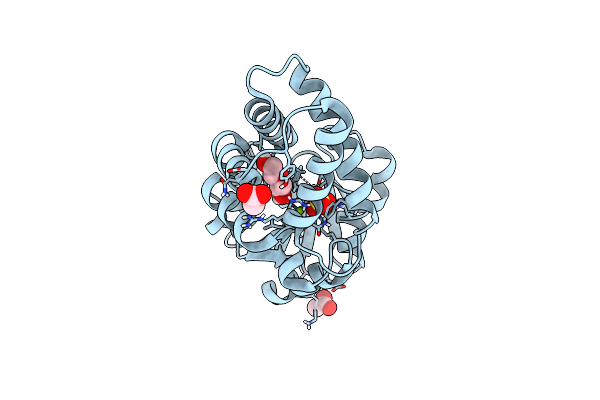
Crystal Structure Of Bhmehis1.8, An Engineered Enzyme For The Morita-Baylis-Hillman Reaction
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.62 Å Release Date: 2024-02-28 Classification: BIOSYNTHETIC PROTEIN Ligands: PGE, EDO |
 |
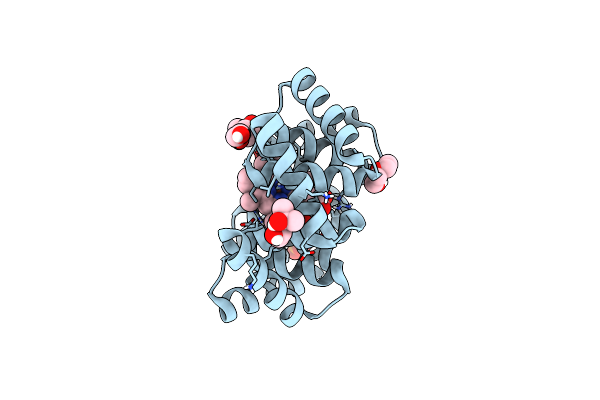
Crystal Structure Of Bhmehis1.0, An Engineered Enzyme For The Morita-Baylis-Hillman Reaction
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.72 Å Release Date: 2024-02-28 Classification: BIOSYNTHETIC PROTEIN Ligands: PEG, ACT, MG, SO4 |
 |
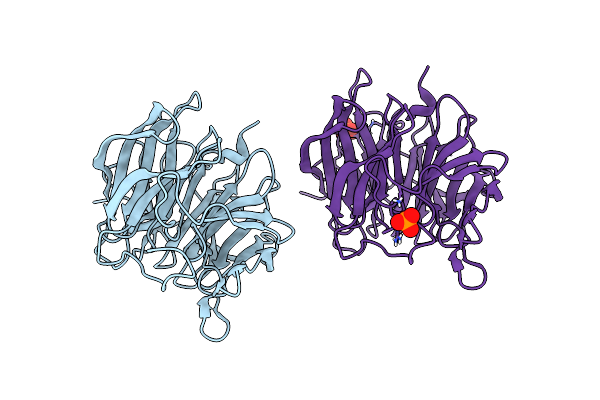
Crystal Structure Of A Computationally Designed Heme Binding Protein, Dnhem1
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2023-07-05 Classification: BIOSYNTHETIC PROTEIN Ligands: IMD, HEM, MPD, PO4, PEG, EDO |
 |
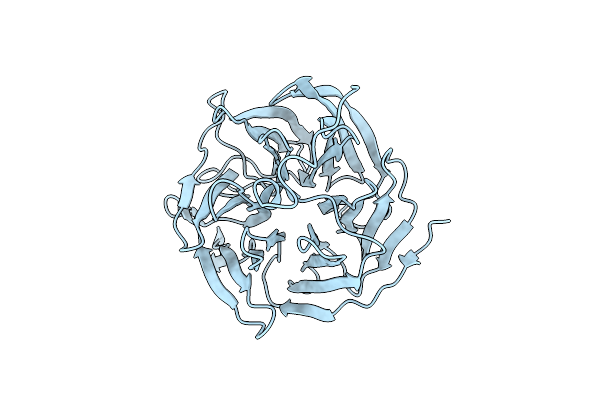
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.54 Å Release Date: 2022-09-28 Classification: BIOSYNTHETIC PROTEIN Ligands: PO4 |
 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2022-09-28 Classification: BIOSYNTHETIC PROTEIN |
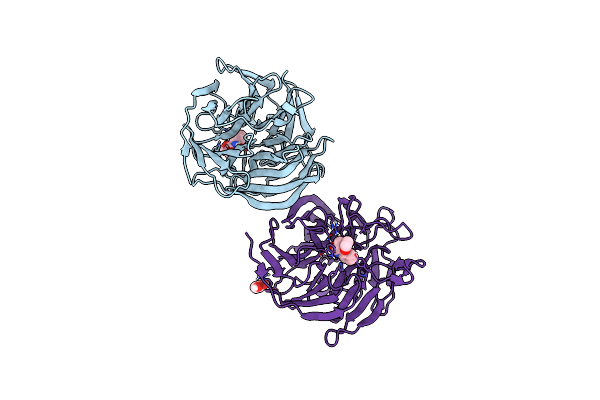 |
Crystal Structure Of Evolved Photoenzyme Ent1.3 (Truncated) With Bound Product
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2022-09-28 Classification: BIOSYNTHETIC PROTEIN Ligands: JRP, EDO |
 |
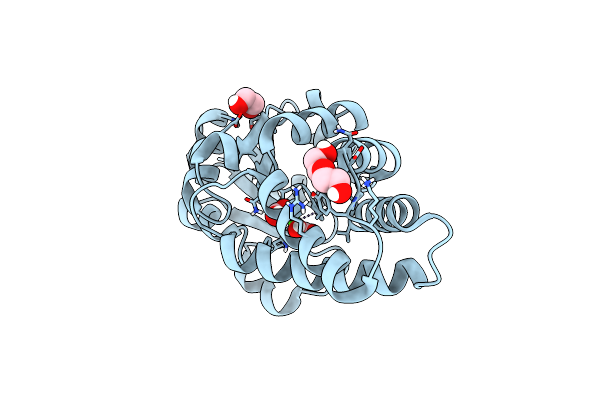
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2021-11-03 Classification: BIOSYNTHETIC PROTEIN |
 |
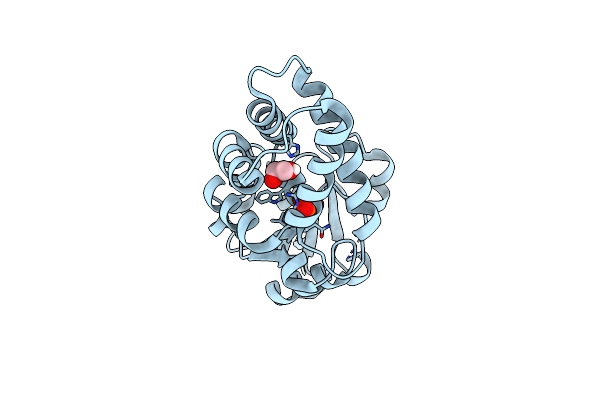
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.48 Å Release Date: 2021-08-25 Classification: BIOSYNTHETIC PROTEIN Ligands: PEG, EDO, CA, FMT |
 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.29 Å Release Date: 2021-08-25 Classification: BIOSYNTHETIC PROTEIN Ligands: PO4, EDO |
 |
Crystal Structure Of Bh32 Alkylated With The Mechanistic Inhibitor 2-Bromoacetophenone
Organism: Pyrococcus horikoshii
Method: X-RAY DIFFRACTION Resolution:2.02 Å Release Date: 2019-06-05 Classification: HYDROLASE Ligands: AC0 |
 |
Organism: Pyrococcus horikoshii
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2019-06-05 Classification: HYDROLASE Ligands: CA |
 |
Organism: Pyrococcus horikoshii
Method: X-RAY DIFFRACTION Resolution:1.96 Å Release Date: 2019-06-05 Classification: HYDROLASE Ligands: PGE, AC0, SO4, EDO, MG |
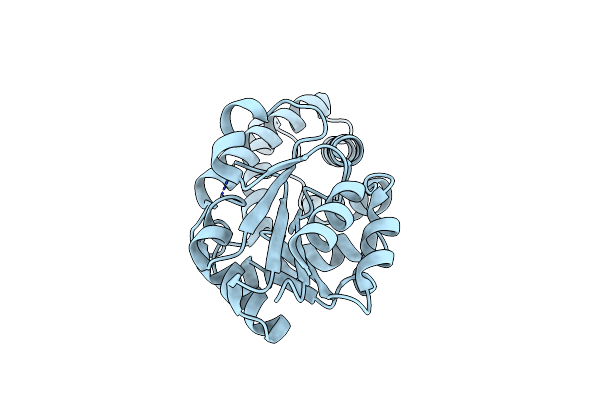 |
Organism: Pyrococcus horikoshii
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2019-06-05 Classification: HYDROLASE |
 |
Crystal Structure Of Oe1.3 Alkylated With The Mechanistic Inhibitor 2-Bromoacetophenone
Organism: Pyrococcus horikoshii
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2019-06-05 Classification: HYDROLASE Ligands: AC0, ACT, PEG |

