Search Count: 35
All
Selected
 |
Organism: Escherichia coli k-12, Enterobacteria phage m
Method: ELECTRON MICROSCOPY Resolution:3.60 Å Release Date: 2026-02-11 Classification: TRANSPORT PROTEIN |
 |
Organism: Escherichia coli k-12, Pseudomonas phage pp7
Method: ELECTRON MICROSCOPY Resolution:3.70 Å Release Date: 2026-02-11 Classification: TRANSPORT PROTEIN |
 |
Structure Of Murj In Complex With Single Gene Lysis Protein From Phage Changjiang3
Organism: Escherichia coli k-12, Changjiang levi-like virus 3
Method: ELECTRON MICROSCOPY Resolution:3.60 Å Release Date: 2026-02-11 Classification: TRANSPORT PROTEIN |
 |
Cryo-Em Structure Of Human Udp-N-Acetylglucosamine-Dolichyl-Phosphate N-Acetylglucosaminephosphotransferase (Dpagt1) In Complex With Appb
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.90 Å Release Date: 2026-02-04 Classification: TRANSFERASE Ligands: A1C3G, A1C3H |
 |
Organism: Hydrogenivirga sp. 128-5-r1-1
Method: ELECTRON MICROSCOPY Resolution:2.91 Å Release Date: 2026-02-04 Classification: TRANSFERASE Ligands: A1C3G |
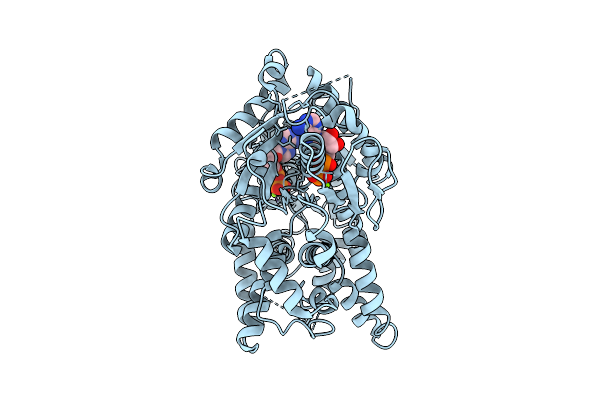 |
Open Conformation Of Arsa From L. Ferriphilum In Complex With Mgadp Determined In The Presence Of Arsenite
Organism: Leptospirillum ferriphilum
Method: ELECTRON MICROSCOPY Release Date: 2025-08-20 Classification: HYDROLASE Ligands: ADP, MG |
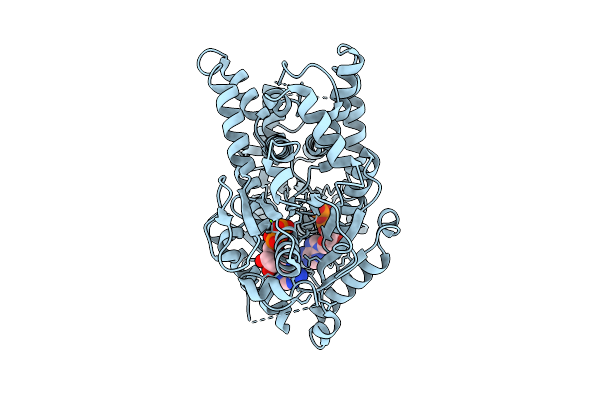 |
Open Conformation Of Arsa From L. Ferriphilum In Complex With Mgadp Determined In Absence Of Arsenite
Organism: Leptospirillum ferriphilum
Method: ELECTRON MICROSCOPY Release Date: 2025-08-20 Classification: HYDROLASE Ligands: ADP, MG |
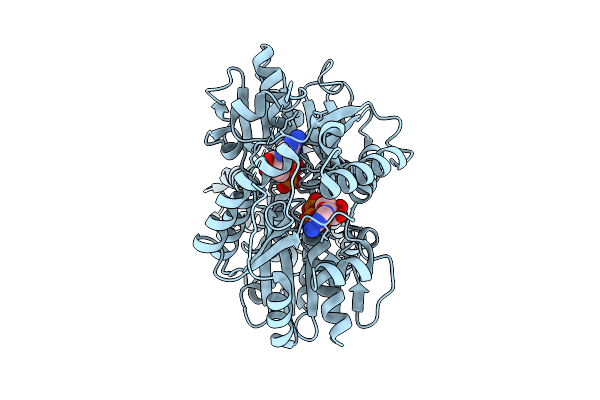 |
Closed Conformation Of Arsa From L. Ferriphilum In Complex With Mgatp And Arsenite
Organism: Leptospirillum ferriphilum
Method: ELECTRON MICROSCOPY Release Date: 2025-08-20 Classification: HYDROLASE Ligands: ATP, MG, ARS |
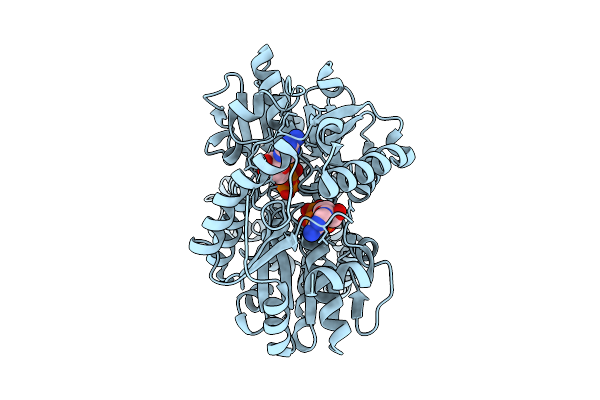 |
Closed Conformation Of Arsa From L. Ferriphilum In Complex With Mgatp And Arsenite At 1.5 Minute Time Point
Organism: Leptospirillum ferriphilum ml-04
Method: ELECTRON MICROSCOPY Release Date: 2025-08-20 Classification: HYDROLASE Ligands: ATP, MG, ARS |
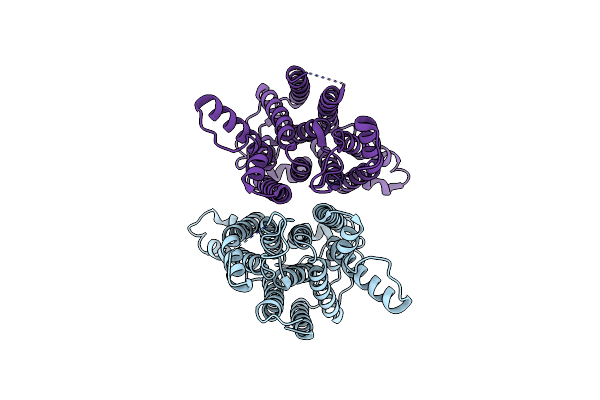 |
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2024-05-01 Classification: MEMBRANE PROTEIN |
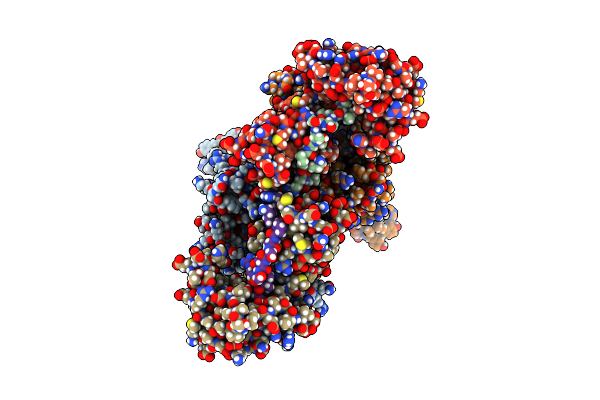 |
Organism: Escherichia coli k-12, Escherichia phage id21
Method: ELECTRON MICROSCOPY Release Date: 2023-07-26 Classification: TRANSFERASE/ISOMERASE |
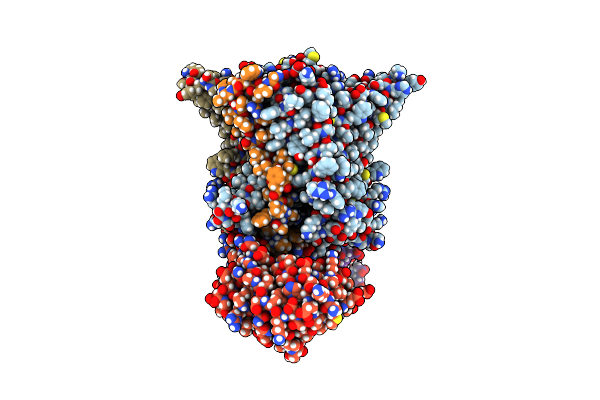 |
Organism: Escherichia coli k-12, Escherichia phage phix174
Method: ELECTRON MICROSCOPY Release Date: 2023-07-26 Classification: TRANSFERASE/ISOMERASE |
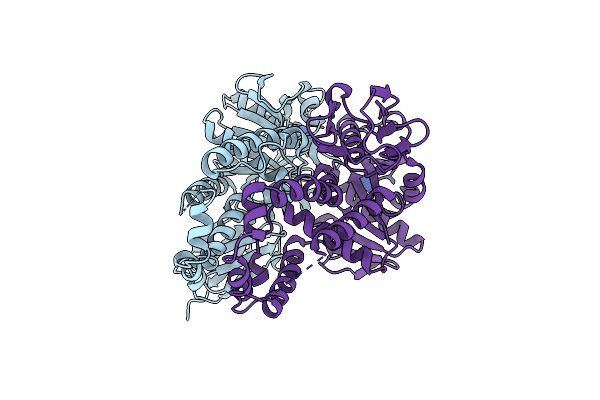 |
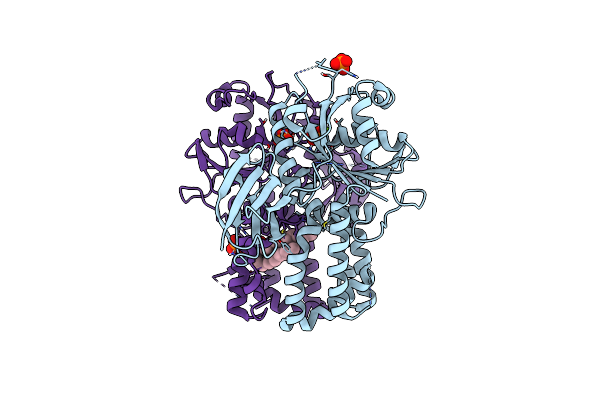
Re-Refinement Of Crystal Structure Of Nosget3D, The All4481 Protein From Nostoc Sp. Pcc 7120
Organism: Nostoc sp. pcc 7120 = fachb-418
Method: X-RAY DIFFRACTION Resolution:1.98 Å Release Date: 2023-05-24 Classification: CHAPERONE Ligands: AZI |
 |
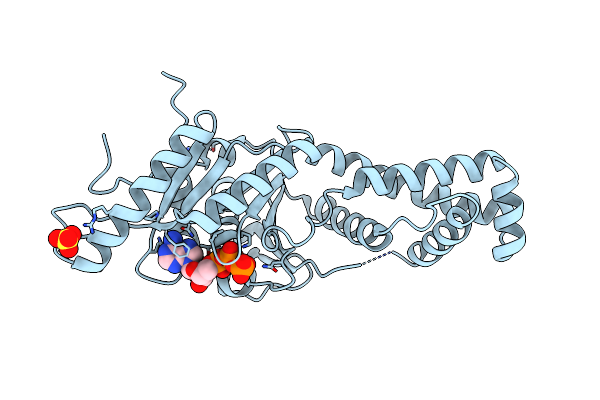
Organism: Arabidopsis thaliana
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2023-05-03 Classification: CHAPERONE Ligands: PO4, MG, XP4 |
 |
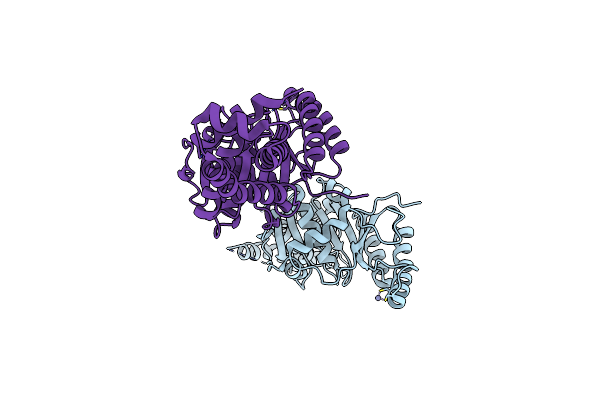
Organism: Giardia intestinalis (strain atcc 50803 / wb clone c6)
Method: X-RAY DIFFRACTION Resolution:2.23 Å Release Date: 2022-07-20 Classification: CHAPERONE Ligands: ATP, MG, ZN, SO4 |
 |
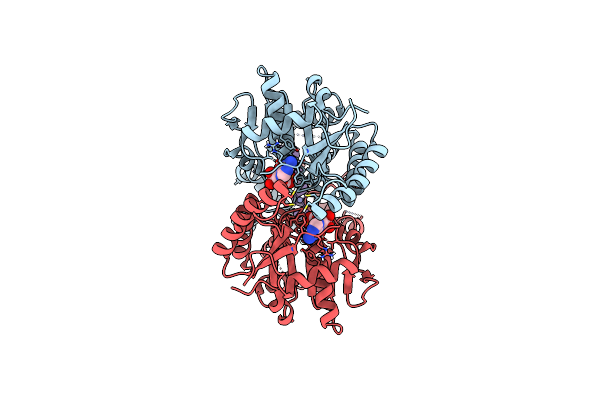
Organism: Giardia lamblia atcc 50803
Method: X-RAY DIFFRACTION Resolution:3.00 Å Release Date: 2022-07-20 Classification: CHAPERONE Ligands: ZN |
 |
Get3 Bound To Adp And The Transmembrane Domain Of The Tail-Anchored Protein Bos1
Organism: Giardia intestinalis (strain atcc 50803 / wb clone c6), Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2022-07-20 Classification: CHAPERONE Ligands: ADP, ZN, MG |
 |
Crystal Structure Of Seca:Adp In An Open Conformation From Bacillus Subtilis
Organism: Bacillus subtilis
Method: X-RAY DIFFRACTION Resolution:2.90 Å Release Date: 2004-08-03 Classification: PROTEIN TRANSPORT Ligands: MG, ADP |
 |
Organism: Bacillus subtilis
Method: X-RAY DIFFRACTION Resolution:2.18 Å Release Date: 2004-08-03 Classification: PROTEIN TRANSPORT |
 |
Crystal Structure Of The Bacterial Fatty Acid Transporter Fadl From Escherichia Coli
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2004-06-15 Classification: LIPID TRANSPORT Ligands: CU, LDA, C8E |

