Search Count: 4,125
All
Selected
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.85 Å Release Date: 2026-03-04 Classification: CELL CYCLE Ligands: A1B7H, CIT |
 |
Cryo-Em Structure Of The Human Hec1-Nuf2 Dimer Bound To The Paclitaxel-Stabilized Microtubule
Organism: Homo sapiens, Sus scrofa
Method: ELECTRON MICROSCOPY Release Date: 2026-02-04 Classification: CELL CYCLE Ligands: GTP, MG, GDP, TA1 |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-02-04 Classification: CELL CYCLE Ligands: A1B7G, ZN |
 |
Organism: Arabidopsis thaliana
Method: X-RAY DIFFRACTION Resolution:1.88 Å Release Date: 2026-02-04 Classification: PLANT PROTEIN Ligands: MLA, GOL |
 |
Crystal Structure Of Cdc48A-N Domain In Complex With Pux5 Shp Box Motif And Ubx Domain
Organism: Arabidopsis thaliana
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2026-02-04 Classification: PLANT PROTEIN |
 |
Organism: Sulfolobus acidocaldarius
Method: ELECTRON MICROSCOPY Release Date: 2026-01-28 Classification: CELL CYCLE |
 |
Organism: Sulfolobus acidocaldarius
Method: ELECTRON MICROSCOPY Release Date: 2026-01-28 Classification: CELL CYCLE |
 |
Organism: Sulfolobus acidocaldarius
Method: ELECTRON MICROSCOPY Release Date: 2026-01-28 Classification: CELL CYCLE |
 |
Organism: Saccharolobus islandicus
Method: X-RAY DIFFRACTION Resolution:2.85 Å Release Date: 2026-01-28 Classification: CELL CYCLE |
 |
Organism: Saccharolobus islandicus
Method: ELECTRON MICROSCOPY Release Date: 2026-01-28 Classification: CELL CYCLE |
 |
Organism: Saccharolobus islandicus
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2026-01-28 Classification: CELL CYCLE |
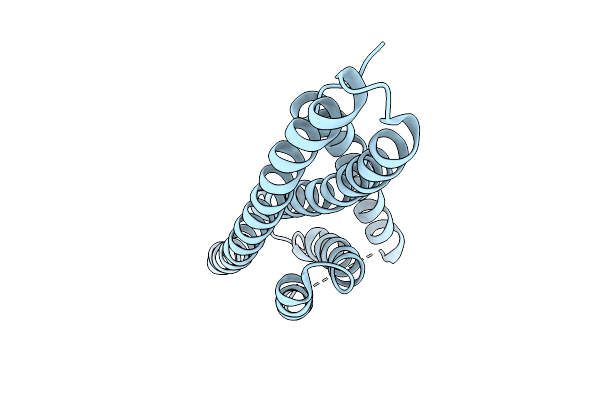 |
Organism: Saccharolobus islandicus
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2026-01-28 Classification: CELL CYCLE |
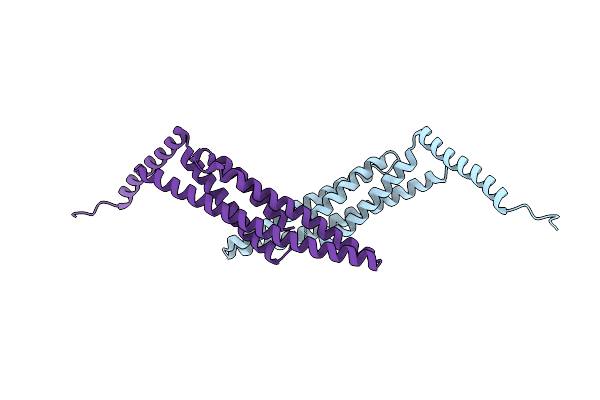 |
Organism: Saccharolobus islandicus
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2026-01-28 Classification: CELL CYCLE |
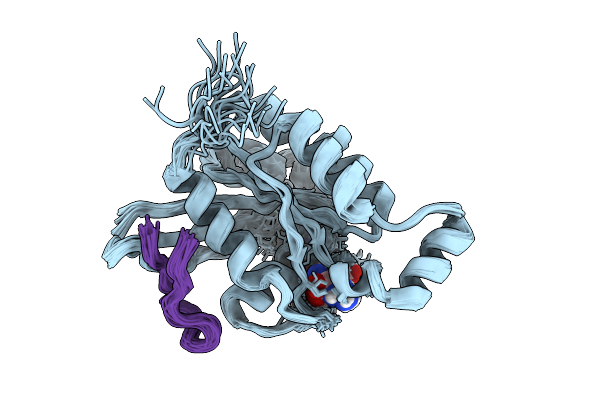 |
Organism: Homo sapiens, Synthetic construct
Method: SOLUTION NMR Release Date: 2026-01-21 Classification: CELL CYCLE Ligands: GNP, MG |
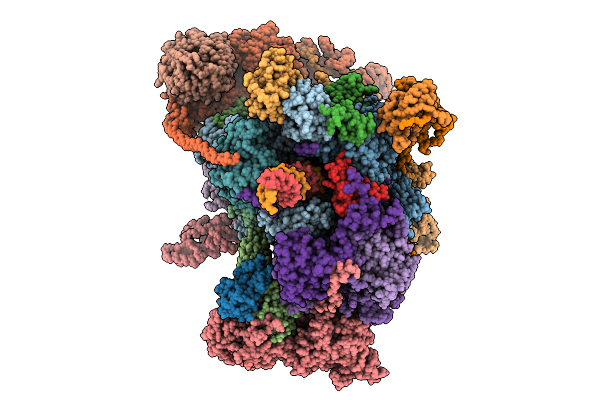 |
Pol Ii-Dsif-Spt6-Paf1C-Tfiis-Iws1-Elof1-Ledgf-Nucleosome Activated Elongation Complex Composite Map T
Organism: Homo sapiens, Xenopus laevis, Synthetic construct, Sus scrofa
Method: ELECTRON MICROSCOPY Release Date: 2026-01-21 Classification: TRANSCRIPTION Ligands: ZN, MG |
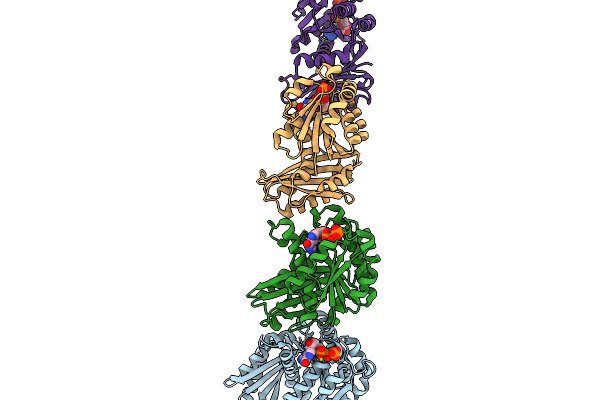 |
Organism: Streptococcus mutans serotype c (strain atcc 700610 / ua159)
Method: X-RAY DIFFRACTION Resolution:3.49 Å Release Date: 2026-01-14 Classification: CELL CYCLE Ligands: GTP, MG |
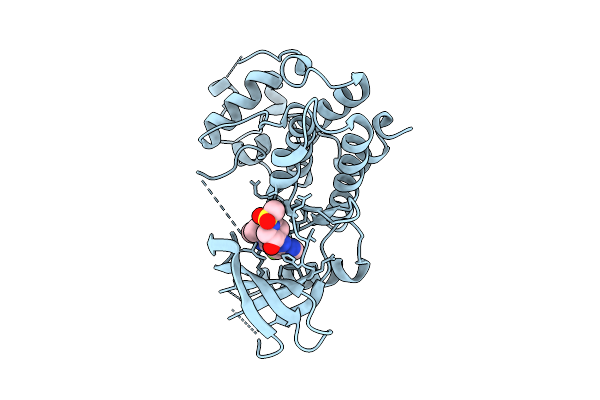 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2025-12-17 Classification: TRANSFERASE/INHIBITOR Ligands: A1CIF |
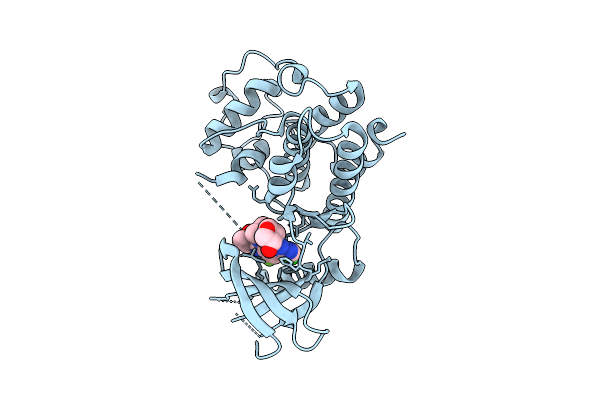 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2025-12-17 Classification: TRANSFERASE/INHIBITOR Ligands: A1AZ4 |
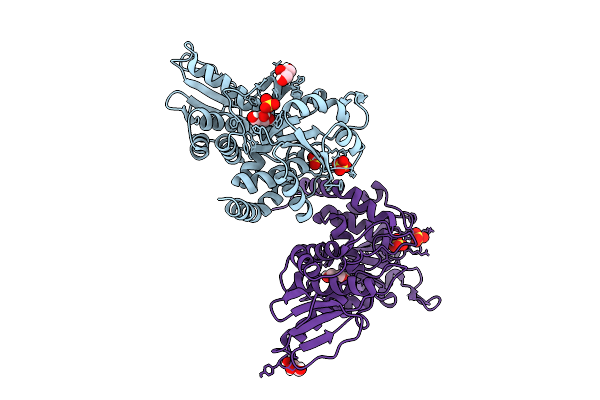 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:1.62 Å Release Date: 2025-12-10 Classification: CELL CYCLE Ligands: GOL, SO4 |
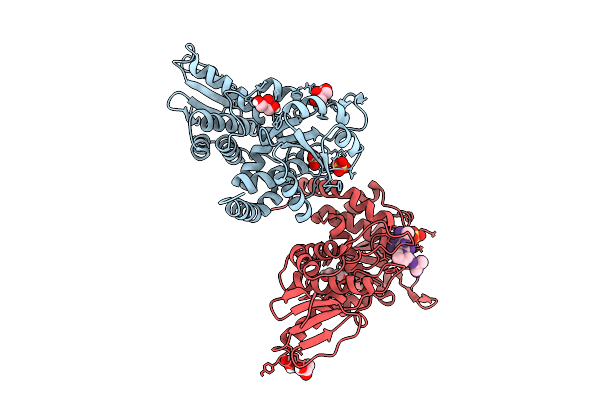 |
Crystal Structure Of C278S Mutant Of Mouse Cdc14A In Complex With A Model Phosphopeptide
Organism: Mus musculus, Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.62 Å Release Date: 2025-12-10 Classification: CELL CYCLE Ligands: GOL, SO4 |

