Search Count: 56
 |
Structure Of Proline Utilization A Complexed With 2,3-Dihydro-1,4-Benzodioxine-5-Carboxylic Acid
Organism: Sinorhizobium meliloti
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2024-11-27 Classification: OXIDOREDUCTASE Ligands: FAD, PEG, 53E, FMT, NAD, SO4, MG, PGE, PG4 |
 |
Structure Of Proline Utilization A Complexed With Quinoline-2-Carboxylic Acid
Organism: Sinorhizobium meliloti
Method: X-RAY DIFFRACTION Resolution:1.77 Å Release Date: 2024-11-27 Classification: OXIDOREDUCTASE Ligands: FAD, PGE, QNC, PEG, FMT, SO4, MG, PG4 |
 |
Organism: Sinorhizobium meliloti
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2024-11-27 Classification: OXIDOREDUCTASE Ligands: PGE, PEG, PYS, FMT, SO4, FAD |
 |
Organism: Sinorhizobium meliloti
Method: X-RAY DIFFRACTION Resolution:1.77 Å Release Date: 2024-11-27 Classification: OXIDOREDUCTASE Ligands: FAD, PGE, FMT, PEG, PG4, A1BDA, SO4 |
 |
Structure Of Proline Utilization A Complexed With Piperidine-3-Carboxylic Acid
Organism: Sinorhizobium meliloti
Method: X-RAY DIFFRACTION Resolution:1.42 Å Release Date: 2024-11-27 Classification: OXIDOREDUCTASE Ligands: FAD, PEG, PGE, FMT, A1BDB, ID7, SO4, NAD, MG, PG4 |
 |
Structure Of Proline Utilization A Complexed With 1-(4-Fluorophenyl)Thiourea
Organism: Sinorhizobium meliloti
Method: X-RAY DIFFRACTION Resolution:1.72 Å Release Date: 2024-11-27 Classification: OXIDOREDUCTASE Ligands: FAD, PEG, PGE, 8P4, FMT, SO4 |
 |
Organism: Sinorhizobium meliloti
Method: X-RAY DIFFRACTION Resolution:1.64 Å Release Date: 2024-11-27 Classification: OXIDOREDUCTASE Ligands: FAD, PEG, A1BDD, PGE, PG4, FMT, NAD, SO4, MG |
 |
Structure Of Proline Utilization A Co-Crystallized With 4-Methoxybenzyl Alcohol
Organism: Sinorhizobium meliloti
Method: X-RAY DIFFRACTION Resolution:1.32 Å Release Date: 2024-11-27 Classification: OXIDOREDUCTASE Ligands: FAD, A1BDD, PGE, FMT, PEG, NAD, SO4, MG, 1PE |
 |
Organism: Sinorhizobium meliloti
Method: X-RAY DIFFRACTION Resolution:1.39 Å Release Date: 2024-11-27 Classification: OXIDOREDUCTASE Ligands: FAD, FMT, PEG, NAD, SO4, MG, PGE, A1BDW |
 |
Structure Of Proline Utilization A Complexed With (2,3-Dihydro-1-Benzofuran-5-Yl)Methanol
Organism: Sinorhizobium meliloti
Method: X-RAY DIFFRACTION Resolution:1.47 Å Release Date: 2024-11-27 Classification: OXIDOREDUCTASE Ligands: FAD, PEG, FMT, A1AQC, NAD, SO4, MG, 1PE, PGE |
 |
Structure Of Proline Utilization A Complexed With 1-Benzofuran-5-Ylmethanol
Organism: Sinorhizobium meliloti
Method: X-RAY DIFFRACTION Resolution:1.32 Å Release Date: 2024-11-27 Classification: OXIDOREDUCTASE Ligands: FAD, PEG, FMT, A1BDX, NAD, SO4, MG, PGE, 1PE |
 |
Organism: Sinorhizobium meliloti
Method: X-RAY DIFFRACTION Resolution:1.33 Å Release Date: 2024-11-27 Classification: OXIDOREDUCTASE Ligands: FAD, PGE, FMT, A1BDV, NAD, SO4, MG, 1PE, PEG |
 |
Organism: Sinorhizobium meliloti
Method: X-RAY DIFFRACTION Resolution:1.37 Å Release Date: 2024-11-27 Classification: OXIDOREDUCTASE Ligands: FAD, PGE, PEG, FMT, A1BDU, NAD, SO4, MG, 1PE |
 |
Minimal Puta Proline Dehydrogenase Domain (Design #2) Complexed With (Prop-2-Ynylthio)Acetic Acid
Organism: Sinorhizobium meliloti sm11
Method: X-RAY DIFFRACTION Resolution:1.38 Å Release Date: 2024-10-30 Classification: OXIDOREDUCTASE Ligands: FAD, PEG, X79 |
 |
Minimal Puta Proline Dehydrogenase Domain (Design #2) Complexed With 2-(Cyanomethylthio)Acetic Acid
Organism: Sinorhizobium meliloti sm11
Method: X-RAY DIFFRACTION Resolution:1.43 Å Release Date: 2024-10-30 Classification: OXIDOREDUCTASE Ligands: FAD, X7K, PEG |
 |
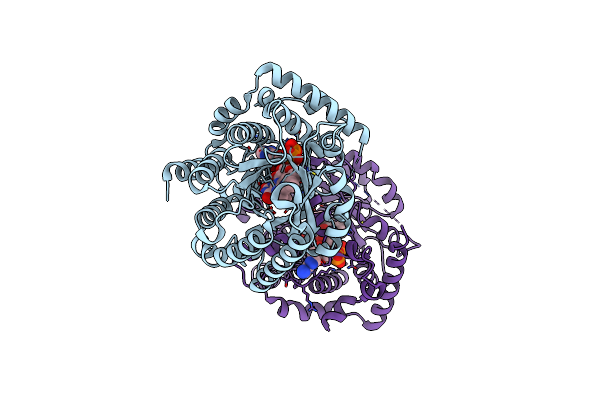
Minimal Puta Proline Dehydrogenase Domain (Design #2) Complexed With (Allylthio)Acetic Acid
Organism: Sinorhizobium meliloti sm11
Method: X-RAY DIFFRACTION Resolution:1.41 Å Release Date: 2024-10-30 Classification: OXIDOREDUCTASE Ligands: FAD, PEG, X7Q |
 |
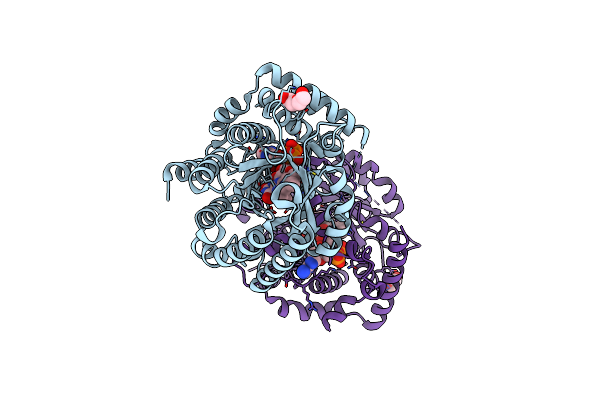
Minimal Puta Proline Dehydrogenase Domain (Design #2) With The Fad N5 Modified With Propanal Resulting From Inactivation With N-Allylglycine(Replicate #1)
Organism: Sinorhizobium meliloti
Method: X-RAY DIFFRACTION Resolution:1.54 Å Release Date: 2024-10-30 Classification: OXIDOREDUCTASE Ligands: FDA, FAD, CBG |
 |
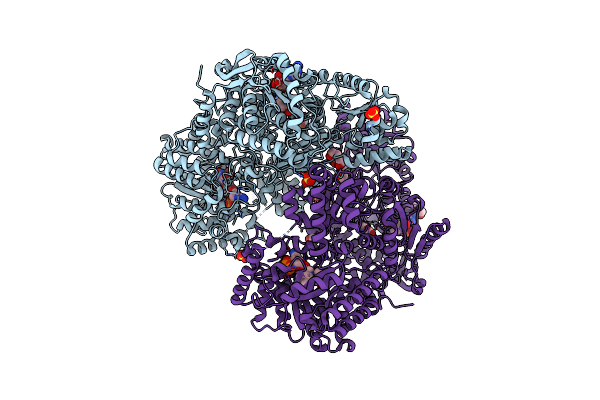
Minimal Puta Proline Dehydrogenase Domain (Design #2) With The Fad N5 Modified With Propanal Resulting From Inactivation With N-Allylglycine (Replicate #2)
Organism: Sinorhizobium meliloti
Method: X-RAY DIFFRACTION Resolution:1.52 Å Release Date: 2024-10-30 Classification: OXIDOREDUCTASE Ligands: FDA, FAD, CBG, PEG |
 |
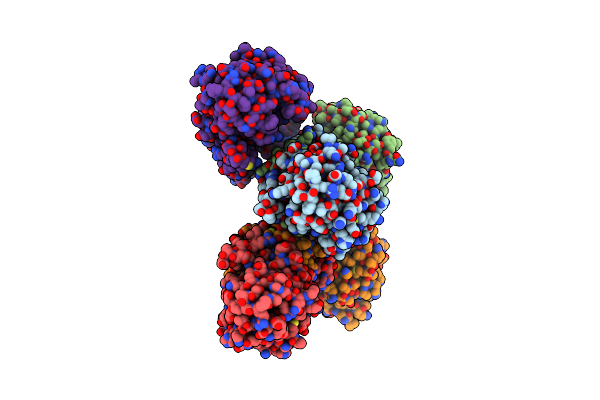
Structure Of Proline Utilization A With The Fad Covalently-Modified By Propanal Resulting From Inactivation With N-Allylglycine
Organism: Sinorhizobium meliloti
Method: X-RAY DIFFRACTION Resolution:1.54 Å Release Date: 2024-10-30 Classification: OXIDOREDUCTASE Ligands: FDA, CBG, NAD, MG, SO4, PEG, A1AV8, PGE |
 |
Structure Of Pycr1 Complexed With Nadh And Cyclobutane-1,1-Dicarboxylic Acid
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2024-07-03 Classification: OXIDOREDUCTASE/INHIBITOR Ligands: NAI, ZPS, SO4 |

