Search Count: 7
 |
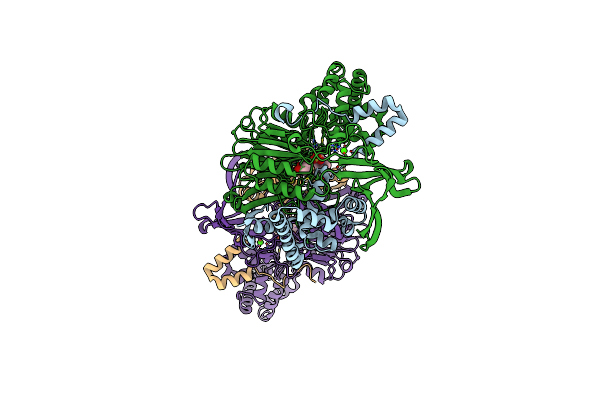
Crystal Structure Of A Surface Entropy Reduction Variant Of Penicillin G Acylase From Bacillaceae I. S. Sp. Fjat-27231
Organism: Bacillus sp. fjat-27231
Method: X-RAY DIFFRACTION Resolution:1.63 Å Release Date: 2023-05-03 Classification: HYDROLASE Ligands: EPE, BU3, CA |
 |
Crystal Structure Of A Variant Of Penicillin G Acylase From Bacillaceae I. S. Sp. Fjat-27231 With Reduced Surface Entropy And Additionally Engineered Crystal Contact
Organism: Bacillus sp. fjat-27231
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2023-05-03 Classification: HYDROLASE Ligands: EPE, BU3, CA |
 |
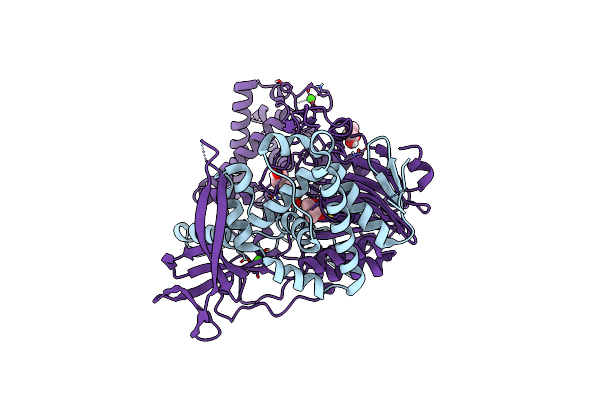
Crystal Structure Of A Variant Of Penicillin G Acylase From Bacillaceae I. S. Sp. Fjat-27231 With Reduced Surface Entropy And Additionally Engineered Crystal Contact.
Organism: Bacillus sp. fjat-27231
Method: X-RAY DIFFRACTION Resolution:1.20 Å Release Date: 2023-05-03 Classification: HYDROLASE Ligands: EPE, BU3, CA |
 |
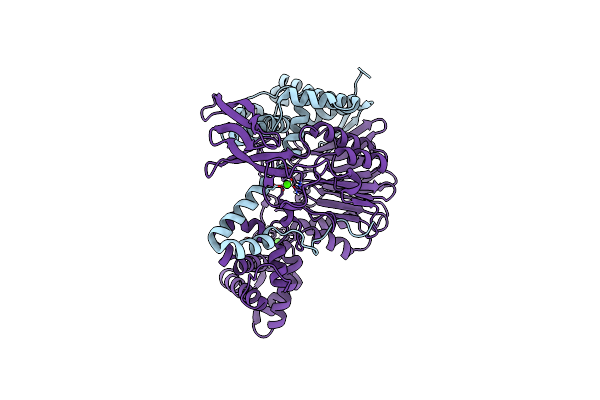
Crystal Structure Of A Variant Of Penicillin G Acylase From Bacillaceae I. S. Sp. Fjat-27231 With Reduced Surface Entropy And Additionally Engineered Crystal Contact
Organism: Bacillus sp. fjat-27231
Method: X-RAY DIFFRACTION Resolution:1.31 Å Release Date: 2023-05-03 Classification: HYDROLASE Ligands: EPE, BU3, CA |
 |
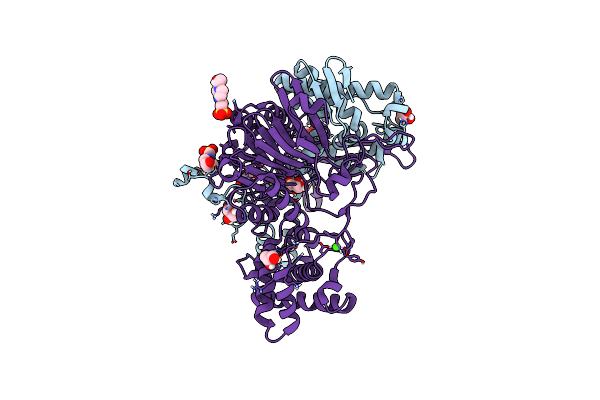
Organism: Bacillus megaterium
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2019-07-03 Classification: HYDROLASE Ligands: CA |
 |
Organism: Bacillus sp. fjat-27231
Method: X-RAY DIFFRACTION Resolution:1.36 Å Release Date: 2019-07-03 Classification: HYDROLASE Ligands: EPE, GOL, CA |
 |
Organism: Quasibacillus thermotolerans
Method: X-RAY DIFFRACTION Resolution:2.27 Å Release Date: 2019-07-03 Classification: HYDROLASE Ligands: CA, GOL |

