Search Count: 84
 |
Proline Utilization A With The Fadh- N5 Atom Covalently Modified By Proline
Organism: Sinorhizobium meliloti sm11
Method: X-RAY DIFFRACTION Release Date: 2025-06-04 Classification: OXIDOREDUCTASE Ligands: NAD, PGE, FDA, A1CAB, PRO, MG, SO4, A1AT4 |
 |
Proline Utilization A With The Covalent Acyl-Enzyme Intermediate In The Aldehyde Dehydrogenase Active Site
Organism: Sinorhizobium meliloti sm11
Method: X-RAY DIFFRACTION Release Date: 2025-06-04 Classification: OXIDOREDUCTASE Ligands: NAD, SO4, PGE, A1AUG, MG, FDA, PEG |
 |
Proline Utilization A Complexed With The Substrate L-Glutamate Gamma-Semialdehyde In The Aldehyde Dehydrogenase Active Site
Organism: Sinorhizobium meliloti sm11
Method: X-RAY DIFFRACTION Release Date: 2025-06-04 Classification: OXIDOREDUCTASE Ligands: NAD, FDA, PGE, PEG, A1AUG, FMT, MG, SO4 |
 |
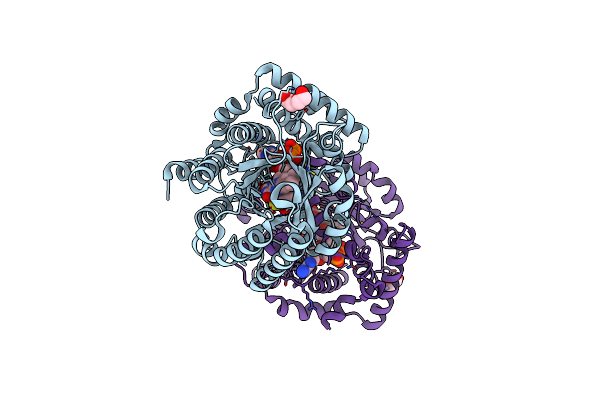
Proline Utilization A Complexed With The Product L-Glutamate In The Aldehyde Dehydrogenase Active Site
Organism: Sinorhizobium meliloti
Method: X-RAY DIFFRACTION Release Date: 2025-03-12 Classification: OXIDOREDUCTASE Ligands: GGL, SO4, PGE, FAD, PEG |
 |
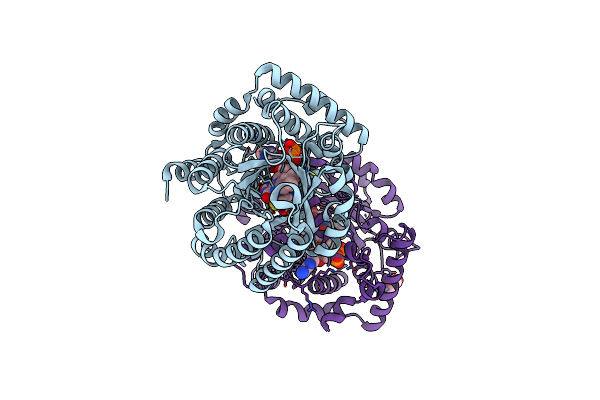
Minimal Puta Proline Dehydrogenase Domain (Design #2) Complexed With (Prop-2-Ynylthio)Acetic Acid
Organism: Sinorhizobium meliloti sm11
Method: X-RAY DIFFRACTION Resolution:1.38 Å Release Date: 2024-10-30 Classification: OXIDOREDUCTASE Ligands: FAD, PEG, X79 |
 |
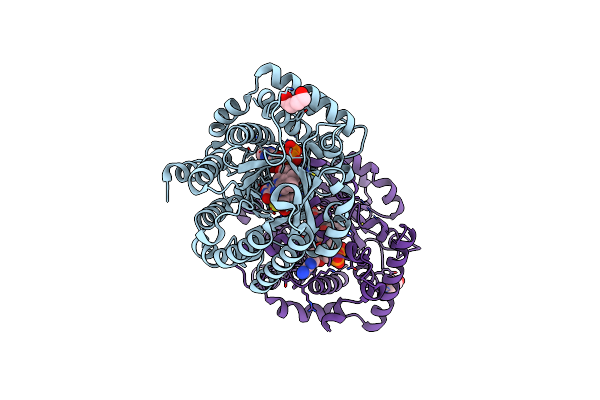
Minimal Puta Proline Dehydrogenase Domain (Design #2) Complexed With 2-(Cyanomethylthio)Acetic Acid
Organism: Sinorhizobium meliloti sm11
Method: X-RAY DIFFRACTION Resolution:1.43 Å Release Date: 2024-10-30 Classification: OXIDOREDUCTASE Ligands: FAD, X7K, PEG |
 |
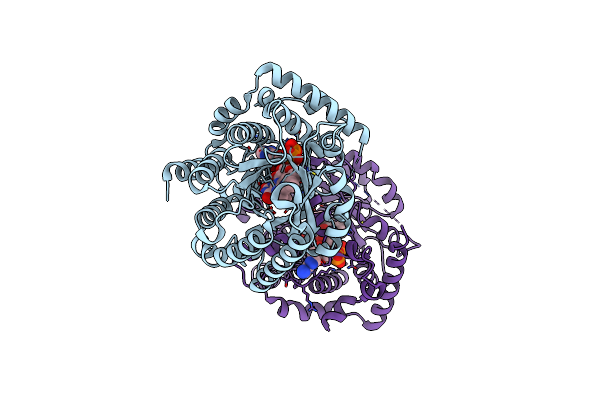
Minimal Puta Proline Dehydrogenase Domain (Design #2) Complexed With (Allylthio)Acetic Acid
Organism: Sinorhizobium meliloti sm11
Method: X-RAY DIFFRACTION Resolution:1.41 Å Release Date: 2024-10-30 Classification: OXIDOREDUCTASE Ligands: FAD, PEG, X7Q |
 |
Minimal Puta Proline Dehydrogenase Domain (Design #2) With The Fad N5 Modified With Propanal Resulting From Inactivation With N-Allylglycine(Replicate #1)
Organism: Sinorhizobium meliloti
Method: X-RAY DIFFRACTION Resolution:1.54 Å Release Date: 2024-10-30 Classification: OXIDOREDUCTASE Ligands: FDA, FAD, CBG |
 |
Minimal Puta Proline Dehydrogenase Domain (Design #2) With The Fad N5 Modified With Propanal Resulting From Inactivation With N-Allylglycine (Replicate #2)
Organism: Sinorhizobium meliloti
Method: X-RAY DIFFRACTION Resolution:1.52 Å Release Date: 2024-10-30 Classification: OXIDOREDUCTASE Ligands: FDA, FAD, CBG, PEG |
 |
Structure Of Proline Utilization A With The Fad Covalently-Modified By Propanal Resulting From Inactivation With N-Allylglycine
Organism: Sinorhizobium meliloti
Method: X-RAY DIFFRACTION Resolution:1.54 Å Release Date: 2024-10-30 Classification: OXIDOREDUCTASE Ligands: FDA, CBG, NAD, MG, SO4, PEG, A1AV8, PGE |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2023-02-08 Classification: OXIDOREDUCTASE Ligands: NAI, SO4 |
 |
Structure Of Proline Utilization A With 1,3-Dithiolane-2-Carboxylate Bound In The Proline Dehydrogenase Active Site
Organism: Sinorhizobium meliloti (strain sm11)
Method: X-RAY DIFFRACTION Resolution:1.63 Å Release Date: 2021-09-29 Classification: OXIDOREDUCTASE Ligands: UJD, FAD, NAD, SO4, MG, PGE, PEG, PG4 |
 |
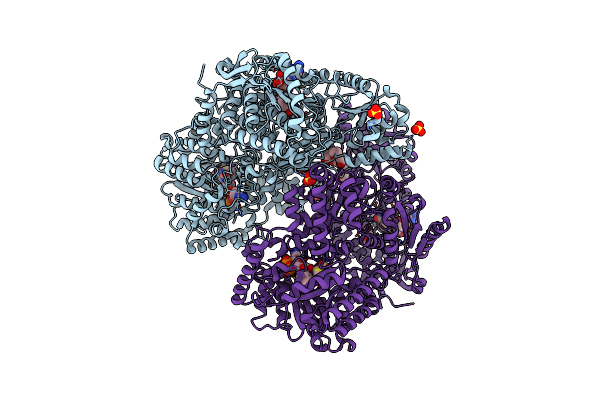
Structure Of Proline Utilization A With The Fad Covalently-Modified By 1,3-Dithiolane
Organism: Sinorhizobium meliloti sm11
Method: X-RAY DIFFRACTION Resolution:1.56 Å Release Date: 2021-09-29 Classification: OXIDOREDUCTASE Ligands: UJJ, MG, NAD, SO4, PGE, PEG, PG4 |
 |
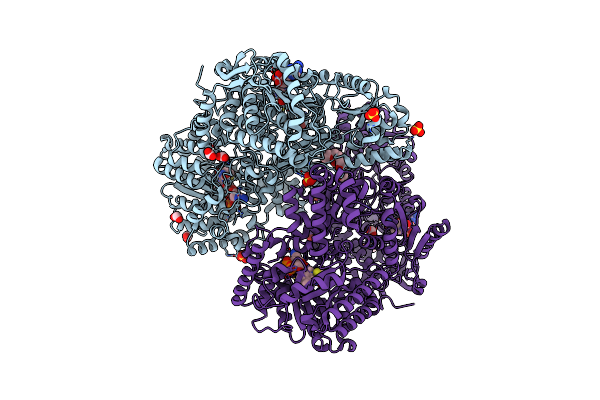
Structure Of Proline Utilization A With Tetrahydrothiophene-2-Carboxylate Bound In The Proline Dehydrogenase Active Site
Organism: Sinorhizobium meliloti (strain sm11)
Method: X-RAY DIFFRACTION Resolution:1.52 Å Release Date: 2021-09-29 Classification: OXIDOREDUCTASE Ligands: UJM, UJP, PGE, FAD, NAD, SO4, MG, PEG |
 |
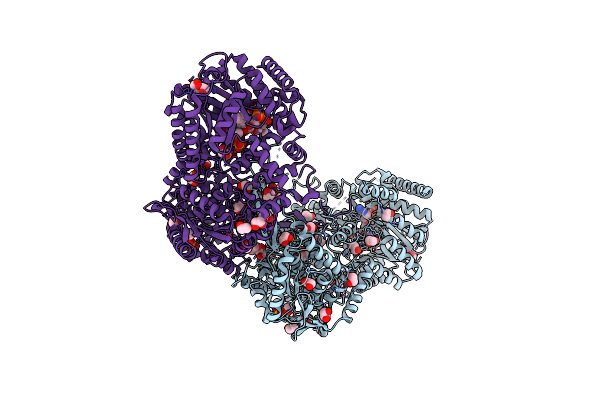
Structure Of Proline Utilization A With The Fad Covalently Modified By Tetrahydrothiophene
Organism: Sinorhizobium meliloti sm11
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2021-09-29 Classification: OXIDOREDUCTASE Ligands: UJG, NAI, SO4, PGE, FMT, MG, PEG |
 |
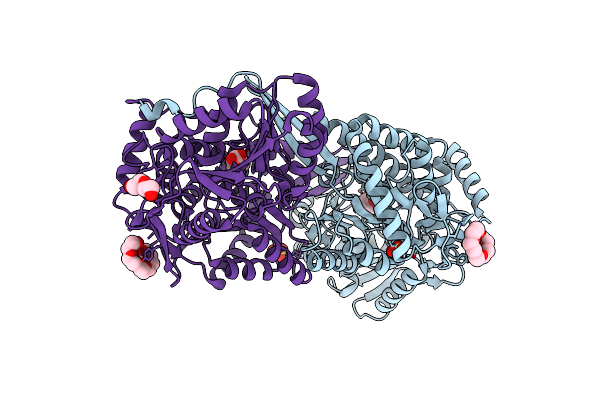
Structure Of Geobacter Sulfurreducens Proline Utilization A (Puta) Variant A206W
Organism: Geobacter sulfurreducens (strain atcc 51573 / dsm 12127 / pca)
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2021-09-22 Classification: OXIDOREDUCTASE Ligands: FAD, SO4, EDO |
 |
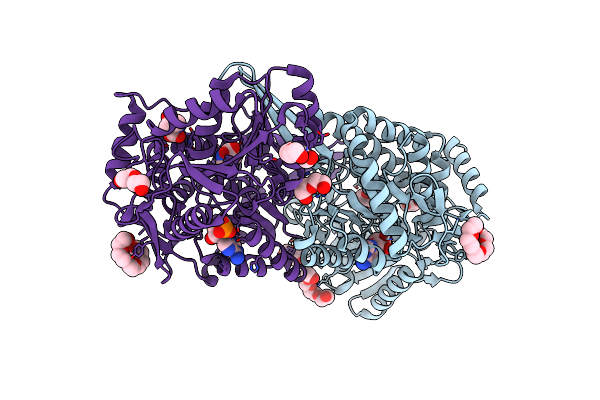
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:1.74 Å Release Date: 2021-06-09 Classification: OXIDOREDUCTASE Ligands: HYP, 1PE, SO4, PEG |
 |
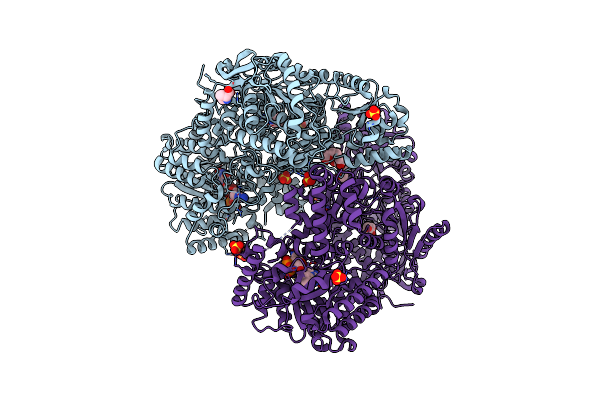
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:1.37 Å Release Date: 2021-06-09 Classification: OXIDOREDUCTASE Ligands: NAD, UY7, PG4, PEG, 1PE, PGE |
 |
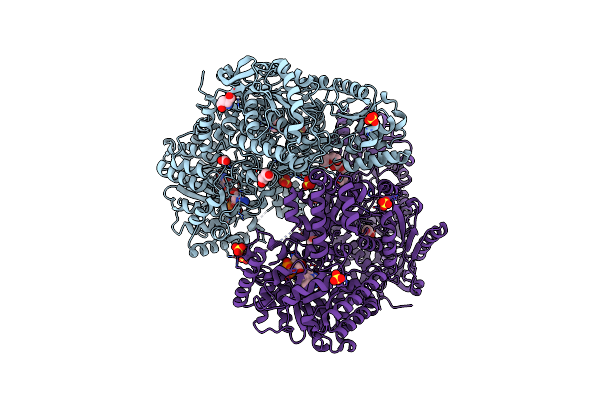
Structure Of Proline Utilization A With D-Proline Bound In The L-Glutamate-Gamma-Semialdehyde Dehydrogenase Active Site
Organism: Sinorhizobium meliloti (strain sm11)
Method: X-RAY DIFFRACTION Resolution:1.56 Å Release Date: 2020-12-30 Classification: OXIDOREDUCTASE Ligands: FAD, SO4, DPR, PGE |
 |
Structure Of Proline Utilization A With Trans-4-Hydroxy-D-Proline Bound In The L-Glutamate-Gamma-Semialdehyde Dehydrogenase Active Site
Organism: Sinorhizobium meliloti (strain sm11)
Method: X-RAY DIFFRACTION Resolution:1.41 Å Release Date: 2020-12-30 Classification: OXIDOREDUCTASE Ligands: FAD, FMT, PEG, UY7, SO4, PGE |

