Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
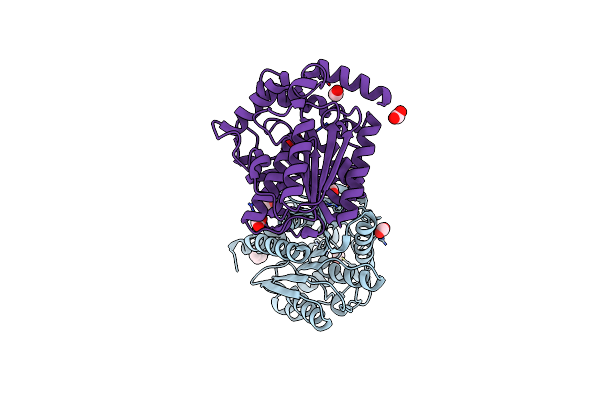 |
Organism: Uncultured bacterium
Method: X-RAY DIFFRACTION Resolution:1.20 Å Release Date: 2021-03-03 Classification: HYDROLASE Ligands: FMT, IPH |
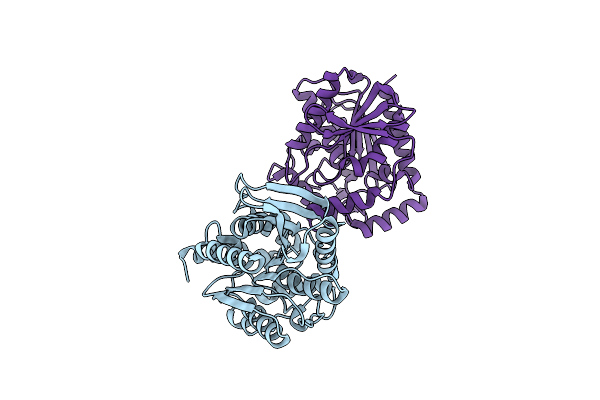 |
Organism: Uncultured bacterium
Method: X-RAY DIFFRACTION Resolution:1.44 Å Release Date: 2021-03-03 Classification: HYDROLASE |
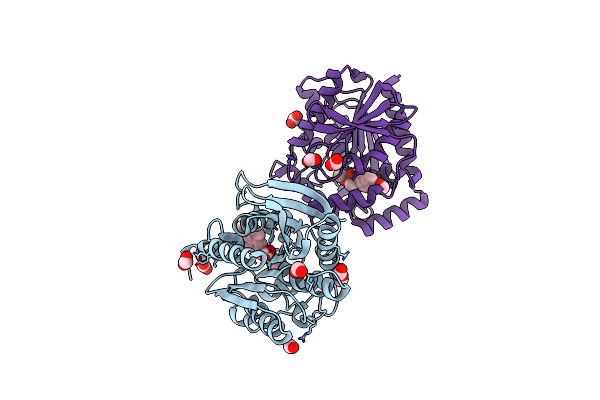 |
Organism: Uncultured bacterium
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2021-03-03 Classification: HYDROLASE Ligands: FMT, NPX, MOH |
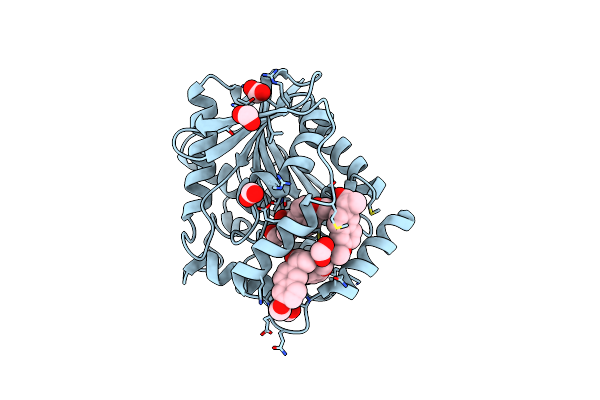 |
Organism: Uncultured bacterium
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2021-03-03 Classification: HYDROLASE Ligands: FMT, NPX |
 |
Organism: Uncultured bacterium
Method: X-RAY DIFFRACTION Resolution:1.45 Å Release Date: 2021-03-03 Classification: HYDROLASE Ligands: FMT, ACT, RXH |
 |
Organism: Uncultured bacterium
Method: X-RAY DIFFRACTION Resolution:1.20 Å Release Date: 2021-03-03 Classification: HYDROLASE Ligands: FMT, NPO, ACT |
 |
Organism: Uncultured bacterium
Method: X-RAY DIFFRACTION Resolution:1.59 Å Release Date: 2021-03-03 Classification: HYDROLASE Ligands: FMT, ACT, RXK |
 |
Organism: Uncultured bacterium
Method: X-RAY DIFFRACTION Resolution:2.28 Å Release Date: 2021-03-03 Classification: HYDROLASE Ligands: FMT, FDS |
 |
Organism: Uncultured bacterium
Method: X-RAY DIFFRACTION Resolution:1.97 Å Release Date: 2021-03-03 Classification: HYDROLASE Ligands: FMT, FDS, ACT |
 |
Organism: Uncultured bacterium
Method: X-RAY DIFFRACTION Resolution:2.13 Å Release Date: 2021-03-03 Classification: HYDROLASE Ligands: U68 |
 |
Organism: Aspergillus niger cbs 513.88
Method: X-RAY DIFFRACTION Resolution:1.71 Å Release Date: 2017-04-19 Classification: HYDROLASE Ligands: NAG, SO4, CL, BTB, GOL |
 |
Structure Of E298Q-Beta-Galactosidase From Aspergillus Niger In Complex With 3-B-Galactopyranosyl Glucose
Organism: Aspergillus niger cbs 513.88
Method: X-RAY DIFFRACTION Resolution:2.45 Å Release Date: 2017-04-19 Classification: HYDROLASE Ligands: NAG, SO4, CL, DMS |
 |
Structure Of E298Q-Beta-Galactosidase From Aspergillus Niger In Complex With Allolactose
Organism: Aspergillus niger cbs 513.88
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2017-04-19 Classification: HYDROLASE Ligands: NAG, SO4, CL, DMS |
 |
Structure Of E298Q-Beta-Galactosidase From Aspergillus Niger In Complex With 6-B-Galactopyranosyl Galactose
Organism: Aspergillus niger cbs 513.88
Method: X-RAY DIFFRACTION Resolution:2.27 Å Release Date: 2017-04-19 Classification: HYDROLASE Ligands: NAG, CL, 1PE |
 |
Structure Of E298Q-Beta-Galactosidase From Aspergillus Niger In Complex With 4-Galactosyl-Lactose
Organism: Aspergillus niger cbs 513.88
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2017-04-19 Classification: HYDROLASE Ligands: NAG |
 |
Structure Of E298Q-Beta-Galactosidase From Aspergillus Niger In Complex With 6-Galactosyl-Lactose
Organism: Aspergillus niger cbs 513.88
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2017-04-19 Classification: HYDROLASE Ligands: NAG, DMS, CL, 1PE |
 |
Structure Of The Beta-Galactosidase From Kluyveromyces Lactis In Complex With Galactose
Organism: Kluyveromyces lactis
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2011-08-17 Classification: HYDROLASE Ligands: GAL, MG, NA, MN |
 |
Organism: Kluyveromyces lactis
Method: X-RAY DIFFRACTION Resolution:2.75 Å Release Date: 2011-08-17 Classification: HYDROLASE Ligands: MN3, GOL |
 |
Organism: Saccharomyces cerevisiae
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2010-06-30 Classification: HYDROLASE Ligands: NAG, GOL, NA, BTB |
 |
Structure Of Alfa-Galactosidase (Mel1) From Saccharomyces Cerevisiae With Melibiose
Organism: Saccharomyces cerevisiae
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2010-06-30 Classification: HYDROLASE Ligands: NAG, NA |