Search Count: 35,679
All
Selected
 |
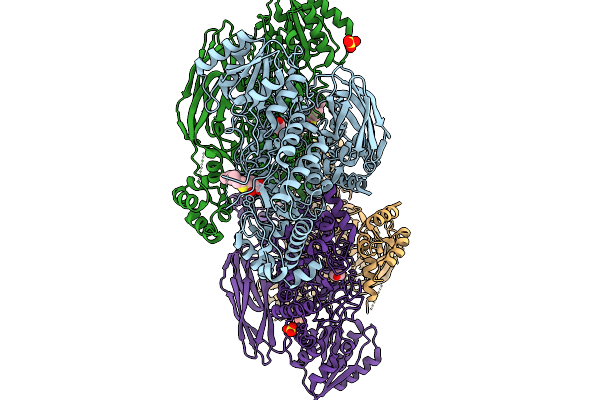
The Structure Of A Bacterial Cyanide Dihydratase From Bacillus Safensis Per-Urp-08
Organism: Bacillus safensis
Method: ELECTRON MICROSCOPY Release Date: 2026-02-25 Classification: HYDROLASE |
 |
Cryo-Em Structure Of Oceanobacillus Iheyensis Group Ii Intron Domains D1-D5
Organism: Oceanobacillus iheyensis
Method: ELECTRON MICROSCOPY Release Date: 2026-02-25 Classification: RNA Ligands: MG, K |
 |
Cryo-Em Structure Of Oceanobacillus Iheyensis Group Ii Intron Domains D1-D5
Organism: Oceanobacillus iheyensis
Method: ELECTRON MICROSCOPY Release Date: 2026-02-25 Classification: RNA Ligands: MG, K |
 |
Organism: Bacillus subtilis subsp. subtilis str. 168
Method: X-RAY DIFFRACTION Resolution:2.64 Å Release Date: 2026-02-25 Classification: PROTEIN BINDING Ligands: ADP, AMP |
 |
Organism: Geobacillus stearothermophilus
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-02-25 Classification: DNA BINDING PROTEIN Ligands: EDO, PGE, CL |
 |
Organism: Bacillus subtilis subsp. subtilis str. 168
Method: X-RAY DIFFRACTION Resolution:2.76 Å Release Date: 2026-02-25 Classification: DNA BINDING PROTEIN Ligands: ADP, PEG, CL |
 |
Organism: Bacillus subtilis subsp. subtilis str. 168
Method: X-RAY DIFFRACTION Resolution:3.35 Å Release Date: 2026-02-25 Classification: DNA BINDING PROTEIN Ligands: B4P, CL |
 |
Organism: Bacillus subtilis subsp. subtilis str. 168
Method: X-RAY DIFFRACTION Resolution:2.62 Å Release Date: 2026-02-25 Classification: DNA BINDING PROTEIN Ligands: ADP, PEG, CL, AMP |
 |
Organism: Bacillus subtilis subsp. subtilis str. 168
Method: X-RAY DIFFRACTION Resolution:3.06 Å Release Date: 2026-02-25 Classification: DNA BINDING PROTEIN Ligands: ADP, EDO, CL, PEG |
 |
Organism: Bacillus subtilis subsp. subtilis str. 168
Method: X-RAY DIFFRACTION Resolution:2.53 Å Release Date: 2026-02-25 Classification: DNA BINDING PROTEIN Ligands: ADP, ATP, EDO, CL, PEG |
 |
Organism: Bacillus subtilis subsp. subtilis str. 168
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2026-02-25 Classification: DNA BINDING PROTEIN Ligands: AMP, PEG, CL |
 |
Organism: Geobacillus phage gbsv1
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: VIRAL PROTEIN |
 |
Organism: Bacillus subtilis subsp. subtilis str. 168
Method: X-RAY DIFFRACTION Resolution:2.41 Å Release Date: 2026-02-18 Classification: BIOSYNTHETIC PROTEIN Ligands: B4P |
 |
Organism: Oceanobacillus iheyensis
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: RNA |
 |
Organism: Paenibacillus
Method: X-RAY DIFFRACTION Resolution:2.04 Å Release Date: 2026-02-18 Classification: HYDROLASE |
 |
Native Structure Of Full-Length Pesticidal Protein Cry1Ca18 At Ph9, From Crystals Formed In Vivo
Organism: Bacillus thuringiensis
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2026-02-11 Classification: TOXIN Ligands: CA, ZN |
 |
Crystal Structure Of Substrate Bound Gh5_22 Exo-Beta-Xylosidase From The Seaweed-Derived Thermophile Geobacillus Thermodenitrificans Os27
Organism: Geobacillus thermodenitrificans
Method: X-RAY DIFFRACTION Resolution:1.52 Å Release Date: 2026-02-04 Classification: HYDROLASE Ligands: GOL, FE |
 |
Crystal Structure Of Gh5_22 Exo-Beta-Xylosidase From The Seaweed-Derived Thermophile Geobacillus Thermodenitrificans Os27
Organism: Geobacillus thermodenitrificans
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2026-02-04 Classification: HYDROLASE Ligands: FE |
 |
Structure Of Stg-Hydrolyzing Beta-Glucosidase 1 (Pstg1) Complexed With Heptyl 1-Thio-Beta-D-Glucopyranoside
Organism: Paenibacillus relictisesami
Method: X-RAY DIFFRACTION Resolution:3.35 Å Release Date: 2026-02-04 Classification: HYDROLASE Ligands: HTG, SO4, EDO, GOL |
 |
Organism: Lactobacillus acidophilus ncfm
Method: X-RAY DIFFRACTION Resolution:1.93 Å Release Date: 2026-02-04 Classification: STRUCTURAL PROTEIN Ligands: GOL, FMT |

