Search Count: 2,253
All
Selected
 |
Cryo-Em Structure Of Amppnp Bound Human Phosphoribosylformylglycinamidine Synthase
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.31 Å Release Date: 2026-02-25 Classification: LIGASE Ligands: ADP, ANP, MG |
 |
Cryo-Em Structure Of Ternary Complex Of Human Phosphoribosylglycinamidine Synthase With The Intermediate (Iminophosphate) And Adp Bound At The Synthase Site.
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.22 Å Release Date: 2026-02-25 Classification: LIGASE Ligands: ADP, A1B7I, MG |
 |
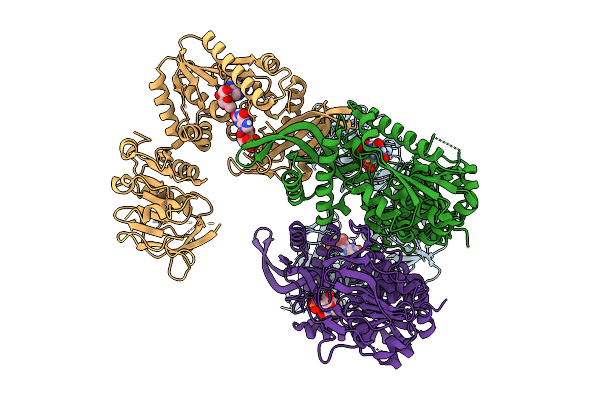
Cryo-Em Structure Of Quaternary Complex Of Human Phosphoribosylglycinamidine Synthase With Thioester Intermediate Bound (At Glutaminase Site) And Amppnp And Fgar (At Synthase Site).
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.19 Å Release Date: 2026-02-25 Classification: LIGASE Ligands: FGR, ANP, ADP, GLN, MG |
 |
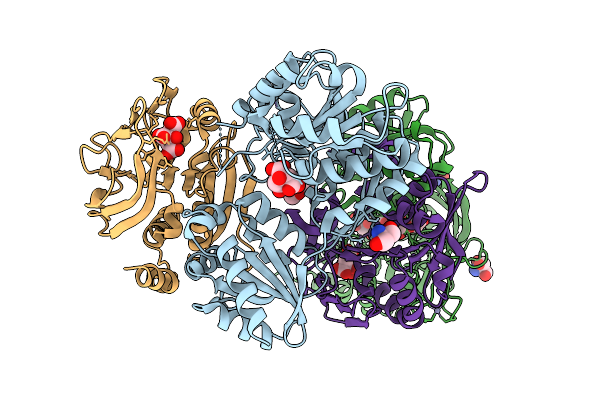
Organism: Escherichia coli k-12
Method: X-RAY DIFFRACTION Resolution:2.51 Å Release Date: 2025-12-24 Classification: LIGASE/ANTIBIOTIC Ligands: XMP, A1L67 |
 |
Crystal Structure Of The Murt/Gatd Enzyme Complex From Streptococcus Pyogenes
Organism: Streptococcus pyogenes mgas10270
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2025-10-08 Classification: LIGASE Ligands: ZN, CIT, TRS, PEG |
 |
Crystal Structure Of The Murt/Gatd Enzyme Complex From Streptococcus Pyogenes With Bound Amppnp
Organism: Streptococcus pyogenes mgas10270
Method: X-RAY DIFFRACTION Resolution:1.98 Å Release Date: 2025-10-08 Classification: LIGASE Ligands: ZN, MG, ANP, GOL |
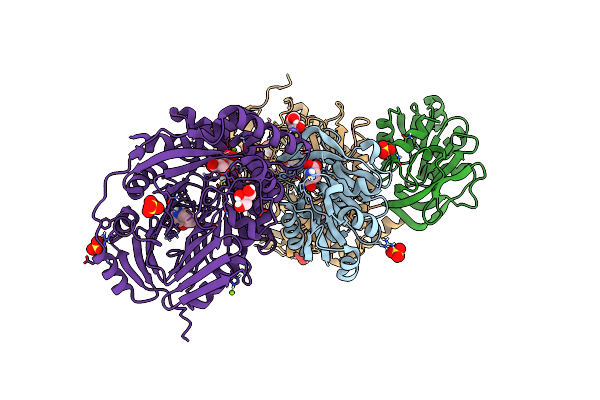 |
Aminodeoxychorismate Synthase Complex From Escherichia Coli, With Glutamine And Chorismate Added
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.64 Å Release Date: 2025-01-29 Classification: TRANSFERASE Ligands: GLU, GOL, SO4, ISJ, TRP, MG, CL |
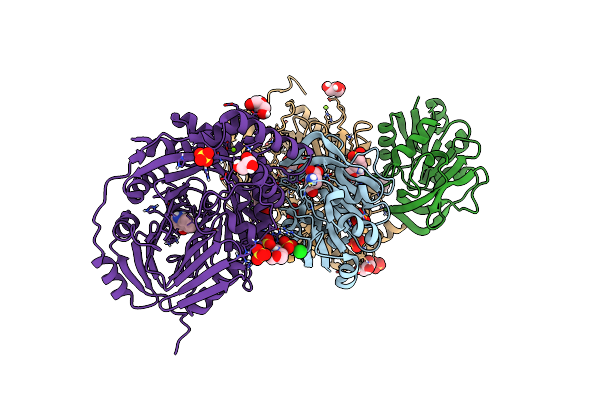 |
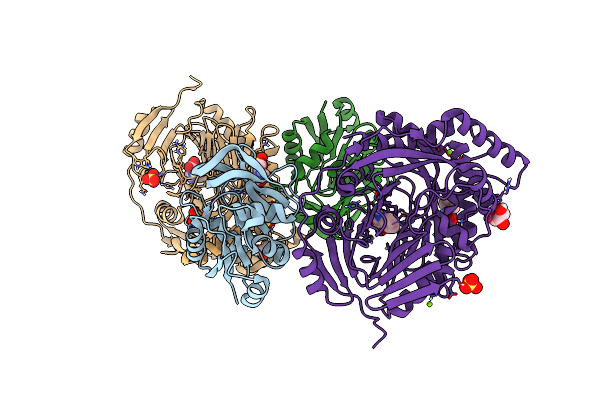
Aminodeoxychorismate Synthase Complex From Escherichia Coli, With Glutamine Added
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.86 Å Release Date: 2025-01-29 Classification: TRANSFERASE Ligands: GLU, CL, SO4, MES, TRP, MG, GOL |
 |
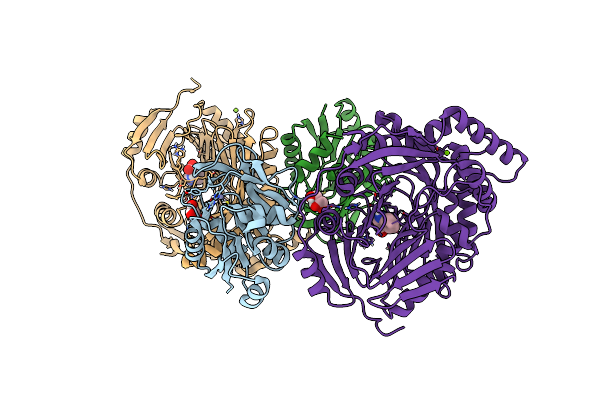
Aminodeoxychorismate Synthase Complex From Escherichia Coli, With Edta Added
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.98 Å Release Date: 2025-01-29 Classification: TRANSFERASE Ligands: CL, SO4, TRP, MG, MES, GOL |
 |
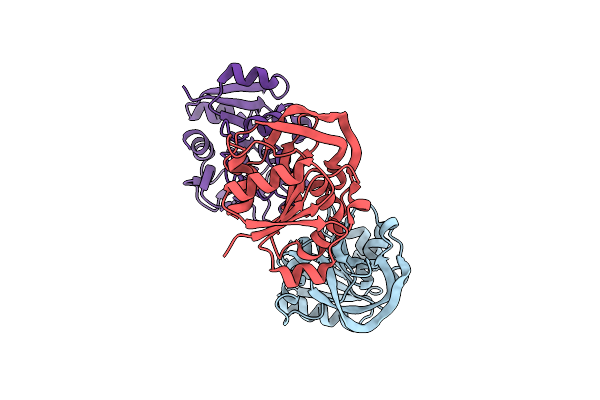
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.45 Å Release Date: 2025-01-29 Classification: TRANSFERASE Ligands: ZN, GOL, TRP, MG, CL |
 |
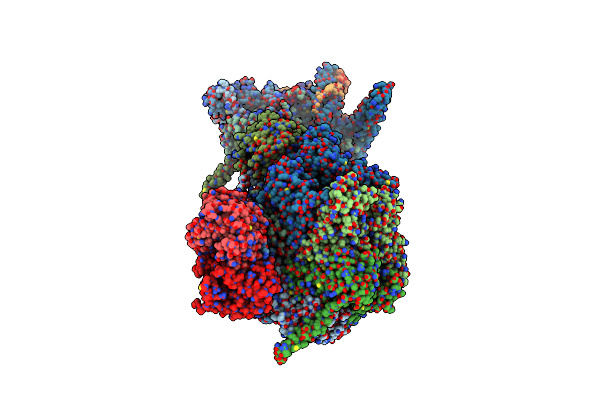
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.82 Å Release Date: 2025-01-29 Classification: TRANSFERASE |
 |
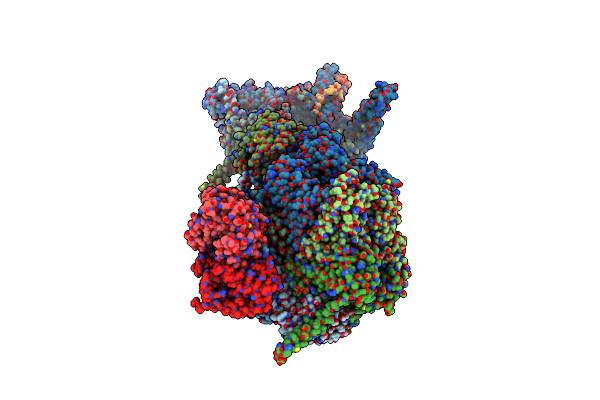
Cryo-Em Structure Of The Ycf2-Ftshi Motor Complex From Arabidopsis In Atp-Bound State
Organism: Arabidopsis thaliana
Method: ELECTRON MICROSCOPY Release Date: 2024-09-11 Classification: MEMBRANE PROTEIN Ligands: MG, ATP, ZN, PX2 |
 |
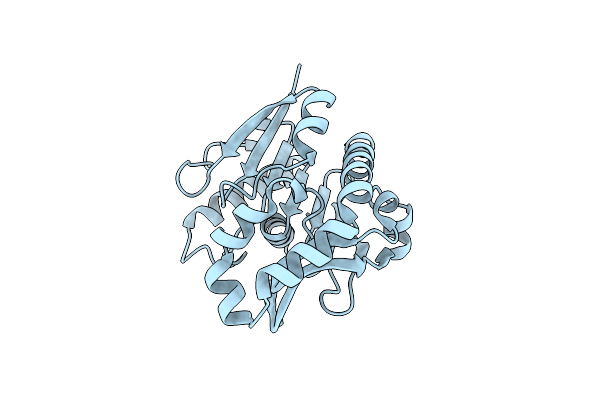
Cryo-Em Structure Of The Ycf2-Ftshi Motor Complex From Arabidopsis In Apo State
Organism: Arabidopsis thaliana
Method: ELECTRON MICROSCOPY Release Date: 2024-09-11 Classification: MEMBRANE PROTEIN Ligands: ZN, PX2 |
 |
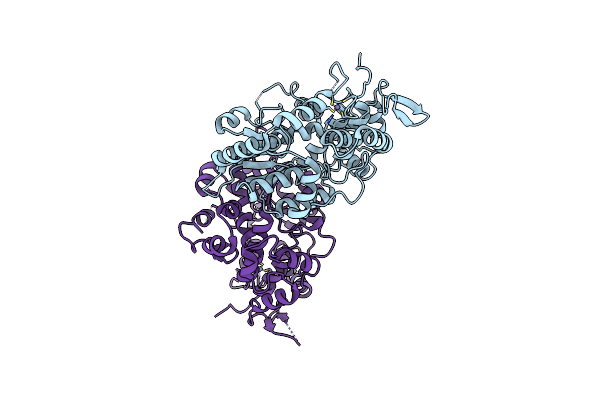
Organism: Alcaligenes ammonioxydans
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2024-09-04 Classification: TRANSFERASE |
 |
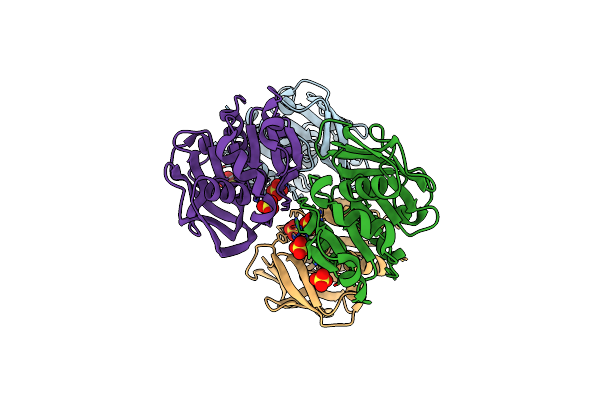
Crystal Structure Of N-Acetyl Sugar Amidotransferase From Legionella Pneumophila
Organism: Legionella pneumophila
Method: X-RAY DIFFRACTION Resolution:2.33 Å Release Date: 2024-05-22 Classification: TRANSFERASE Ligands: ZN |
 |
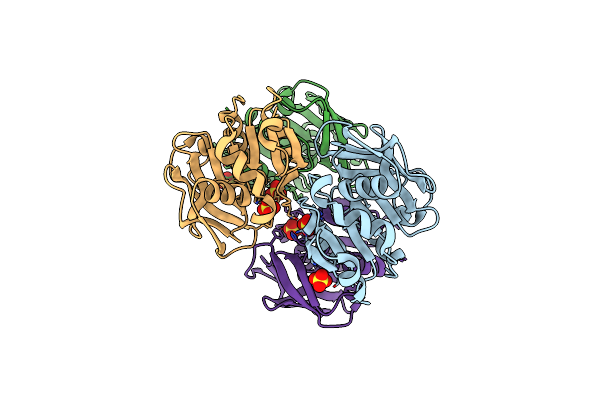
Crystal Structure Of D110V/K151L Mutant Of Gatase Subunit Of Methanocaldococcus Jannaschii Gmp Synthetase
Organism: Methanocaldococcus jannaschii dsm 2661
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2023-07-12 Classification: TRANSFERASE Ligands: SO4 |
 |
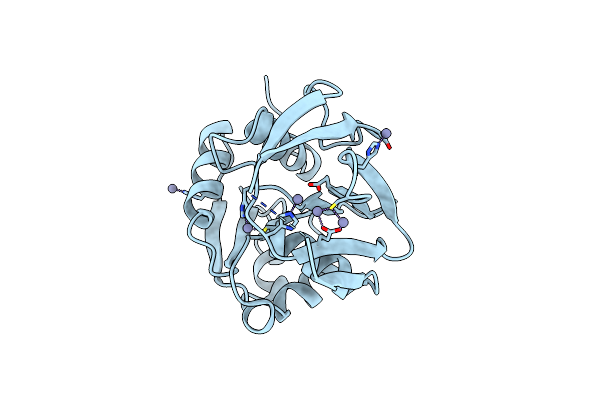
Crystal Structure Of K151L/Y158F Mutant Of Gatase Subunit Of Methanocaldococcus Jannaschii Gmp Synthetase
Organism: Methanocaldococcus jannaschii dsm 2661
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2023-07-12 Classification: TRANSFERASE Ligands: SO4 |
 |
Crystal Structure Of D110P Mutant Of Gatase Subunit Of Methanocaldococcus Jannaschii Gmp Synthetase
Organism: Methanocaldococcus jannaschii
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2023-06-14 Classification: TRANSFERASE Ligands: ZN |
 |
Crystal Structure Of The Murt-Gatd Enzyme Complex From Staphylococcus Aureus Col Strain
Organism: Staphylococcus aureus subsp. aureus col
Method: X-RAY DIFFRACTION Resolution:2.90 Å Release Date: 2023-05-24 Classification: LIGASE Ligands: ZN, CL |
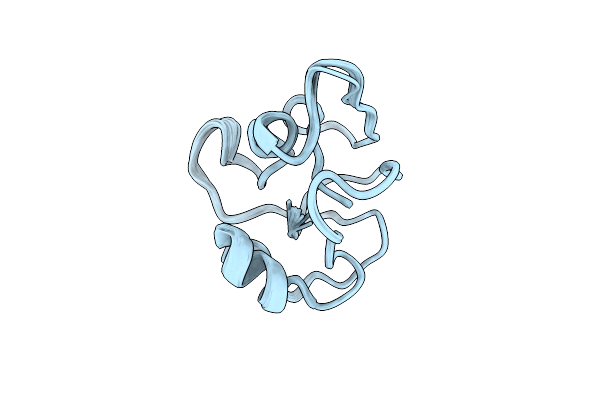 |
Solution Nmr Structure Of Purs Subunit Of Phosphoribosylformylglycinamidine Synthase Enzyme From Staphylococcus Aureus
Organism: Staphylococcus aureus subsp. aureus mu50
Method: SOLUTION NMR Release Date: 2023-05-17 Classification: LYASE |

