Search Count: 8,844
All
Selected
 |
Ecoli Dnab Helicase And Phage Lambda Loader P With Adp-Mg In A 6:5 Stoichiometry Ratio
Organism: Escherichia coli, Escherichia phage lambda
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: DNA BINDING PROTEIN Ligands: ADP, MG |
 |
Crystal Structure Of The Ribokinase Rbk1 In Complex With Adp From Saccharomyces Cerevisiae
Organism: Saccharomyces cerevisiae s288c
Method: X-RAY DIFFRACTION Resolution:2.66 Å Release Date: 2026-03-04 Classification: PROTEIN BINDING Ligands: ADP, MG, PO4, NA |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: DNA BINDING PROTEIN Ligands: ZN |
 |
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: PROTON TRANSPORT Ligands: ADP |
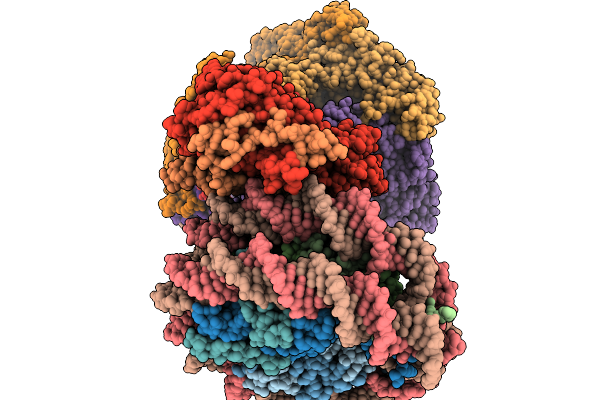 |
Organism: Saccharomyces cerevisiae, Xenopus laevis, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: DNA BINDING PROTEIN/DNA Ligands: ADP |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.27 Å Release Date: 2026-03-04 Classification: TRANSLATION Ligands: ADP, ZN, CA, MG |
 |
Phosphorylation Of A Conserved Aspartate At The Eukaryotic Elongation Factor 2 Kinase Catalytic Site
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2026-03-04 Classification: TRANSLATION Ligands: ADP, MN, ZN, CA |
 |
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: DNA BINDING PROTEIN/DNA Ligands: ADP, MG, ATP |
 |
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: DNA BINDING PROTEIN/DNA Ligands: ADP, MG, ATP |
 |
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: DNA BINDING PROTEIN/DNA Ligands: ADP, MG, ATP |
 |
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: DNA BINDING PROTEIN/DNA Ligands: ADP, CA, ATP |
 |
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: DNA BINDING PROTEIN/DNA Ligands: ADP, MG, ATP |
 |
Organism: Homo sapiens, Oryctolagus cuniculus
Method: ELECTRON MICROSCOPY Resolution:3.13 Å Release Date: 2026-03-04 Classification: PROTEIN FIBRIL Ligands: ADP, MG |
 |
Organism: Homo sapiens, Oryctolagus cuniculus
Method: ELECTRON MICROSCOPY Resolution:3.68 Å Release Date: 2026-03-04 Classification: PROTEIN FIBRIL Ligands: ADP, MG |
 |
Organism: Homo sapiens, Oryctolagus cuniculus
Method: ELECTRON MICROSCOPY Resolution:3.50 Å Release Date: 2026-03-04 Classification: PROTEIN FIBRIL Ligands: ADP, MG |
 |
Organism: Homo sapiens, Oryctolagus cuniculus
Method: ELECTRON MICROSCOPY Resolution:3.48 Å Release Date: 2026-03-04 Classification: PROTEIN FIBRIL Ligands: ADP, MG |
 |
Organism: Homo sapiens, Oryctolagus cuniculus
Method: ELECTRON MICROSCOPY Resolution:3.02 Å Release Date: 2026-03-04 Classification: PROTEIN FIBRIL Ligands: ADP, MG |
 |
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: TRANSPORT PROTEIN Ligands: ADP, MG |
 |
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: TRANSPORT PROTEIN Ligands: ADP, MG |
 |
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: TRANSPORT PROTEIN Ligands: ADP, MG |

