Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
 |
Organism: Bacillus, Bacillaceae bacterium
Method: X-RAY DIFFRACTION Release Date: 2026-01-14 Classification: VIRAL PROTEIN Ligands: ZN, ACT, IMD, GOL |
 |
Organism: Mesorhizobium metallidurans
Method: X-RAY DIFFRACTION Release Date: 2025-12-10 Classification: OXIDOREDUCTASE Ligands: P6G, PEG, EPE, NA, TRS |
 |
Lystt72, A Lytic Endopeptidase From Thermus Thermophilus Mat72 Phage Vb_Tt72
Organism: Thermus phage tt72, Escherichia coli bl21(de3)
Method: X-RAY DIFFRACTION Release Date: 2025-03-05 Classification: VIRAL PROTEIN Ligands: ZN, PO4 |
 |
Lystt72, A Lytic Endopeptidase From Thermus Thermophilus Mat72 Phage Vb_Tt72
Organism: Thermus phage tt72, Escherichia coli bl21(de3)
Method: X-RAY DIFFRACTION Release Date: 2025-03-05 Classification: VIRAL PROTEIN Ligands: ZN, PO4 |
 |
Lystt72, A Lytic Endopeptidase From Thermus Thermophilus Mat72 Phage Vb_Tt72
Organism: Thermus phage tt72, Escherichia coli bl21(de3)
Method: X-RAY DIFFRACTION Release Date: 2025-03-05 Classification: VIRAL PROTEIN Ligands: ZN, PO4 |
 |
Organism: Amycolatopsis thermoflava n1165
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2024-09-25 Classification: LIGASE Ligands: PEG |
 |
Crystal Structure Of Feruoyl-Coa Synthetase Complexed With Amp From Amycolatopsis Thermoflava
Organism: Amycolatopsis thermoflava n1165
Method: X-RAY DIFFRACTION Resolution:2.29 Å Release Date: 2024-09-25 Classification: LIGASE Ligands: AMP, MG |
 |
Organism: Amycolatopsis thermoflava n1165
Method: X-RAY DIFFRACTION Resolution:1.98 Å Release Date: 2024-07-24 Classification: OXIDOREDUCTASE Ligands: HEM, GOL, ACT |
 |
Organism: Amycolatopsis thermoflava n1165
Method: X-RAY DIFFRACTION Resolution:1.26 Å Release Date: 2024-07-24 Classification: OXIDOREDUCTASE Ligands: HEM, 3DM, NO3, MG |
 |
Organism: Amycolatopsis thermoflava n1165
Method: X-RAY DIFFRACTION Resolution:1.12 Å Release Date: 2024-07-24 Classification: OXIDOREDUCTASE Ligands: HEM, UB9, NO3 |
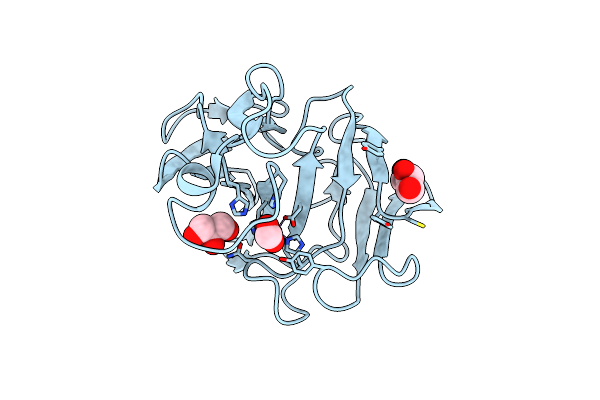 |
Organism: Lysobacter capsici
Method: X-RAY DIFFRACTION Resolution:1.57 Å Release Date: 2023-08-16 Classification: HYDROLASE Ligands: ZN, CL, GOL, FMT |
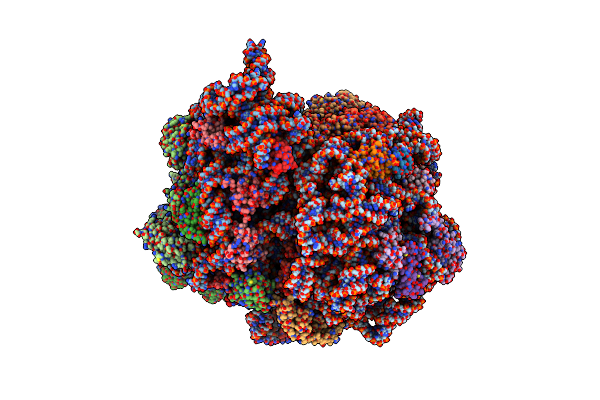 |
Late Translocation Intermediate With Ef-G Partially Dissociated (Structure V)
Organism: Escherichia coli 113303, Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2022-02-23 Classification: RIBOSOME |
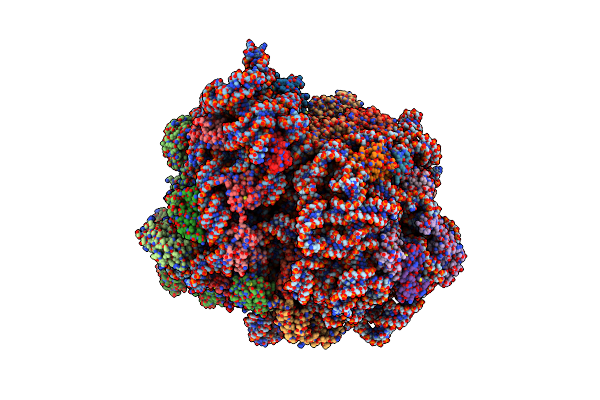 |
Organism: Escherichia coli 113303, Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2022-02-23 Classification: RIBOSOME Ligands: GDP |
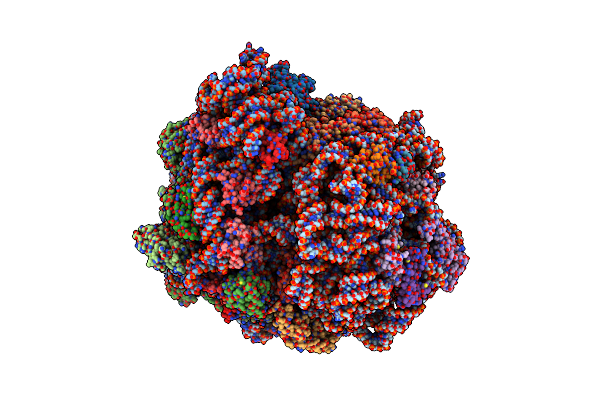 |
Pre Translocation Intermediate With Ef-G Bound To Gdp And Pi (Structure Iii)
Organism: Escherichia coli 113303, Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2022-02-23 Classification: RIBOSOME Ligands: GDP, PO4 |
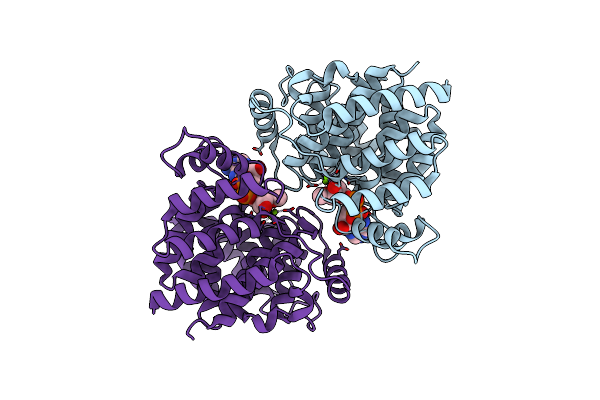 |
Adp-Ribosylserine Hydrolase Arh3 Of Latimeria Chalumnae In Complex With Alpha-1''-O-Methyl-Adp-Ribose (Meadpr)
Organism: Latimeria chalumnae
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2021-06-16 Classification: HYDROLASE Ligands: MG, RVK |
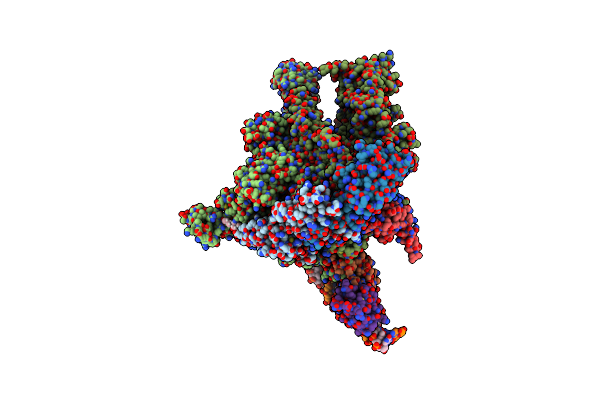 |
Cryo-Em Structure Of Escherichia Coli Rnap Polymerase Bound To Rpstp2 Promoter Dna
Organism: Escherichia coli, Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2020-02-26 Classification: TRANSCRIPTION Ligands: 1N7, MG, ZN |
 |
Organism: Shigella flexneri 2a
Method: X-RAY DIFFRACTION Resolution:2.51 Å Release Date: 2019-03-06 Classification: HYDROLASE Ligands: SO4, PG4, PEG, EDO, GOL, CL |
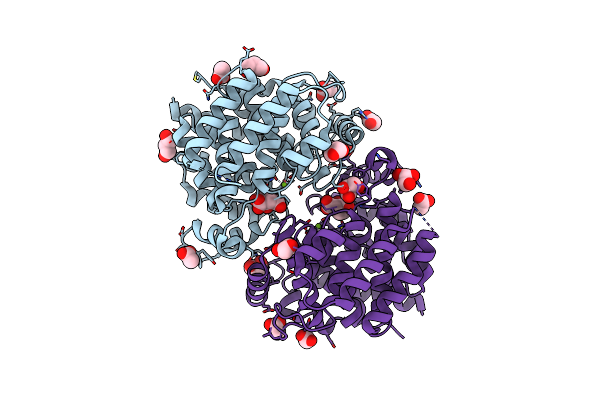 |
Organism: Latimeria chalumnae
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2018-11-28 Classification: HYDROLASE Ligands: MG, CIT, ACT, GOL |
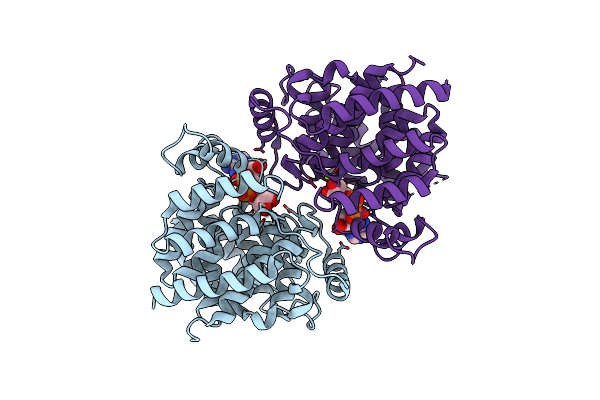 |
Adp-Ribosylserine Hydrolase Arh3 Of Latimeria Chalumnae In Complex With Adp-Ribose
Organism: Latimeria chalumnae
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2018-11-28 Classification: HYDROLASE Ligands: MG, AR6 |
 |
Adp-Ribosylserine Hydrolase Arh3 Of Latimeria Chalumnae In Complex With Adp-Ribose
Organism: Latimeria chalumnae
Method: X-RAY DIFFRACTION Resolution:1.86 Å Release Date: 2018-11-28 Classification: HYDROLASE Ligands: MG, AR6, GOL |