Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
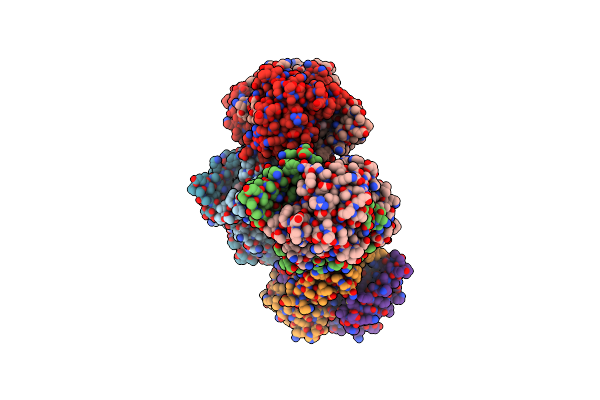 |
E. Coli Nfsb With The Unnatural Amino Acid P-Nitrophenylalanine At Position 124.
Organism: Escherichia coli dh5[alpha]
Method: X-RAY DIFFRACTION Resolution:2.74 Å Release Date: 2024-08-14 Classification: OXIDOREDUCTASE Ligands: FMN, ACT |
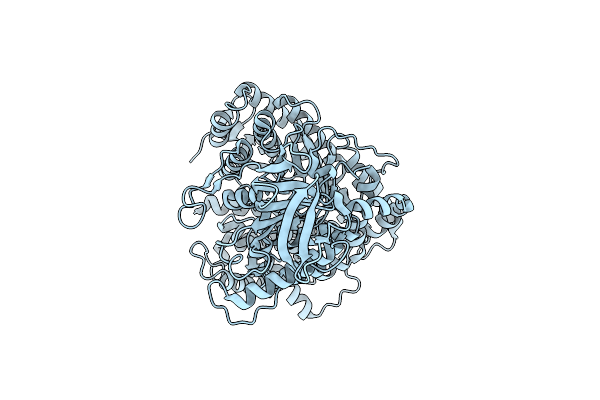 |
Organism: Escherichia coli dh5[alpha]
Method: ELECTRON MICROSCOPY Release Date: 2023-03-29 Classification: BIOSYNTHETIC PROTEIN |
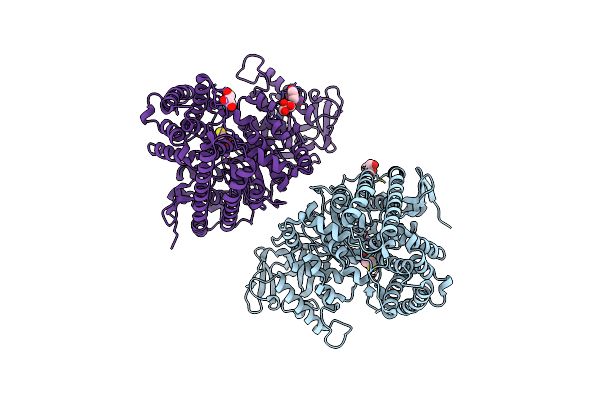 |
Organism: Escherichia coli dh5[alpha]
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2023-03-29 Classification: BIOSYNTHETIC PROTEIN Ligands: CL, NA, BME, GOL |
 |
Organism: Escherichia coli dh5[alpha]
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2022-02-02 Classification: TRANSLATION Ligands: 0O2 |
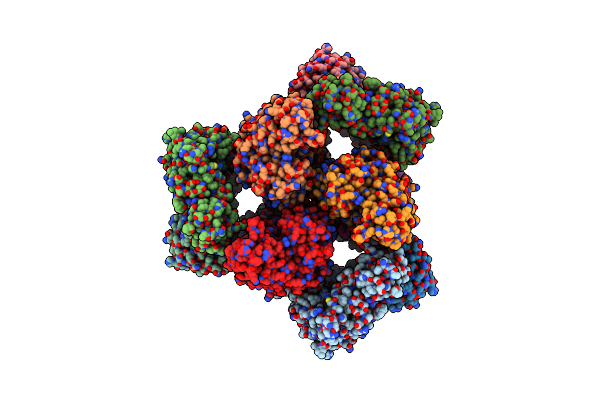 |
Organism: Escherichia coli dh5[alpha]
Method: ELECTRON MICROSCOPY Release Date: 2019-07-31 Classification: ANTIBIOTIC |
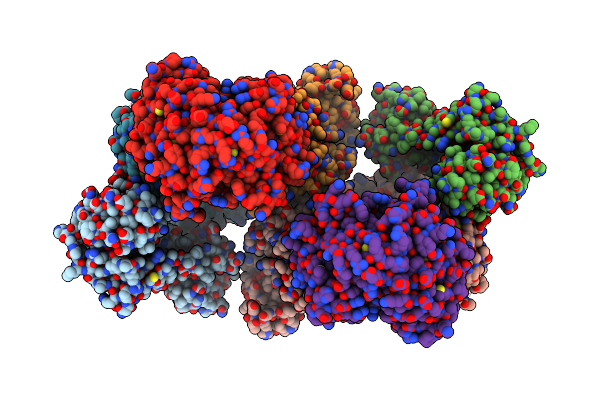 |
Organism: Escherichia coli dh5[alpha]
Method: ELECTRON MICROSCOPY Release Date: 2019-07-31 Classification: ANTIBIOTIC |
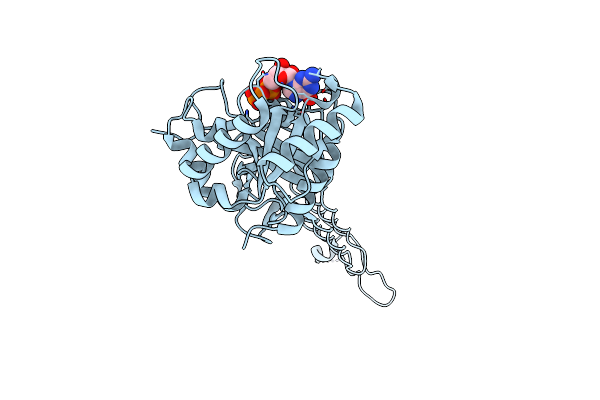 |
Organism: Escherichia coli dh5[alpha]
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2017-03-01 Classification: HYDROLASE Ligands: GDP, MG |
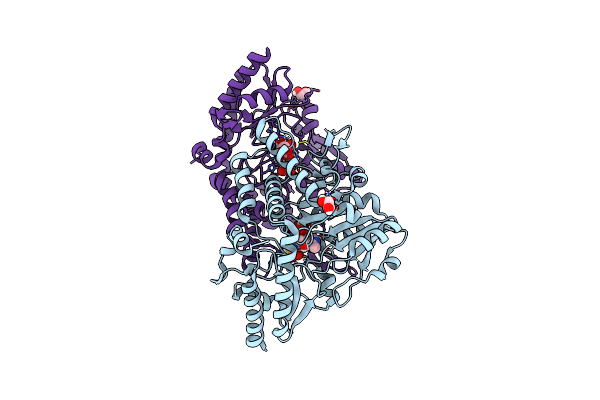 |
Crystal Structure Of Plp-Bound E. Coli Sufs (Cysteine Persulfide Intermediate) In Space Group P21
Organism: Escherichia coli dh5[alpha]
Method: X-RAY DIFFRACTION Resolution:2.75 Å Release Date: 2016-08-24 Classification: TRANSFERASE, LYASE Ligands: PLP, EDO, CYS, CIT |
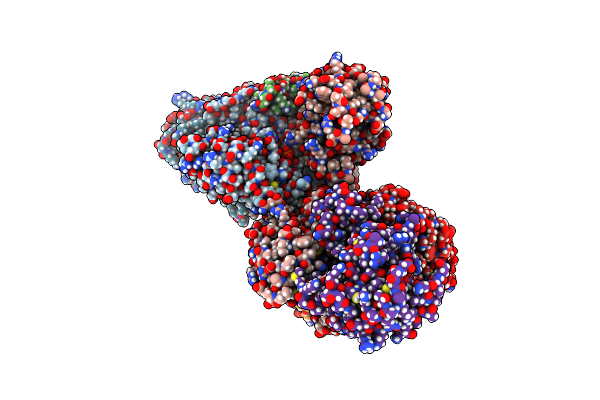 |
Organism: Escherichia coli dh5[alpha]
Method: X-RAY DIFFRACTION Resolution:2.55 Å Release Date: 2016-08-24 Classification: TRANSFERASE Ligands: PO4, PEG, CL, GOL |
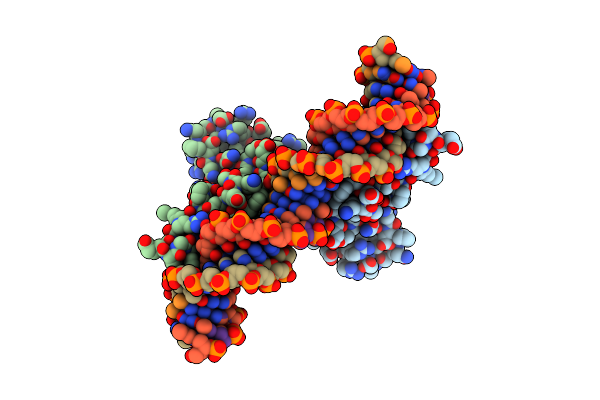 |
Crystal Structure Of The Metal-Free (Repressor) Form Of E. Coli Cuer, A Copper Efflux Regulator, Bound To Copa Promoter Dna
Organism: Escherichia coli dh5[alpha], Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.11 Å Release Date: 2015-09-02 Classification: Transcription Regulator/DNA |
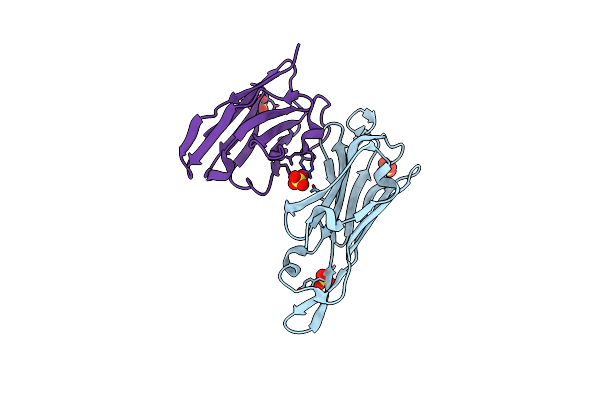 |
Co-Complex Structure Of The Lectin Domain Of F18 Fimbrial Adhesin Fedf With Inhibitory Nanobody Nbfedf9
Organism: Escherichia coli dh5[alpha], Lama glama
Method: X-RAY DIFFRACTION Resolution:1.57 Å Release Date: 2014-12-17 Classification: CELL ADHESION Ligands: SO4 |
 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2013-10-09 Classification: HYDROLASE/DNA Ligands: GOL, SO4 |
 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.96 Å Release Date: 2013-10-09 Classification: HYDROLASE/DNA Ligands: SO4, ZN, IPA, PO4 |
 |
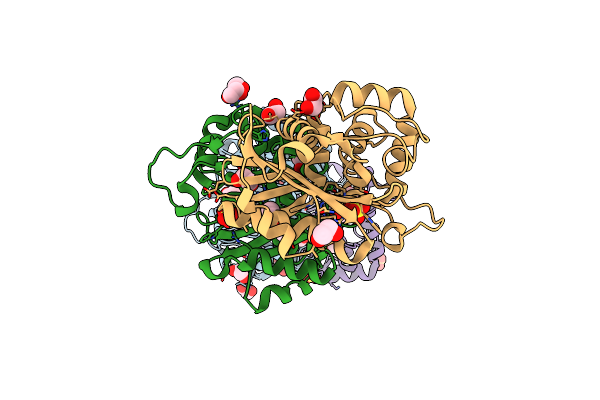
Dual-Target Inhibitor Of Murd And Mure Ligases: Design, Synthesis And Binding Mode Studies
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.49 Å Release Date: 2011-12-28 Classification: LIGASE Ligands: T26, DMS, SO4 |
 |
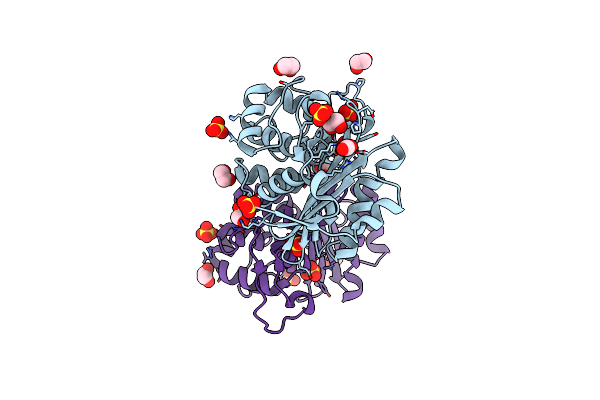
Crystal Structure Of An Engineered Oxa-10 Variant With Carbapenemase Activity, Oxa-10Loop24
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2011-11-02 Classification: HYDROLASE Ligands: SO4, EDO |
 |
Crystal Structure Of A Rationally Designed Oxa-10 Variant Showing Carbapenemase Activity, Oxa-10Loop48
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2011-11-02 Classification: HYDROLASE Ligands: SO4, EDO, CO2 |
 |
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:9.80 Å Release Date: 2011-08-24 Classification: RIBOSOME/HYDROLASE |
 |
Discovery Of Novel 5-Benzylidenerhodanine- And 5-Benzylidene- Thiazolidine-2,4-Dione Inhibitors Of Murd Ligase
Organism: Escherichia coli dh5[alpha]
Method: X-RAY DIFFRACTION Resolution:1.46 Å Release Date: 2010-09-15 Classification: LIGASE Ligands: VSV, AZI, SO4, SO3, CL |
 |
Crystal Structure Of Murd Ligase In Complex With D-Glu Containing Rhodanine Inhibitor
Organism: Escherichia coli dh5[alpha]
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2010-08-25 Classification: LIGASE Ligands: D17, DMS, AZI, SO4, CL |