Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
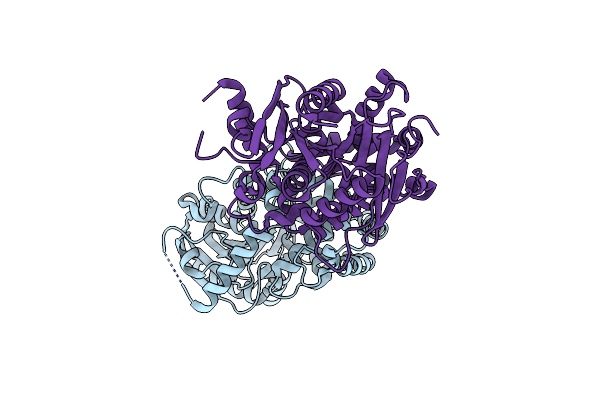 |
Crystal Structure Of An Epoxide Hydrolase Mutant A250Ic/L344V From Aspergillus Usamii E001 At 2.17 Angstroms Resolution
Organism: Aspergillus usamii
Method: X-RAY DIFFRACTION Resolution:2.18 Å Release Date: 2024-02-07 Classification: HYDROLASE |
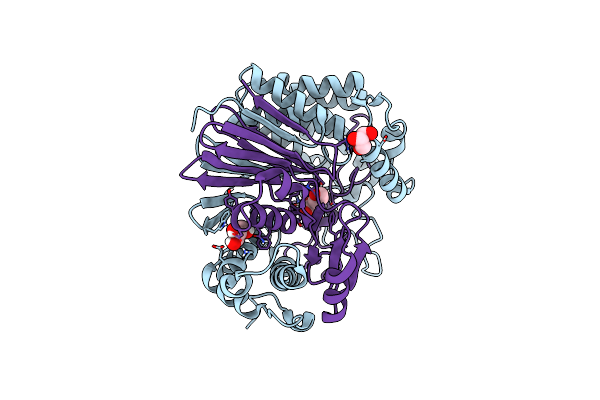 |
Gamma-Glutamyltranspeptidase From Pseudomonas Nitroreducens Complexed With L-Don
Organism: Pseudomonas nitroreducens
Method: X-RAY DIFFRACTION Release Date: 2021-06-23 Classification: TRANSFERASE Ligands: GOL, DON |
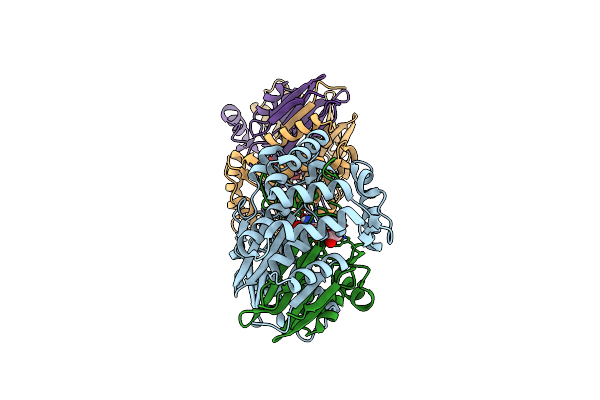 |
Gamma-Glutamyltranspeptidase From Pseudomonas Nitroreducens Complexed With L-Don
Organism: Pseudomonas nitroreducens
Method: X-RAY DIFFRACTION Release Date: 2021-06-23 Classification: TRANSFERASE Ligands: DON, GLY |
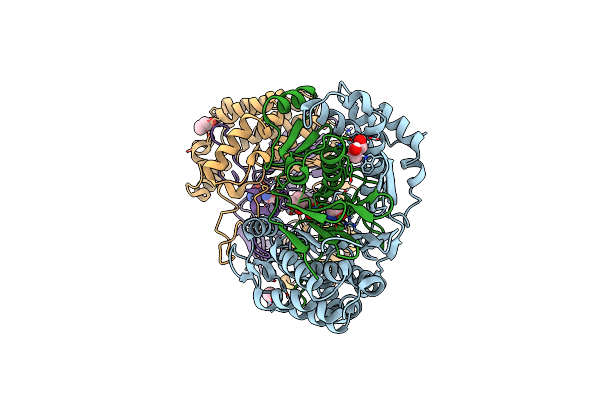 |
Highly Active Mutant W525D Of Gamma-Glutamyltranspeptidase From Pseudomonas Nitroreducens
Organism: Pseudomonas nitroreducens
Method: X-RAY DIFFRACTION Resolution:1.74 Å Release Date: 2021-06-23 Classification: TRANSFERASE Ligands: GBL, GLY, GOL |
 |
Organism: Megabalanus rosa
Method: SOLUTION NMR Release Date: 2020-01-15 Classification: STRUCTURAL PROTEIN |
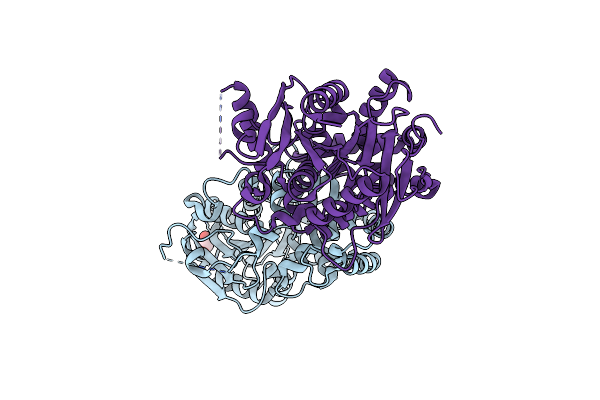 |
Structure Of The A214C/A250I Mutant Of An Epoxide Hydrolase From Aspergillus Usamii E001 (Aueh2) At 1.48 Angstroms Resolution
Organism: Aspergillus usamii
Method: X-RAY DIFFRACTION Resolution:1.48 Å Release Date: 2019-12-11 Classification: HYDROLASE Ligands: GOL |
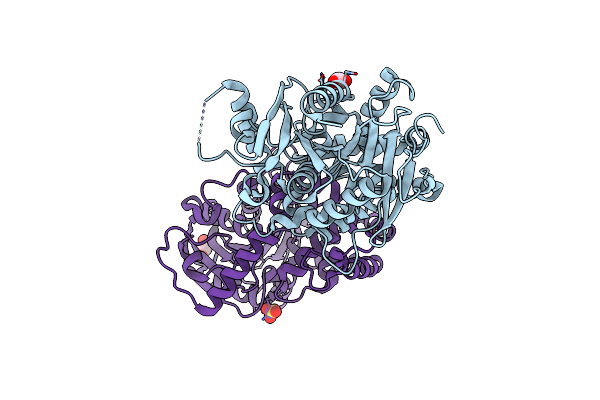 |
Structure Of An Epoxide Hydrolase From Aspergillus Usamii E001 (Aueh2) At 1.51 Angstroms Resolution
Organism: Aspergillus usamii
Method: X-RAY DIFFRACTION Resolution:1.51 Å Release Date: 2019-12-11 Classification: HYDROLASE Ligands: ACT, GOL, SO4, CL, NA |
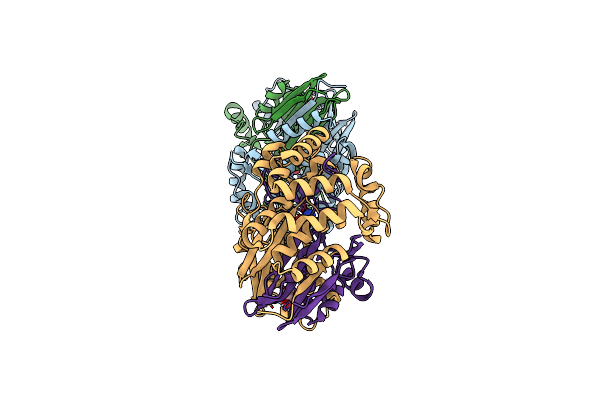 |
Gamma-Glutamyltranspeptidase From Pseudomonas Nitroreducens Complexed With Gly-Gly
Organism: Pseudomonas nitroreducens
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2019-01-16 Classification: TRANSFERASE Ligands: GLY, GOL |
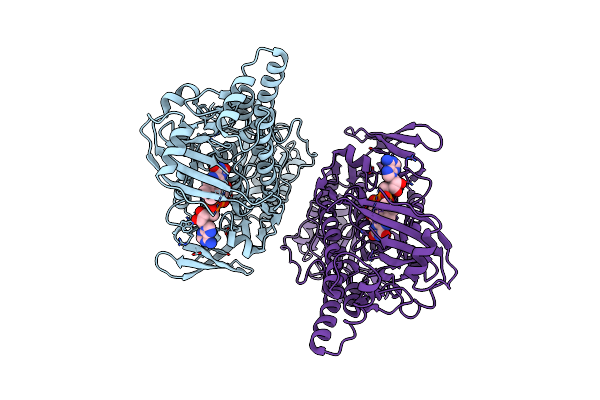 |
Crystal Structure Of 2-Hydroxybiphenyl 3-Monooxygenase M321A From Pseudomonas Azelaica
Organism: Pseudomonas nitroreducens
Method: X-RAY DIFFRACTION Resolution:2.78 Å Release Date: 2018-01-10 Classification: OXIDOREDUCTASE Ligands: FAD |
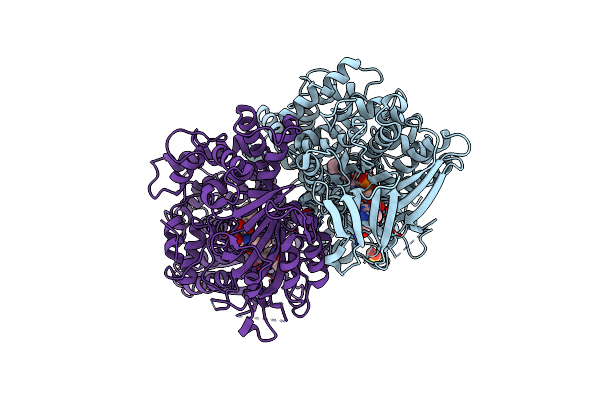 |
X-Ray Crystal Structures Of L-Phenylalanine Oxidase (Deaminating And Decaboxylating) From Pseudomonas Sp. P501. Structures Of The Enzyme-Ligand Complex And Catalytic Mechanism
Organism: Pseudomonas
Method: X-RAY DIFFRACTION Resolution:1.25 Å Release Date: 2011-08-31 Classification: OXIDOREDUCTASE Ligands: SO4, FAD, HCI, GOL |
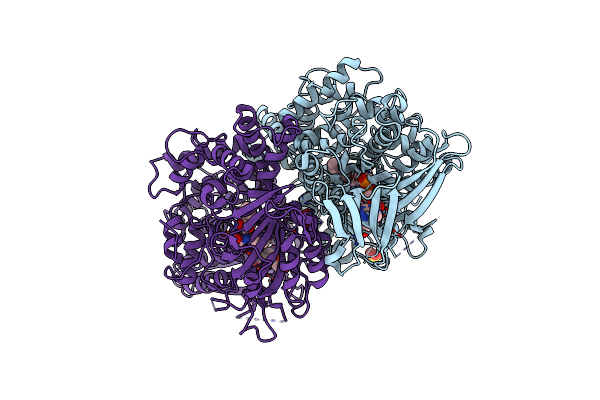 |
X-Ray Crystal Structures Of L-Phenylalanine Oxidase (Deaminating And Decaboxylating) From Pseudomonas Sp. P501. Structures Of The Enzyme-Ligand Complex And Catalytic Mechanism
Organism: Pseudomonas
Method: X-RAY DIFFRACTION Resolution:1.10 Å Release Date: 2011-08-31 Classification: OXIDOREDUCTASE Ligands: SO4, FAD, PHE, GOL |
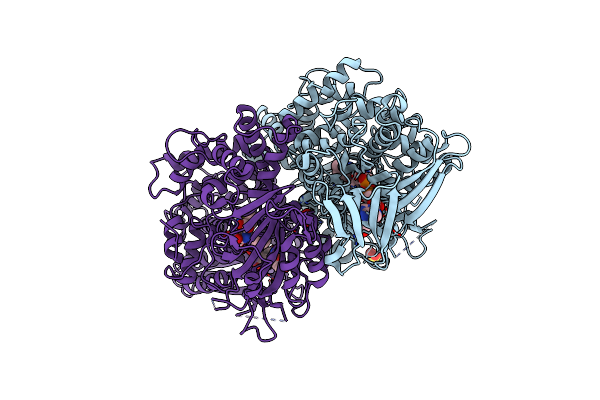 |
X-Ray Crystal Structures Of L-Phenylalanine Oxidase (Deaminating And Decaboxylating) From Pseudomonas Sp. P501. Structures Of The Enzyme-Ligand Complex And Catalytic Mechanism
Organism: Pseudomonas
Method: X-RAY DIFFRACTION Resolution:1.25 Å Release Date: 2011-08-31 Classification: OXIDOREDUCTASE Ligands: SO4, FAD, MET, GOL |
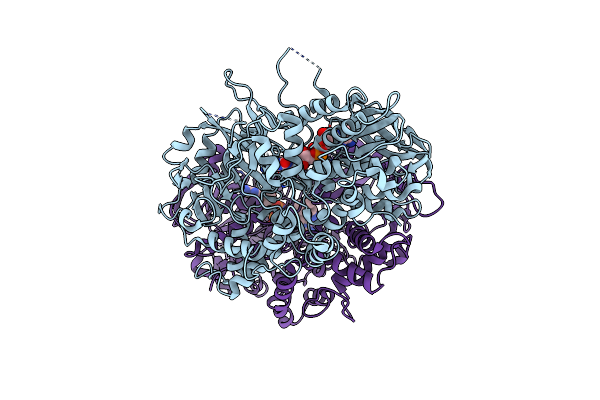 |
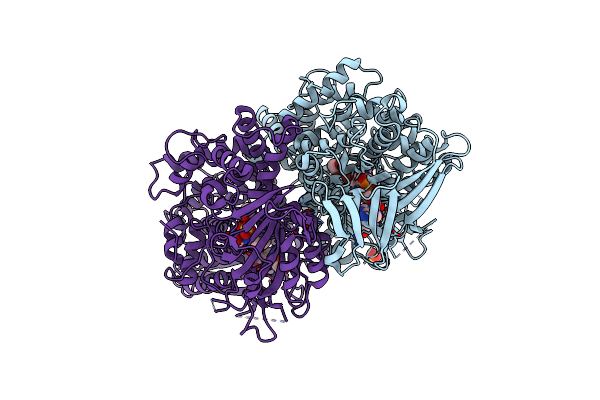
Organism: Pseudomonas sp. p-501
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2008-04-15 Classification: OXIDOREDUCTASE Ligands: FAD, SO4 |
 |
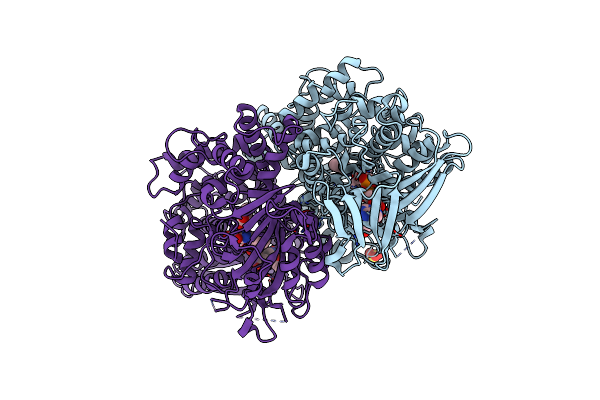
Organism: Pseudomonas sp. p-501
Method: X-RAY DIFFRACTION Resolution:1.25 Å Release Date: 2008-04-15 Classification: OXIDOREDUCTASE Ligands: SO4, FAD, GOL |
 |
Organism: Pseudomonas sp. p-501
Method: X-RAY DIFFRACTION Resolution:1.35 Å Release Date: 2008-04-15 Classification: OXIDOREDUCTASE Ligands: SO4, FAD, BE2, GOL |
 |
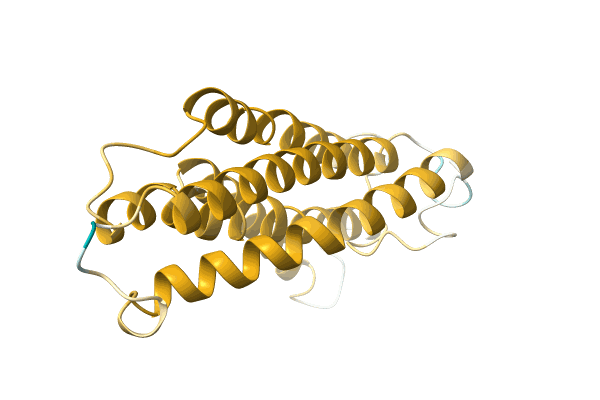 |
 |
 |
 |