Search Count: 3,086
All
Selected
 |
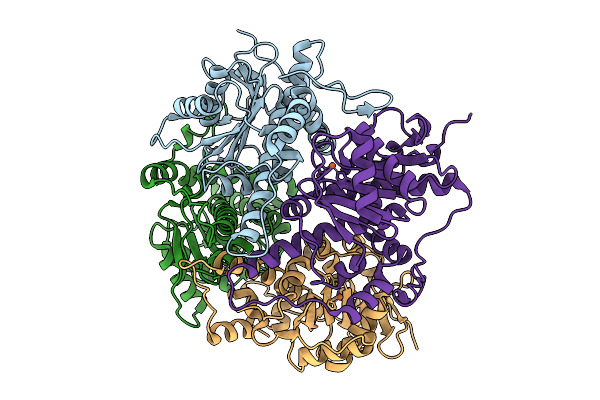
Crystal Structure Of None-Heme Iron Enzyme (Tqam) From Trichoderma Atroviride Bound With Iron And 2-Aminoisobutyric Acid
Organism: Trichoderma atroviride
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2026-02-04 Classification: METAL BINDING PROTEIN Ligands: FE, AIB |
 |
Crystal Structure Of None-Heme Iron Enzyme (Tqam) From Trichoderma Atroviride Bound With Iron
Organism: Trichoderma atroviride
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2026-01-28 Classification: METAL BINDING PROTEIN Ligands: FE |
 |
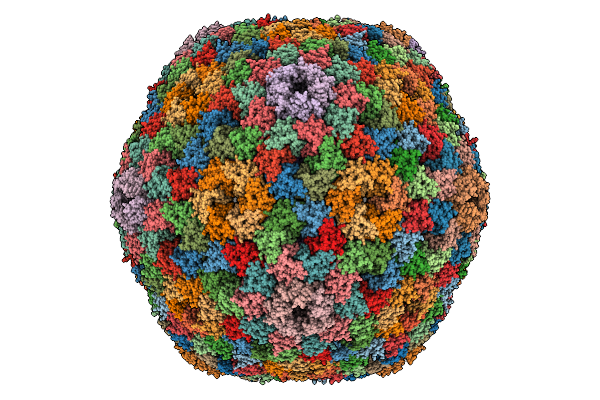
Organism: Zea mays
Method: X-RAY DIFFRACTION Resolution:2.08 Å Release Date: 2026-01-07 Classification: OXIDOREDUCTASE Ligands: FE |
 |
Organism: Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Release Date: 2025-12-31 Classification: STRUCTURAL PROTEIN |
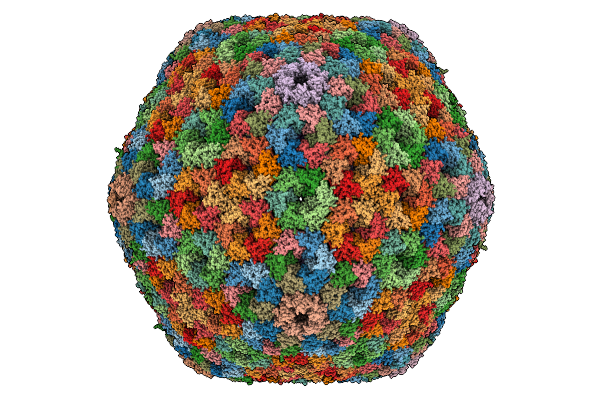 |
Cryo-Em Structure Of Carboxysomal Mid-Shell: T = 16 Shell Under C1 Symmetry.
Organism: Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Release Date: 2025-12-31 Classification: STRUCTURAL PROTEIN |
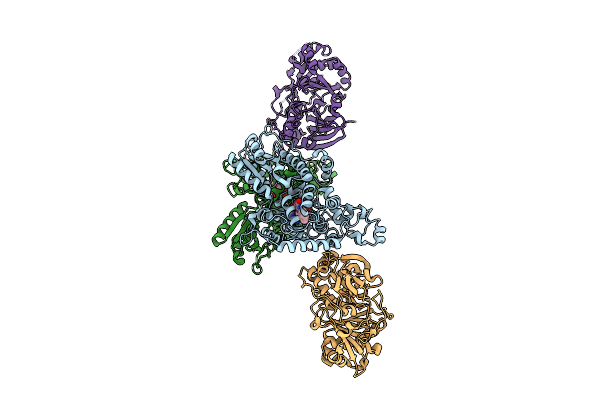 |
Organism: Zea mays
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2025-11-05 Classification: PLANT PROTEIN Ligands: FAD |
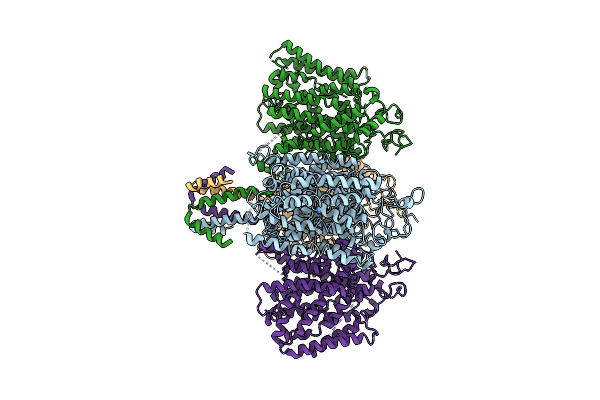 |
Organism: Zea mays
Method: ELECTRON MICROSCOPY Resolution:3.41 Å Release Date: 2025-11-05 Classification: CHOLINE-BINDING PROTEIN Ligands: URE |
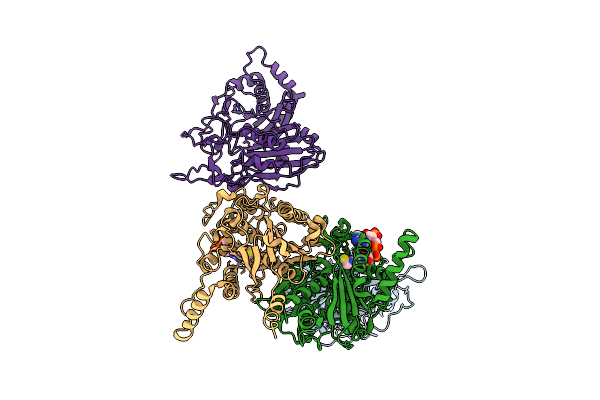 |
Organism: Zea mays
Method: ELECTRON MICROSCOPY Release Date: 2025-10-29 Classification: PLANT PROTEIN Ligands: COA |
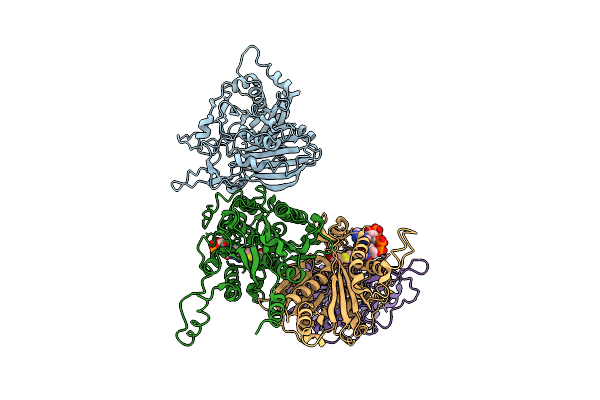 |
Cryo-Em Structure Of The Maize Cer6-Gl2 Complex (Inactive C222A Mutant) Bound With Malonyl-Coa
Organism: Zea mays
Method: ELECTRON MICROSCOPY Release Date: 2025-10-29 Classification: PLANT PROTEIN Ligands: MLC |
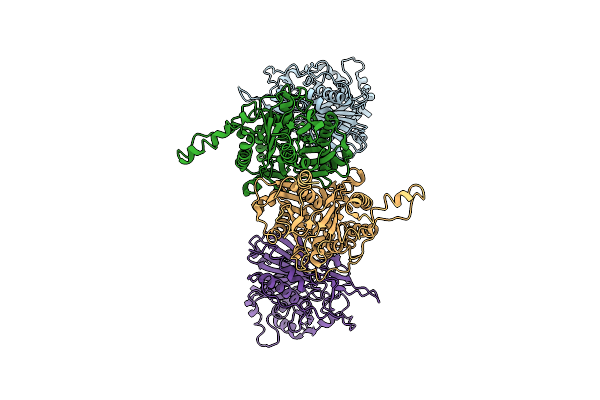 |
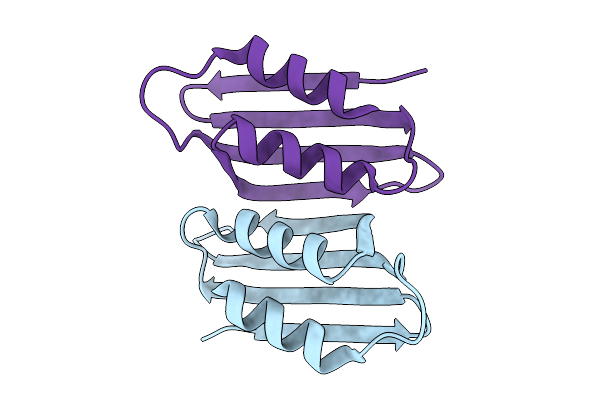
Cryo-Em Structure Of The Maize Cer6-Gl2 Complex In The Presence Of 30:0 Coa
Organism: Zea mays
Method: ELECTRON MICROSCOPY Release Date: 2025-10-29 Classification: PLANT PROTEIN |
 |
Organism: Zea mays
Method: X-RAY DIFFRACTION Resolution:2.29 Å Release Date: 2025-10-08 Classification: LIPID BINDING PROTEIN |
 |
Crystal Structure Of Beta-Lactamase-Like Domain Of Comec From Moorella Glycerini
Organism: Neomoorella glycerini
Method: X-RAY DIFFRACTION Resolution:1.86 Å Release Date: 2025-10-01 Classification: METAL BINDING PROTEIN Ligands: PO4 |
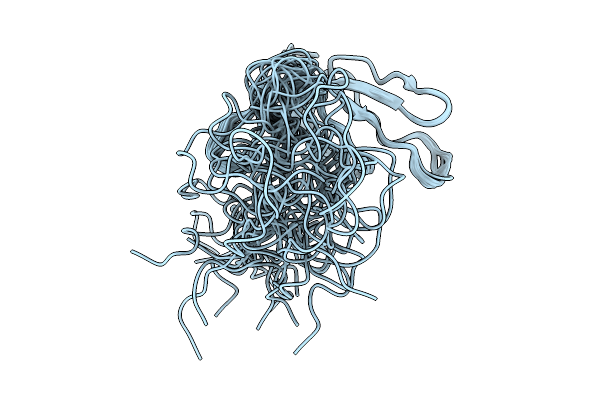 |
Organism: Moorella glycerini
Method: SOLUTION NMR Release Date: 2025-10-01 Classification: DNA BINDING PROTEIN |
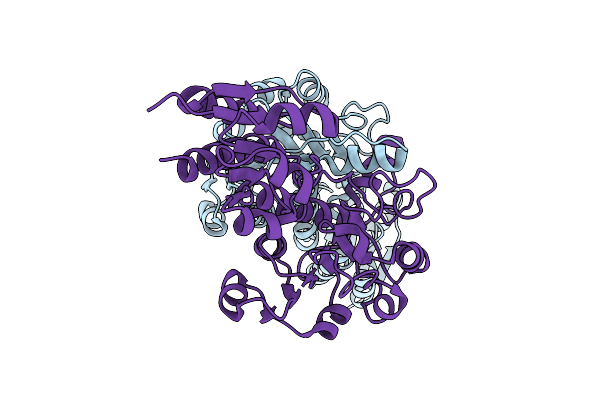 |
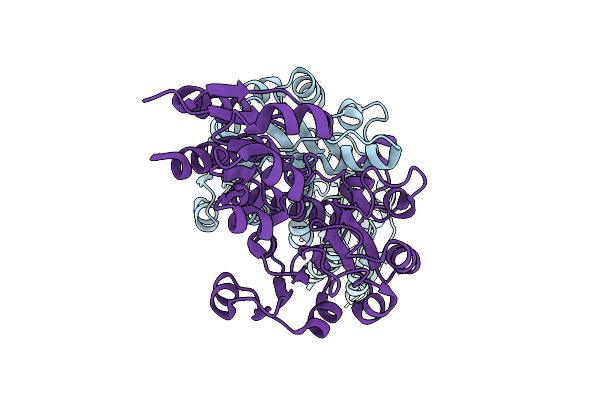
Organism: Zea mays
Method: X-RAY DIFFRACTION Resolution:3.49 Å Release Date: 2025-07-16 Classification: OXIDOREDUCTASE |
 |
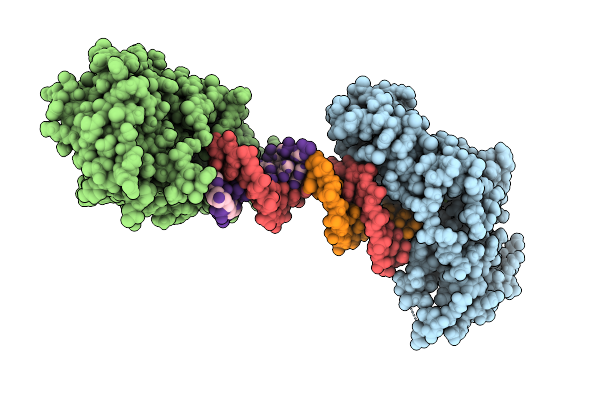
Crystal Structure Of Zea Mays 3-Phosphoglycerate Dehydrogenase S282L Mutant
Organism: Zea mays
Method: X-RAY DIFFRACTION Resolution:3.48 Å Release Date: 2025-07-16 Classification: OXIDOREDUCTASE |
 |
Organism: Zea mays, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.87 Å Release Date: 2025-07-16 Classification: PLANT PROTEIN/DNA Ligands: MG |
 |
Organism: Zea mays
Method: X-RAY DIFFRACTION Resolution:2.42 Å Release Date: 2025-07-16 Classification: PLANT PROTEIN Ligands: MN |
 |
Organism: Zea mays
Method: X-RAY DIFFRACTION Resolution:1.45 Å Release Date: 2025-07-02 Classification: OXIDOREDUCTASE Ligands: NAP, EDO |
 |
Crystal Structure Of Zmet2 In Complex With Unmethylated Ctg Dna And A Histone H3Kc9Me2 Peptide
Organism: Zea mays, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.71 Å Release Date: 2025-05-21 Classification: TRANSFERASE Ligands: SAH |
 |
Crystal Structure Of Zea Mays Adenosine Kinase 3 (Zmadk3) In Complex With Ap5A
Organism: Zea mays
Method: X-RAY DIFFRACTION Resolution:2.26 Å Release Date: 2025-01-29 Classification: TRANSFERASE Ligands: AP5, EDO, PEG, CL |

