Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
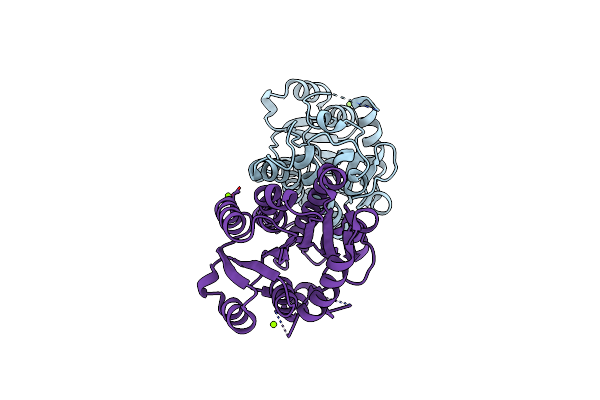 |
Discovery And Characterization Of A New Carbonyl Reductase From Rhodotorula Toluroides Reducing Fluoroketones, And X-Ray Analysis Of The Variant By Rational Engineering
Organism: Rhodotorula toruloides
Method: X-RAY DIFFRACTION Resolution:1.27 Å Release Date: 2024-02-07 Classification: OXIDOREDUCTASE Ligands: MG |
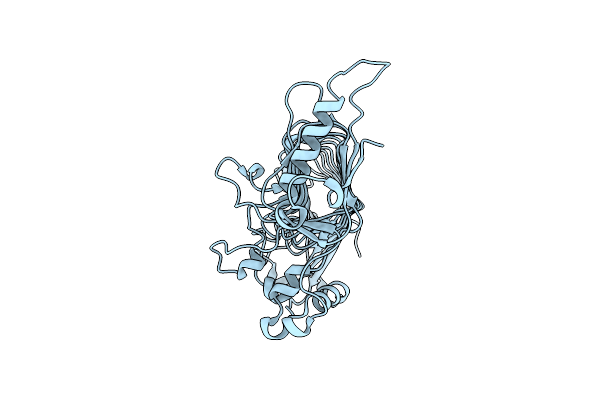 |
Organism: Arthrobacter chlorophenolicus a6
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2018-12-19 Classification: HYDROLASE |
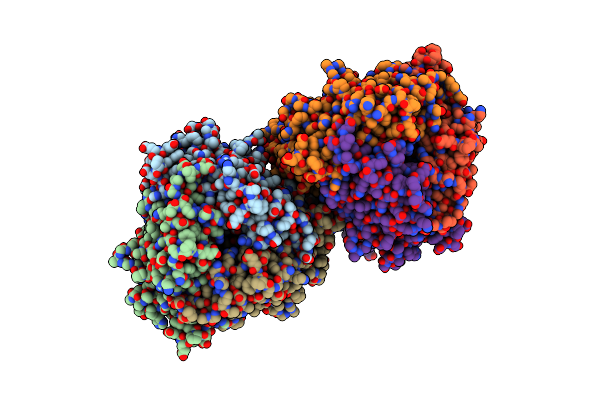 |
Crystal Structure Of Dfa-Iiiase From Arthrobacter Chlorophenolicus A6 In Complex With Dfa-Iii
Organism: Arthrobacter chlorophenolicus a6
Method: X-RAY DIFFRACTION Resolution:2.32 Å Release Date: 2018-12-19 Classification: HYDROLASE Ligands: 9F3 |
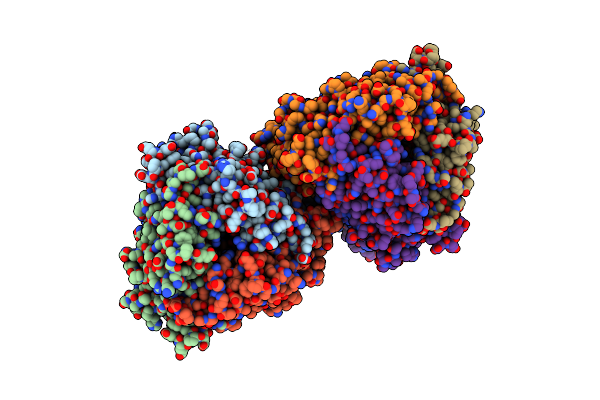 |
Crystal Structure Of Dfa-Iiiase From Arthrobacter Chlorophenolicus A6 In Complex With Gf2
Organism: Arthrobacter chlorophenolicus a6
Method: X-RAY DIFFRACTION Resolution:1.86 Å Release Date: 2018-12-19 Classification: HYDROLASE |
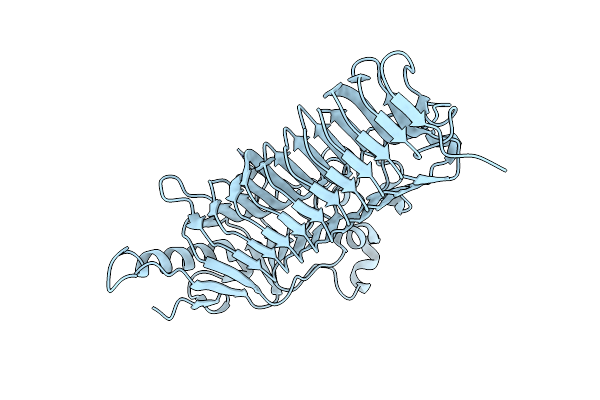 |
Crystal Structure Of Dfa-Iiiase From Arthrobacter Chlorophenolicus A6 Without Its Lid
Organism: Arthrobacter chlorophenolicus a6
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2018-12-19 Classification: HYDROLASE |
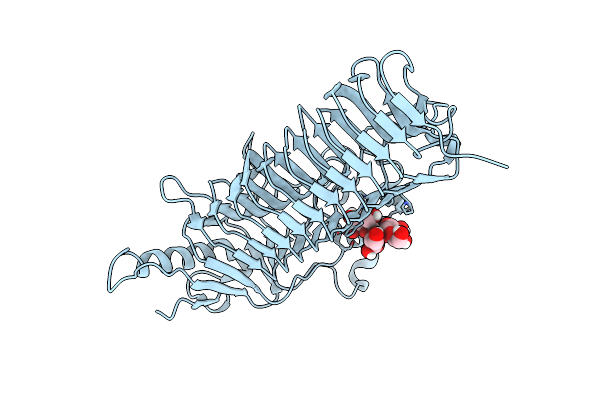 |
Crystal Structure Of Dfa-Iiiase From Arthrobacter Chlorophenolicus A6 Wihout Its Lid In Complex With Gf2
Organism: Arthrobacter chlorophenolicus a6
Method: X-RAY DIFFRACTION Resolution:3.00 Å Release Date: 2018-12-19 Classification: HYDROLASE |
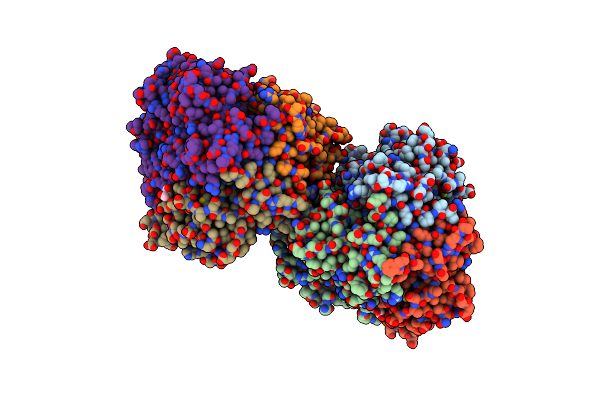 |
Crystal Structure Of Dfa-Iiiase Mutant C387A From Arthrobacter Chlorophenolicus A6
Organism: Arthrobacter chlorophenolicus a6
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2018-12-19 Classification: HYDROLASE Ligands: GOL |
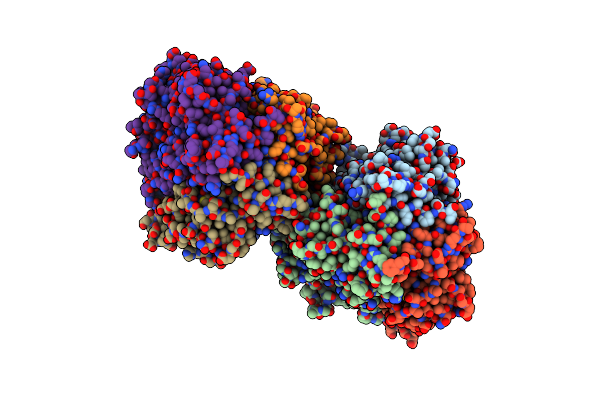 |
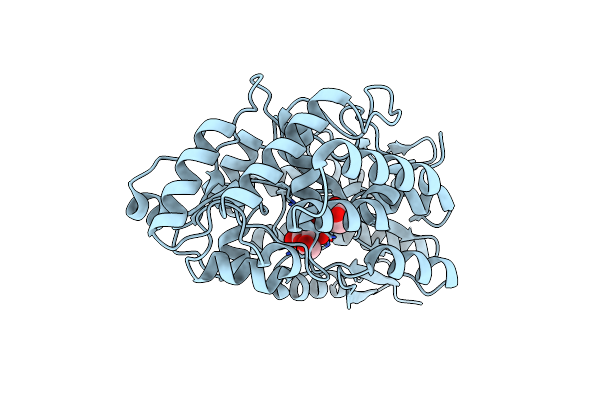
Crystal Structure Of Mutant C387A Of Dfa-Iiiase From Arthrobacter Chlorophenolicus A6 In Complex With Dfa-Iii
Organism: Arthrobacter chlorophenolicus a6
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2018-12-19 Classification: HYDROLASE Ligands: 9F3 |
 |
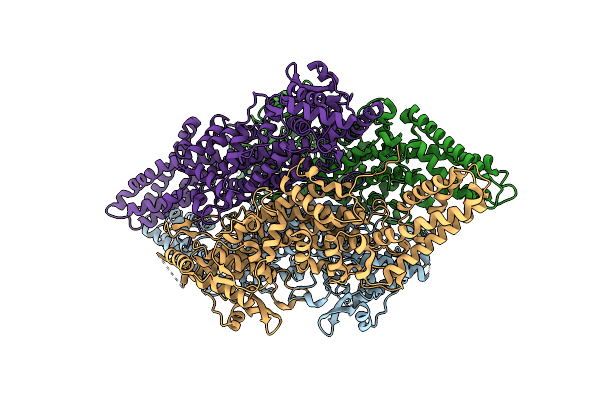
Crystal Structure Of Sugar Transporter Achl_0255 From Arthrobacter Chlorophenolicus A6, Target Efi-510633, With Bound Laminaribiose
Organism: Arthrobacter chlorophenolicus
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2014-08-27 Classification: TRANSPORT PROTEIN |
 |
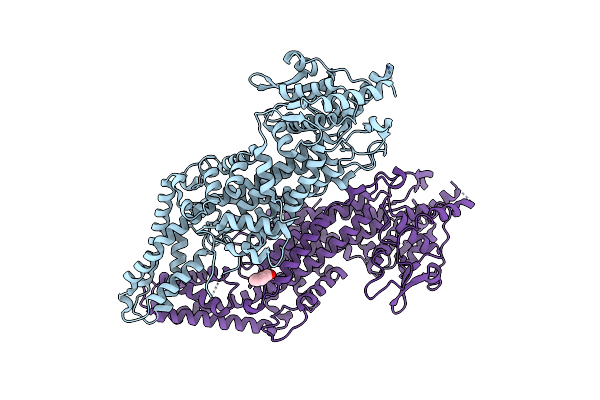
Crystal Structure Of Phenylalanine Ammonia-Lyase From Yeast Rhododporidium Toruloides
Organism: Rhodosporidium toruloides
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2005-11-01 Classification: LYASE |
 |
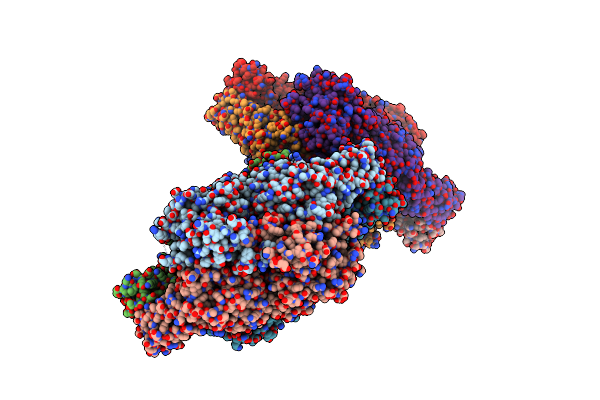
Crystal Structure Of Phenylalanine Ammonia Lyase From Rhodosporidium Toruloides
Organism: Rhodosporidium toruloides
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2004-10-12 Classification: LYASE Ligands: CIN |
 |
Crystal Structure Of Phenylalanine Ammonia Lyase From Rhodosporidium Toruloides
Organism: Rhodosporidium toruloides
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2004-10-12 Classification: LYASE |
 |
Crystal Structure Of D-Amino Acid Oxidase In Complex With Two Anthranylate Molecules
Organism: Rhodosporidium toruloides
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2002-02-27 Classification: OXIDOREDUCTASE Ligands: FAD, BE2 |
 |
Crystal Structure Analysis Of D-Amino Acid Oxidase In Complex With L-Lactate
Organism: Rhodosporidium toruloides
Method: X-RAY DIFFRACTION Resolution:1.46 Å Release Date: 2000-11-22 Classification: OXIDOREDUCTASE Ligands: FAD, LAC |
 |
D-Amino Acid Oxidase: Structure Of Substrate Complexes At Very High Resolution Reveal The Chemical Reacttion Mechanism Of Flavin Dehydrogenation
Organism: Rhodosporidium toruloides
Method: X-RAY DIFFRACTION Resolution:1.73 Å Release Date: 2000-11-22 Classification: OXIDOREDUCTASE Ligands: FLA, FAD |
 |
D-Amino Acic Oxidase In Complex With D-Alanine And A Partially Occupied Biatomic Species
Organism: Rhodosporidium toruloides
Method: X-RAY DIFFRACTION Resolution:1.20 Å Release Date: 2000-11-22 Classification: OXIDOREDUCTASE Ligands: FAD, DAL, PER, GOL |
 |
 |
 |
 |