Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
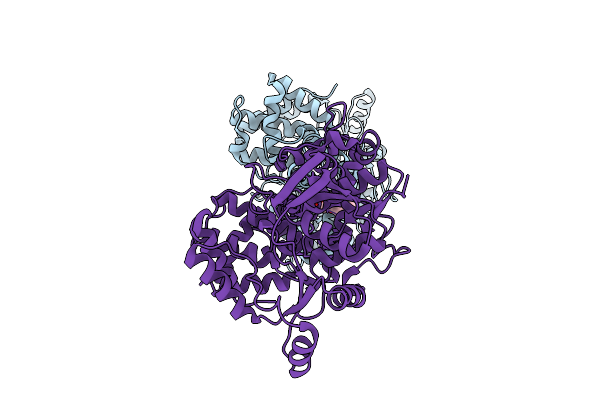 |
Crystal Structure Of Glutamate--Trna Ligase (Gltx) From Moraxella Catarrhalis (Apo)
Organism: Moraxella catarrhalis
Method: X-RAY DIFFRACTION Release Date: 2025-12-24 Classification: LIGASE Ligands: SO4, P6G, CL |
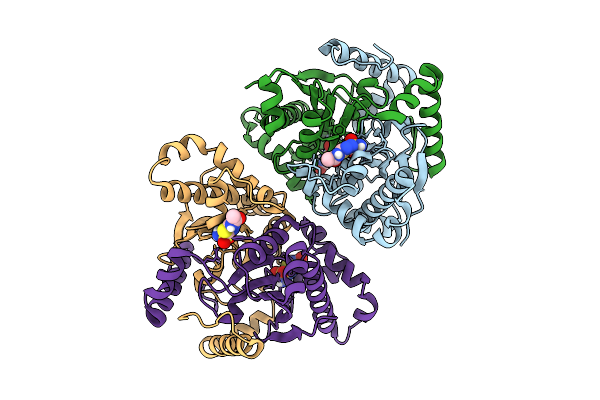 |
Organism: Candida parapsilosis
Method: X-RAY DIFFRACTION Release Date: 2025-11-26 Classification: LYASE Ligands: ZN, AZM |
 |
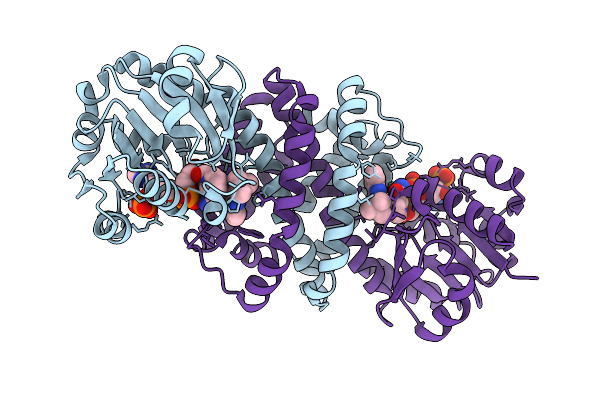
Crystal Structure Of Glutamate--Trna Ligase (Gltx) From Moraxella Catarrhalis In Complex With 5'-O-(N-Glutamyl)Sulfamoyladeonosine
Organism: Moraxella catarrhalis
Method: X-RAY DIFFRACTION Release Date: 2025-10-22 Classification: LIGASE Ligands: GSU, SO4, P6G, 2PE |
 |
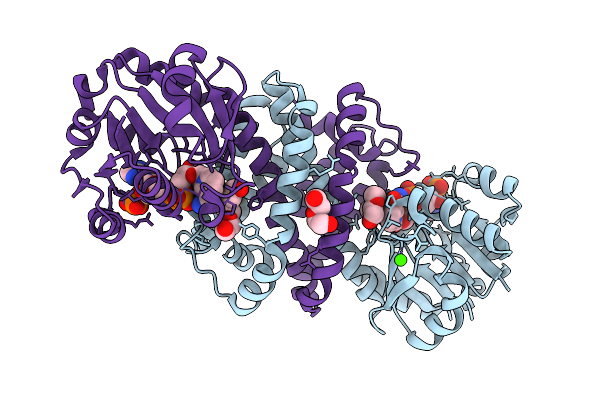
Organism: Amycolatopsis thermoflava
Method: X-RAY DIFFRACTION Release Date: 2025-07-23 Classification: OXIDOREDUCTASE Ligands: NAP, A1ECB |
 |
Organism: Amycolatopsis thermoflava
Method: X-RAY DIFFRACTION Release Date: 2025-07-23 Classification: OXIDOREDUCTASE Ligands: PGE, PEG, CL, CA, NDP |
 |
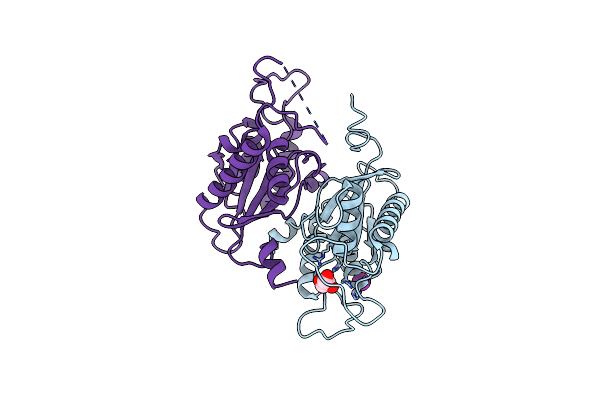
Organism: Acinetobacter baumannii ab0057
Method: X-RAY DIFFRACTION Resolution:3.40 Å Release Date: 2025-03-19 Classification: LIGASE Ligands: PGE, GOL, MG, EDO |
 |
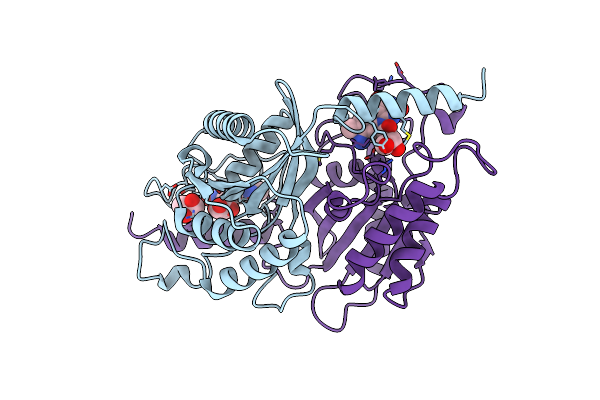
Organism: Streptomyces davaonensis
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2025-03-12 Classification: HYDROLASE Ligands: IOD, NA, GOL |
 |
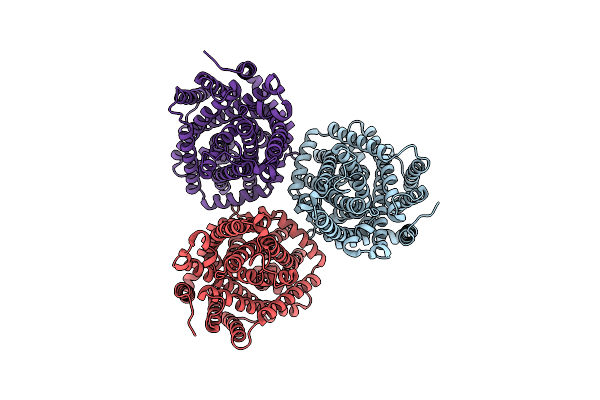
Organism: Streptomyces davaonensis
Method: X-RAY DIFFRACTION Resolution:1.25 Å Release Date: 2025-03-12 Classification: HYDROLASE Ligands: RS3, GOL |
 |
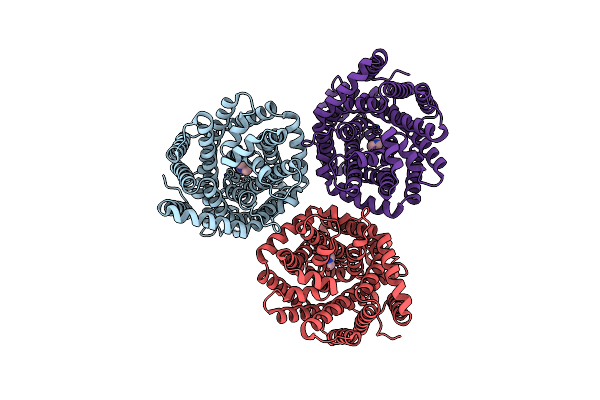
Organism: Myroides profundi
Method: ELECTRON MICROSCOPY Release Date: 2024-11-06 Classification: ELECTRON TRANSPORT |
 |
Organism: Myroides profundi
Method: ELECTRON MICROSCOPY Release Date: 2024-11-06 Classification: ELECTRON TRANSPORT Ligands: KEN |
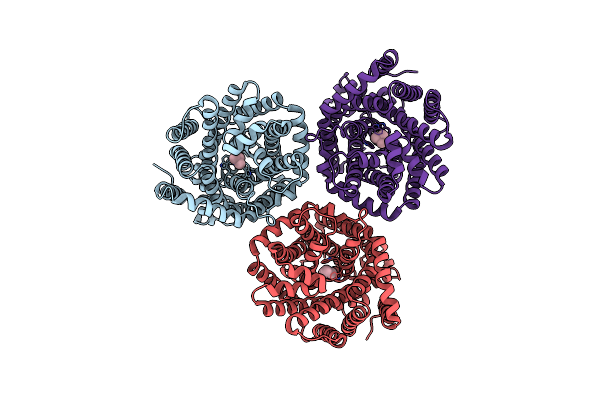 |
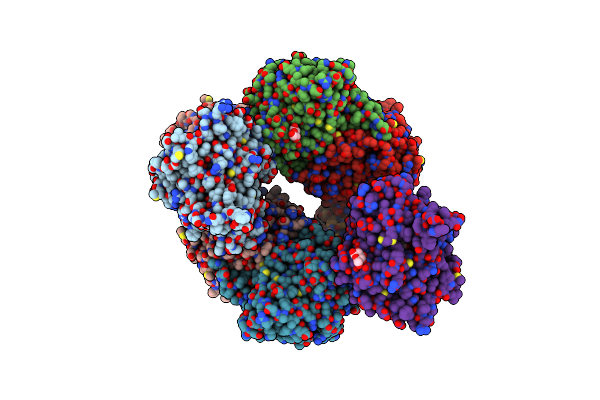
Organism: Myroides profundi
Method: ELECTRON MICROSCOPY Release Date: 2024-11-06 Classification: ELECTRON TRANSPORT Ligands: KEN |
 |
Organism: Sphingomonas sp. y57
Method: X-RAY DIFFRACTION Resolution:1.48 Å Release Date: 2024-11-06 Classification: HYDROLASE Ligands: ACT, BGC, ZN, K |
 |
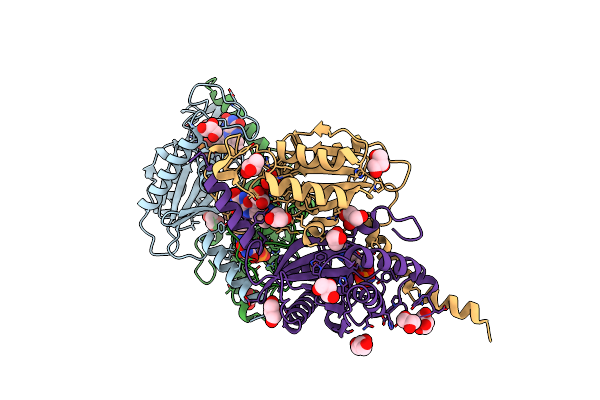
Crystal Structure Of Urethanase From Candida Parapsilosis And Structure-Based Engineering To Improve The Catalytic Activity And Stability
Organism: Candida parapsilosis
Method: X-RAY DIFFRACTION Resolution:2.66 Å Release Date: 2024-10-23 Classification: HYDROLASE |
 |
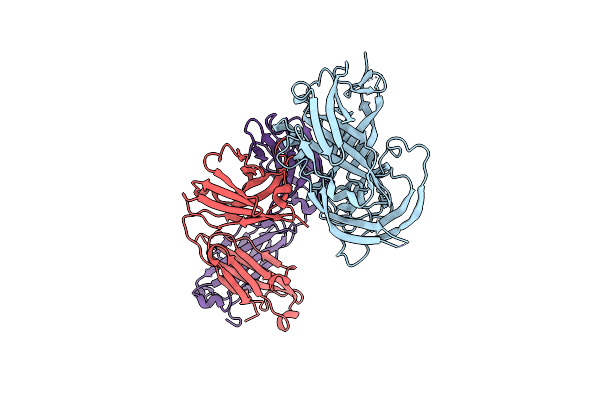
Organism: Streptomyces davaonensis
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2024-09-04 Classification: HYDROLASE Ligands: RS3, PO4, GOL |
 |
Organism: Orthohantavirus hantanense, Homo sapiens
Method: X-RAY DIFFRACTION Resolution:3.51 Å Release Date: 2024-07-24 Classification: VIRAL PROTEIN/IMMUNE SYSTEM |
 |
Crystal Structure Of Cis-Epoxysuccinic Hydrolase From Bradyrhizobium Mercantei
Organism: Bradyrhizobium mercantei
Method: X-RAY DIFFRACTION Release Date: 2024-06-12 Classification: HYDROLASE |
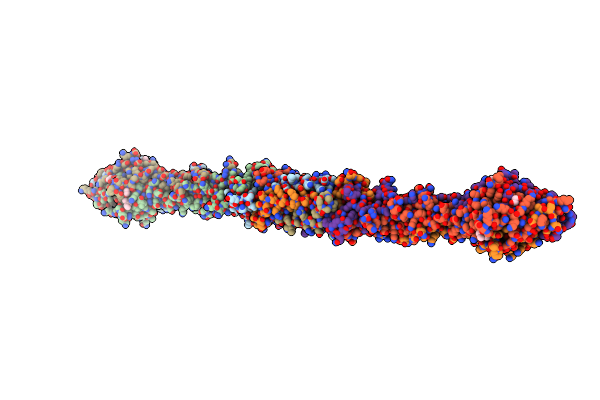 |
Structure Of The Tail Fibre From An Extracellular Contractile Injection System From Photorhabdus Bacteria
Organism: Photorhabdus asymbiotica subsp. asymbiotica atcc 43949
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2024-04-10 Classification: CELL ADHESION Ligands: MPD, EDO, UNX |
 |
Crystal Structure Of The A/Moscow/10/1999(H3N2) Influenza Virus Hemagglutinin With Human Receptor Analog 6'-Slnln
Organism: Influenza a virus
Method: X-RAY DIFFRACTION Resolution:2.56 Å Release Date: 2024-02-14 Classification: VIRAL PROTEIN Ligands: NAG |
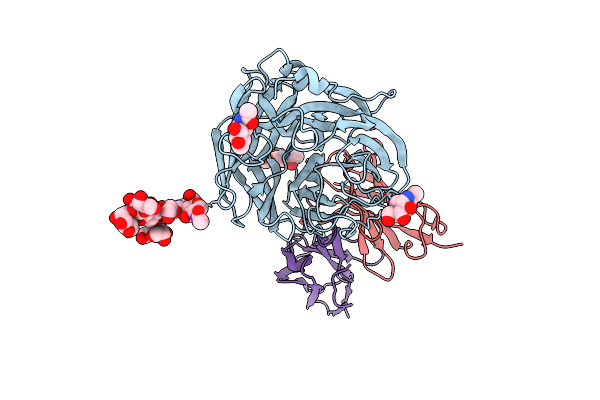 |
Structure Of 3A10 Fab In Complex With A/Moscow/10/1999 (H3N2) Influenza Virus Neuraminidase
Organism: Homo sapiens, Influenza a virus (a/moscow/10/1999(h3n2))
Method: ELECTRON MICROSCOPY Release Date: 2023-11-08 Classification: Hydrolase/Immune System Ligands: MAN, NAG |
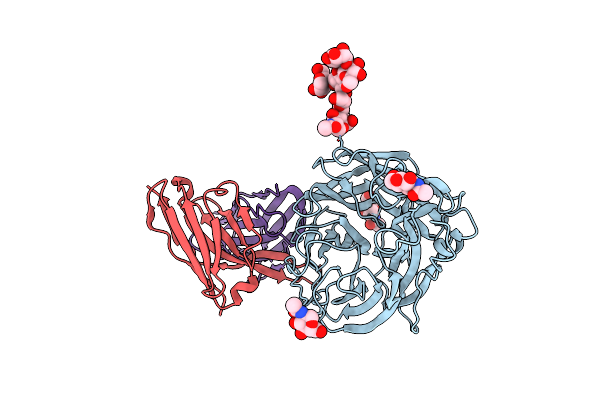 |
Structure Of 1F04 Fab In Complex With A/Moscow/10/1999 (H3N2) Influenza Virus Neuraminidase
Organism: Homo sapiens, Influenza a virus (a/moscow/10/1999(h3n2))
Method: ELECTRON MICROSCOPY Release Date: 2023-11-08 Classification: Hydrolase/Immune System Ligands: MAN, NAG |