Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
 |
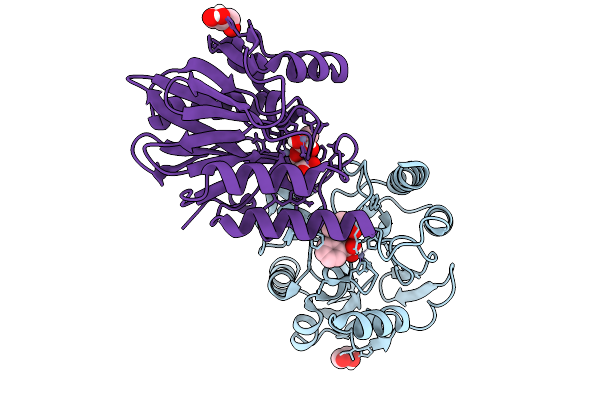
Hexamer Msp1 From S.Cerevisiae(With A Catalytic Dead Mutation) In Complex With An Unknown Peptide Substrate
Organism: Saccharomyces cerevisiae (strain atcc 204508 / s288c), Escherichia coli bl21(de3)
Method: ELECTRON MICROSCOPY Release Date: 2025-12-31 Classification: MEMBRANE PROTEIN Ligands: ATP, MG |
 |
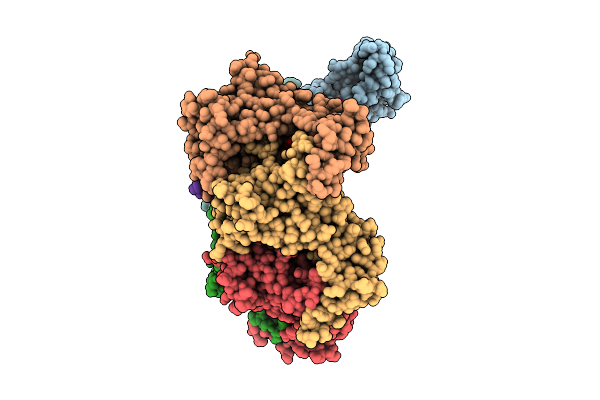
Organism: Escherichia coli bl21(de3)
Method: X-RAY DIFFRACTION Release Date: 2025-12-31 Classification: HYDROLASE Ligands: ZN, A1EJI, GOL |
 |
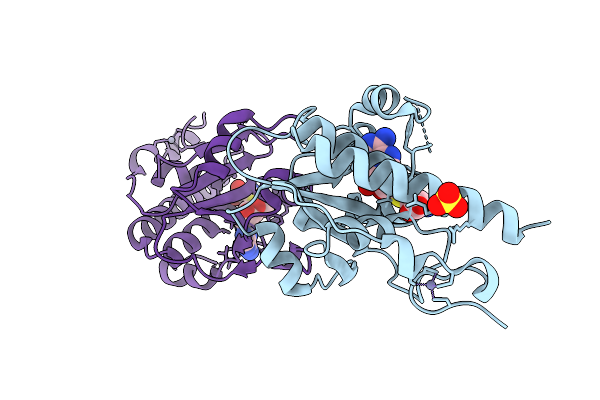
Hexamer Msp1 From S.Cerevisiae (With A Catalytic Dead Mutation) In Complex With An Unknown Peptide Substrate
Organism: Escherichia coli bl21(de3), Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2025-12-24 Classification: MEMBRANE PROTEIN Ligands: ATP, MG |
 |
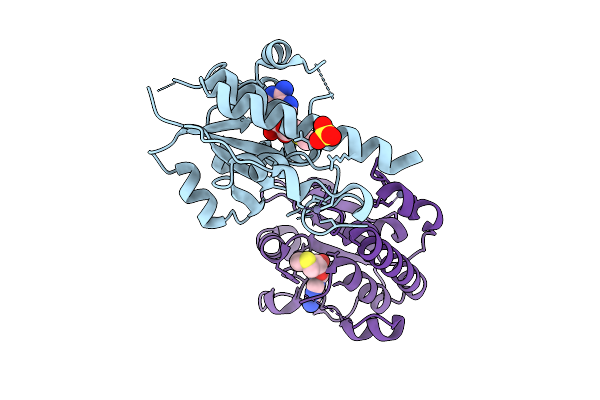
Organism: Escherichia coli (strain b / bl21-de3)
Method: X-RAY DIFFRACTION Release Date: 2025-12-17 Classification: TRANSFERASE Ligands: SAM, ZN, SO4 |
 |
Organism: Escherichia coli (strain b / bl21-de3)
Method: X-RAY DIFFRACTION Release Date: 2025-12-17 Classification: TRANSFERASE Ligands: MTA, ZN, SO4 |
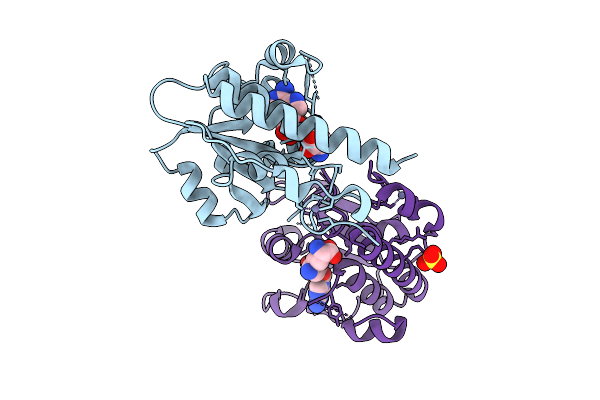 |
Organism: Escherichia coli bl21(de3)
Method: X-RAY DIFFRACTION Release Date: 2025-12-17 Classification: TRANSFERASE Ligands: SFG, ZN, SO4 |
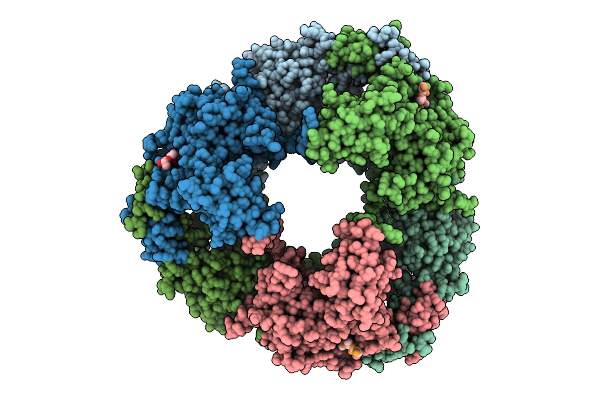 |
Cryo-Em Structure Of Ctpa S300A/K325A/Q329A Mutant From Helicobacter Pylori
Organism: Helicobacter pylori g27, Escherichia coli bl21(de3)
Method: ELECTRON MICROSCOPY Release Date: 2025-12-03 Classification: HYDROLASE |
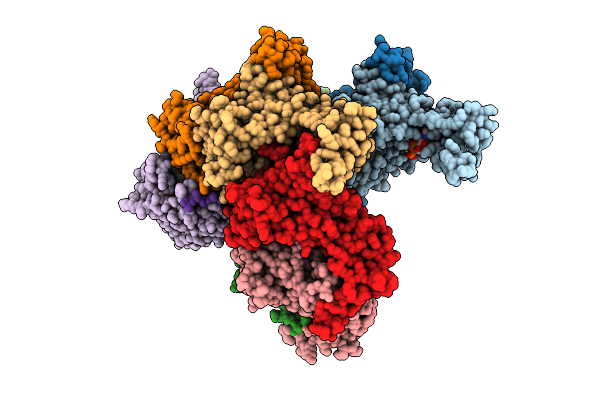 |
Nanomer Msp1 From S.Cerevisiae (With A Catalytic Dead Mutation) In Complex With An Unknown Peptide Substrate
Organism: Saccharomyces cerevisiae s288c, Escherichia coli bl21(de3)
Method: ELECTRON MICROSCOPY Release Date: 2025-11-26 Classification: MEMBRANE PROTEIN Ligands: ATP, MG |
 |
Organism: Escherichia coli bl21(de3)
Method: X-RAY DIFFRACTION Release Date: 2025-10-22 Classification: METAL BINDING PROTEIN Ligands: CO, CA, CL, NA |
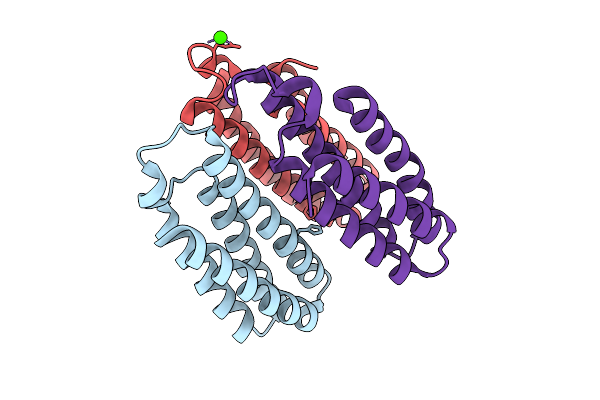 |
Organism: Escherichia coli bl21(de3)
Method: X-RAY DIFFRACTION Release Date: 2025-10-22 Classification: METAL BINDING PROTEIN Ligands: NI, CL, CA |
 |
Organism: Escherichia coli bl21(de3)
Method: X-RAY DIFFRACTION Release Date: 2025-10-22 Classification: METAL BINDING PROTEIN |
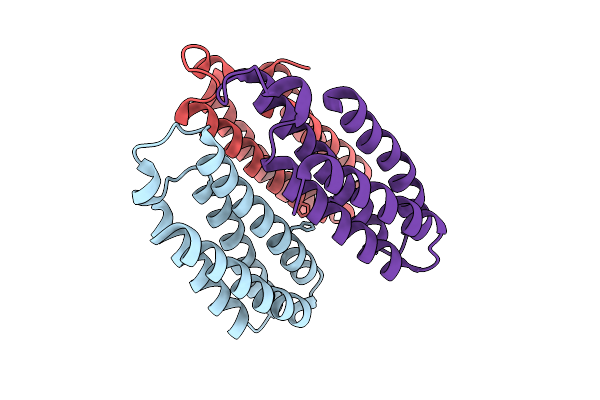 |
Organism: Escherichia coli bl21(de3)
Method: X-RAY DIFFRACTION Release Date: 2025-10-22 Classification: METAL BINDING PROTEIN Ligands: CU |
 |
Organism: Escherichia coli bl21(de3)
Method: X-RAY DIFFRACTION Release Date: 2025-10-22 Classification: METAL BINDING PROTEIN Ligands: ZN, CA, NA |
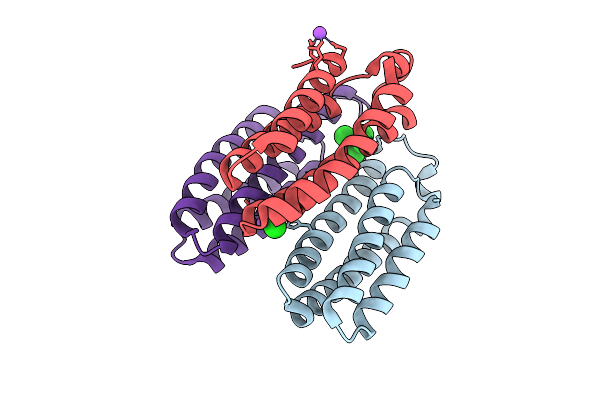 |
Organism: Escherichia coli bl21(de3)
Method: X-RAY DIFFRACTION Release Date: 2025-10-22 Classification: METAL BINDING PROTEIN Ligands: ZN, CL, BS3, NA |
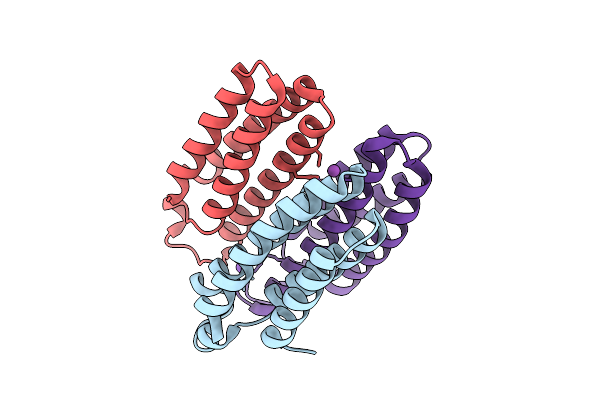 |
Organism: Escherichia coli bl21(de3)
Method: X-RAY DIFFRACTION Release Date: 2025-10-22 Classification: METAL BINDING PROTEIN Ligands: BS3, NA, CO |
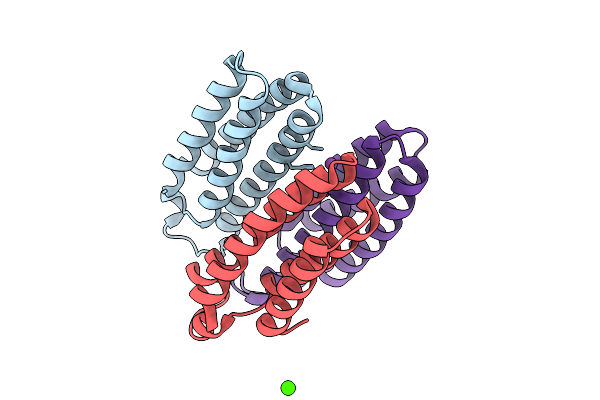 |
Organism: Escherichia coli bl21(de3)
Method: X-RAY DIFFRACTION Release Date: 2025-10-22 Classification: METAL BINDING PROTEIN Ligands: BS3, NI, CA |
 |
Organism: Escherichia coli bl21(de3)
Method: X-RAY DIFFRACTION Release Date: 2025-10-22 Classification: METAL BINDING PROTEIN Ligands: CL, NA, BS3 |
 |
Organism: Helicobacter pylori g27, Escherichia coli bl21(de3)
Method: X-RAY DIFFRACTION Release Date: 2025-10-01 Classification: HYDROLASE |
 |
Organism: Escherichia coli bl21(de3)
Method: X-RAY DIFFRACTION Release Date: 2025-08-27 Classification: ISOMERASE Ligands: QCF, PE8, SO4 |
 |
Organism: Escherichia coli bl21(de3)
Method: X-RAY DIFFRACTION Release Date: 2025-08-20 Classification: TRANSCRIPTION |