Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
 |
Organism: Teredinibacter turnerae (strain atcc 39867 / t7901)
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2019-10-09 Classification: OXIDOREDUCTASE |
 |
Organism: Teredinibacter turnerae t7901
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2019-03-20 Classification: HYDROLASE Ligands: 1PE, EDO, GOL, BR, PEG |
 |
Organism: Teredinibacter turnerae (strain atcc 39867 / t7901)
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2018-10-10 Classification: HYDROLASE Ligands: EDO, NA |
 |
Organism: Teredinibacter turnerae (strain atcc 39867 / t7901)
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2018-10-10 Classification: HYDROLASE Ligands: EDO |
 |
Organism: Teredinibacter turnerae (strain atcc 39867 / t7901)
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2018-10-10 Classification: HYDROLASE |
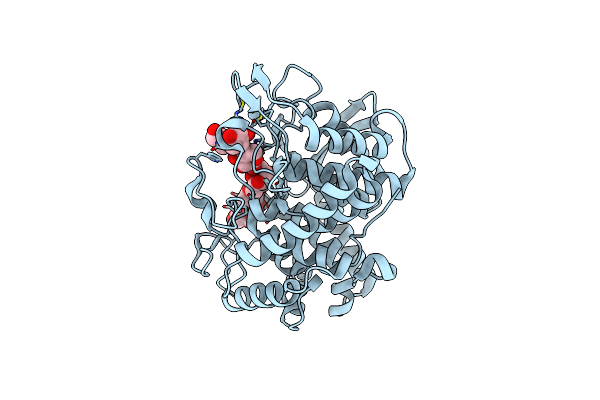 |
Crystal Structure Of A Gh8 Catalytic Mutant Xylohexaose Complex Xylanase From Teredinibacter Turnerae
Organism: Teredinibacter turnerae (strain atcc 39867 / t7901)
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2018-10-10 Classification: HYDROLASE Ligands: XYP, GOL |
 |
Crystal Structure Of Periplasmic Dipeptide Transport Protein From Yersinia Pestis
Organism: Yersinia pestis
Method: X-RAY DIFFRACTION Resolution:1.96 Å Release Date: 2015-12-23 Classification: HYDROLASE Ligands: EDO, GOL, PEG, CL, GLN |
 |
Crystal Structure Of Homoserine Kinase From Yersinia Pestis Nepal516, Nysgrc Target 032715
Organism: Yersinia pestis
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2014-11-12 Classification: TRANSFERASE Ligands: CIT |
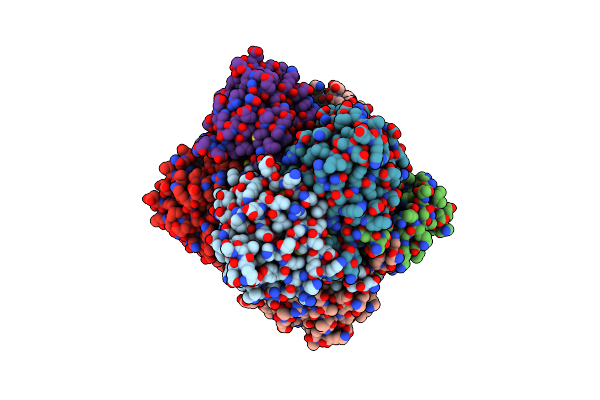 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:3.06 Å Release Date: 2009-09-01 Classification: HYDROLASE Ligands: MG, CL |
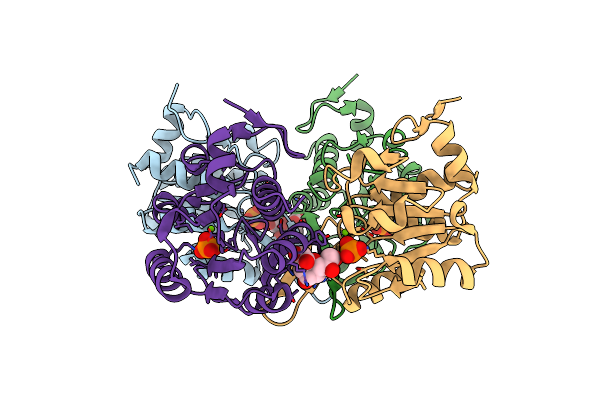 |
Crystal Structure Of Yrbi Lacking The Last 8 Residues, In Complex With Kdo And Inorganic Phosphate
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.49 Å Release Date: 2009-09-01 Classification: HYDROLASE Ligands: MG, PO4, KDO |
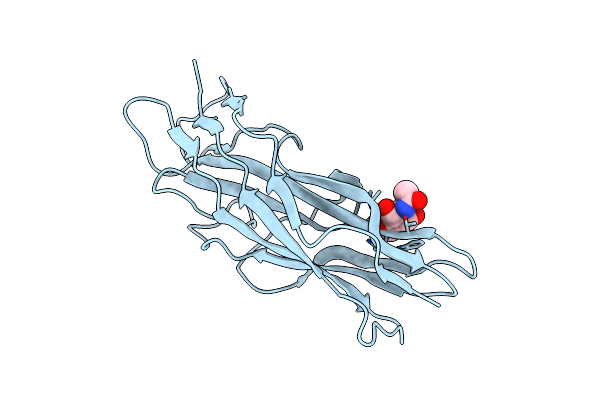 |
Organism: Escherichia coli b
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2005-08-19 Classification: SUGAR BINDING PROTEIN Ligands: NAG |
 |
 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2001-01-17 Classification: HYDROLASE Ligands: URA |
 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.06 Å Release Date: 2000-07-12 Classification: OXIDOREDUCTASE Ligands: FMN |
 |
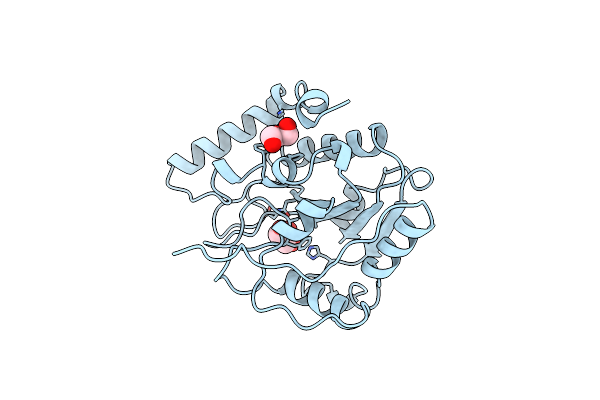
Crystal Structure Of Escherichia Coli Uracil Dna Glycosylase And Its Complexes With Uracil And Glycerol: Structure And Glycosylase Mechanism Revisited
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 1999-10-13 Classification: HYDROLASE Ligands: URA |
 |
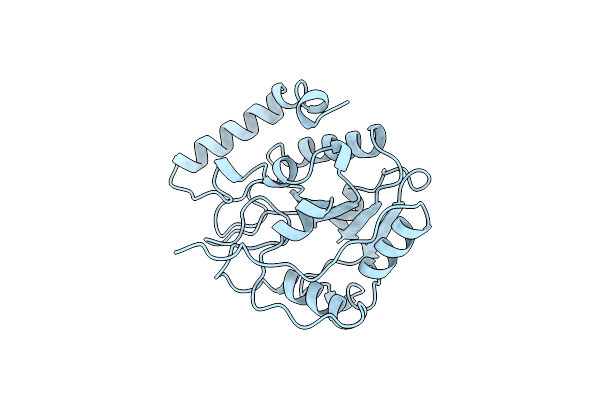
Crystal Structure Of Escherichia Coli Uracil Dna Glycosylase And Its Complexes With Uracil And Glycerol: Structure And Glycosylase Mechanism Revisited
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.43 Å Release Date: 1999-10-13 Classification: HYDROLASE Ligands: GOL |
 |
Crystal Structure Of Escherichia Coli Uracil Dna Glycosylase And Its Complexes With Uracil And Glycerol: Structure And Glycosylase Mechanism Revisited
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 1999-10-12 Classification: HYDROLASE |
 |
Crystallographic And Enzymatic Studies Of An Active Site Variant H187Q Of Escherichia Coli Uracil Dna Glycosylase: Crystal Structures Of Mutant H187Q And Its Uracil Complex
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 1999-07-23 Classification: HYDROLASE |
 |
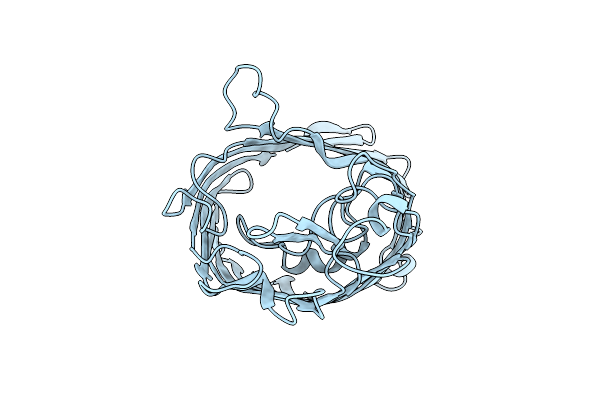
Crystallographic And Enzymatic Studies Of An Active Site Variant H187Q Of Escherichia Coli Uracil Dna Glycosylase: Crystal Structures Of Mutant H187Q And Its Uracil Complex
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 1999-07-23 Classification: HYDROLASE Ligands: URA |
 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:3.00 Å Release Date: 1999-01-13 Classification: MEMBRANE PROTEIN |