Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
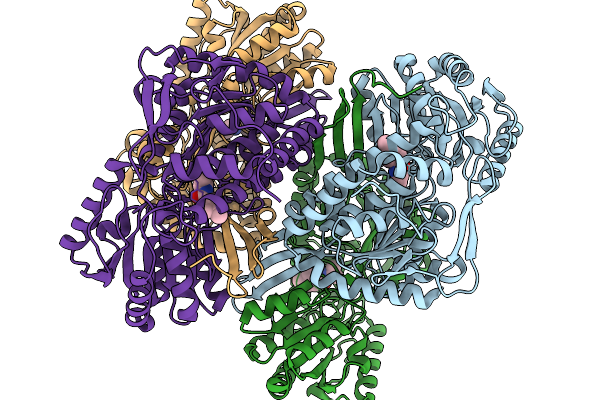 |
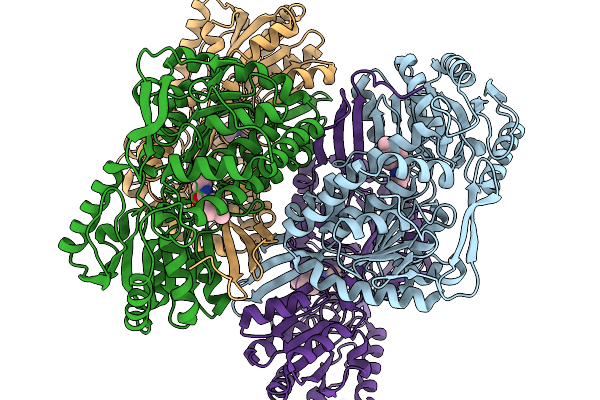
Co-Crystal Structure Of Phbdd-L163G With Product N-Isopropyl-4-Methylbenzamide
Organism: Pseudomonas putida
Method: X-RAY DIFFRACTION Release Date: 2026-01-14 Classification: OXIDOREDUCTASE Ligands: A1EBJ |
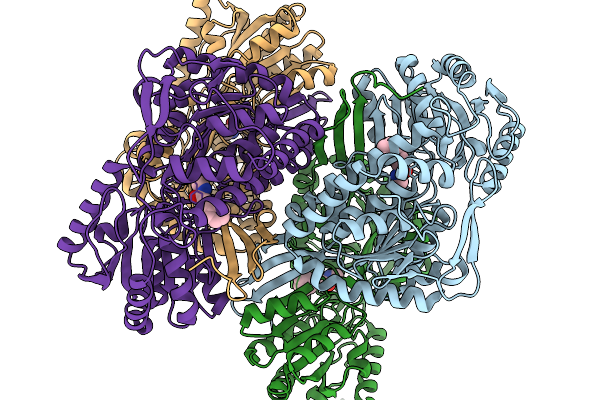 |
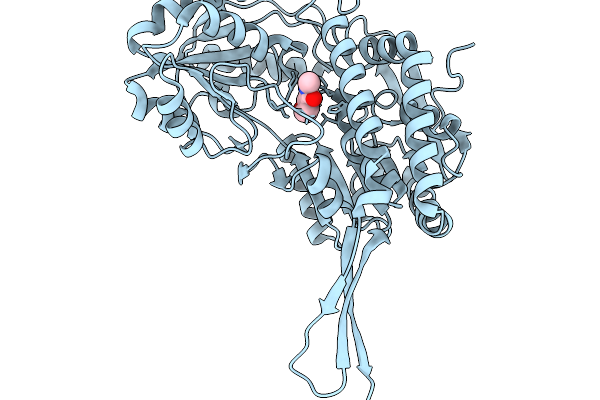
Organism: Pseudomonas putida
Method: X-RAY DIFFRACTION Release Date: 2026-01-14 Classification: OXIDOREDUCTASE Ligands: A1EBQ |
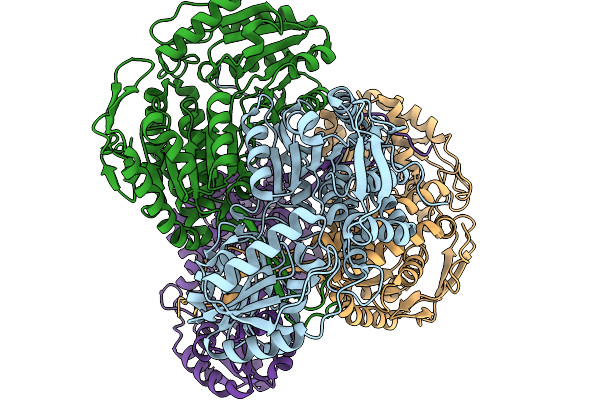 |
Organism: Pseudomonas putida
Method: X-RAY DIFFRACTION Release Date: 2026-01-14 Classification: OXIDOREDUCTASE |
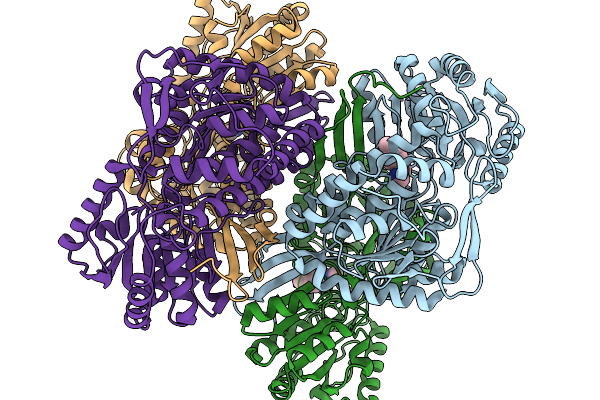 |
Organism: Pseudomonas putida
Method: X-RAY DIFFRACTION Release Date: 2026-01-14 Classification: OXIDOREDUCTASE Ligands: A1EC9 |
 |
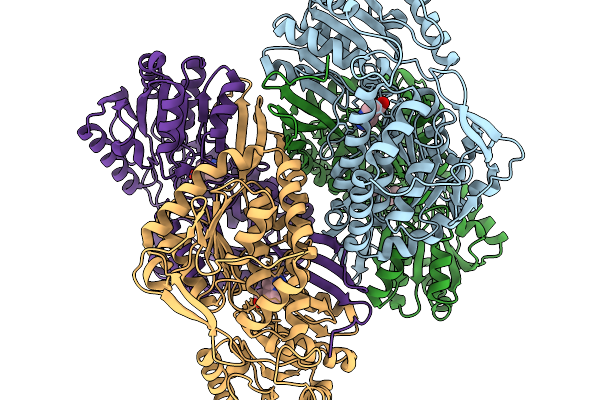
Co-Crystal Structure Of Phbdd-M1-L163G-T234A With N-Cyclopropyl-4-Methylbenzamide
Organism: Pseudomonas putida
Method: X-RAY DIFFRACTION Release Date: 2026-01-14 Classification: OXIDOREDUCTASE Ligands: A1EDB |
 |
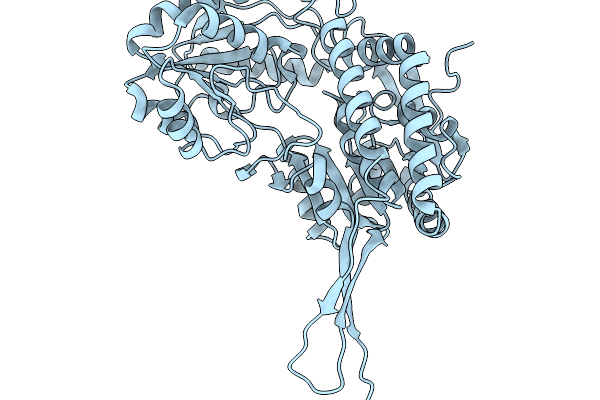
Organism: Pseudomonas putida
Method: X-RAY DIFFRACTION Release Date: 2026-01-14 Classification: OXIDOREDUCTASE Ligands: A1EDA, SO4 |
 |
Co-Crystal Structure Of Phbdd-M1 With 1H-Pyrrolo[2,3-B]Pyridine-4-Carbaldehyde
Organism: Pseudomonas putida
Method: X-RAY DIFFRACTION Release Date: 2026-01-14 Classification: OXIDOREDUCTASE Ligands: A1EDC |
 |
Organism: Pseudomonas putida
Method: X-RAY DIFFRACTION Release Date: 2026-01-14 Classification: OXIDOREDUCTASE |
 |
Organism: Pseudomonas putida
Method: X-RAY DIFFRACTION Release Date: 2026-01-14 Classification: ISOMERASE Ligands: A1CF8, EDO, MG |
 |
Organism: Pseudomonas putida
Method: X-RAY DIFFRACTION Release Date: 2026-01-07 Classification: OXIDOREDUCTASE Ligands: HEM, ACT |
 |
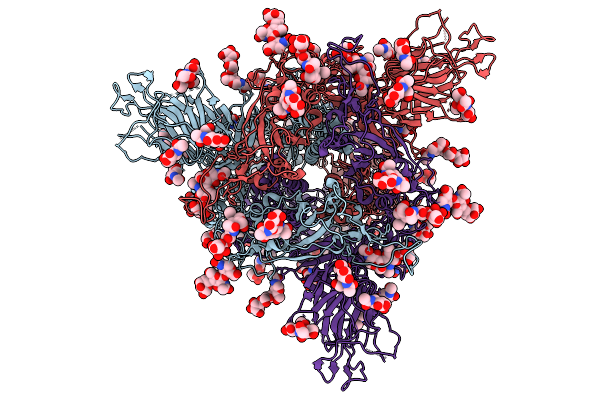
Organism: Homo sapiens, Human metapneumovirus a
Method: ELECTRON MICROSCOPY Release Date: 2025-12-31 Classification: VIRAL PROTEIN |
 |
Organism: Sarbecovirus sp. rhgb07
Method: ELECTRON MICROSCOPY Release Date: 2025-09-24 Classification: VIRAL PROTEIN Ligands: NAG |
 |
Crystal Structure Of The Dyp-Type Peroxidase Pross Variant From Pseudomonas Putida
Organism: Pseudomonas putida
Method: X-RAY DIFFRACTION Release Date: 2025-04-30 Classification: OXIDOREDUCTASE Ligands: OXY, HEM, CL |
 |
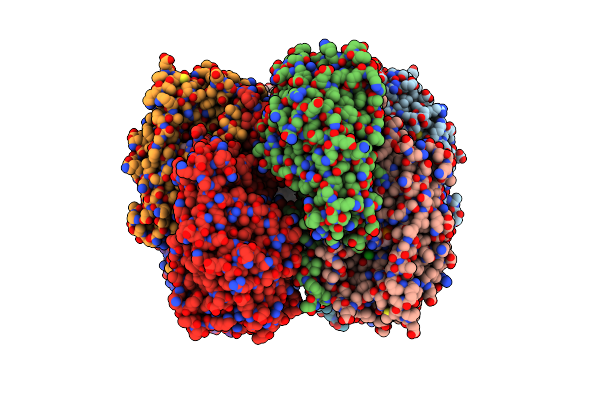
Crystal Structure Of A Dyp-Type Peroxidase Fireprot Variant From Pseudomonas Putida
Organism: Pseudomonas putida
Method: X-RAY DIFFRACTION Release Date: 2025-04-30 Classification: OXIDOREDUCTASE Ligands: GOL, HEM, CL |
 |
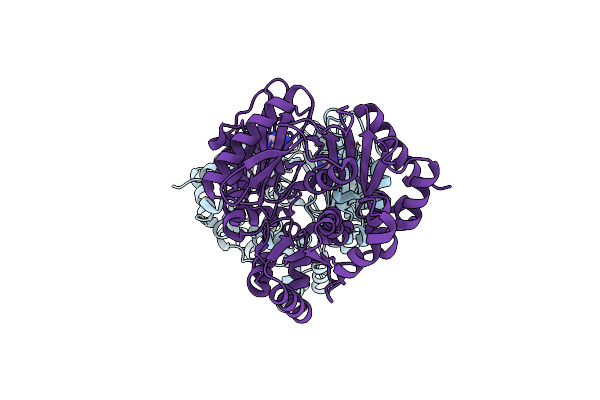
Organism: Penicillium steckii
Method: X-RAY DIFFRACTION Resolution:2.29 Å Release Date: 2025-03-05 Classification: OXIDOREDUCTASE Ligands: FMN, GOL, CL, CA |
 |
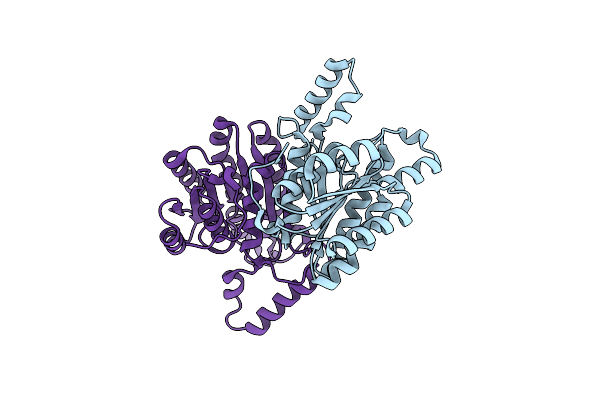
Organism: Streptantibioticus cattleyicolor
Method: X-RAY DIFFRACTION Release Date: 2025-03-05 Classification: LIGASE Ligands: ADE |
 |
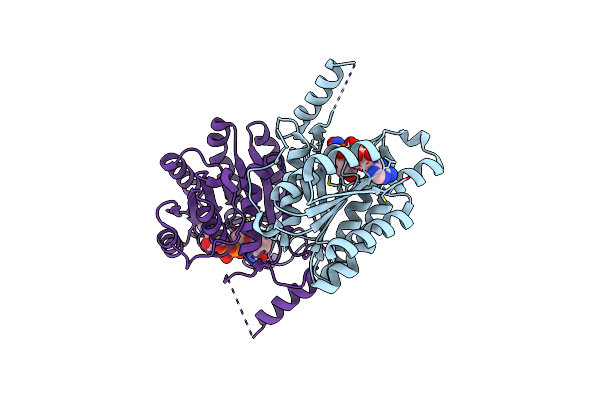
Crystal Structure Of Sulfoquinovose-1-Dehydrogenase From Pseudomonas Putida (Sulfo-Ed Pathway)
Organism: Pseudomonas putida
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2025-02-12 Classification: OXIDOREDUCTASE |
 |
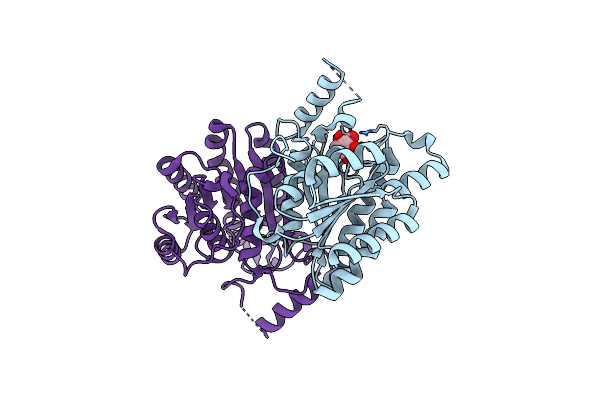
Crystal Structure Of Sulfoquinovose-1-Dehydrogenase From Pseudomonas Putida In Complex With Nad+ (Sulfo-Ed Pathway)
Organism: Pseudomonas putida
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2025-02-12 Classification: OXIDOREDUCTASE Ligands: NAD |
 |
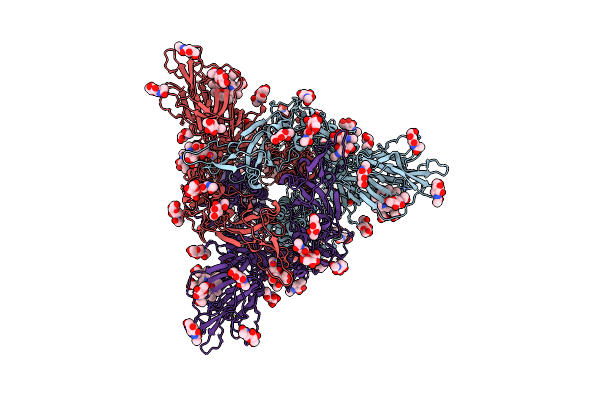
Crystal Structure Of Sulfoquinovose-1-Dehydrogenase From Pseudomonas Putida In Complex With Sulfoquinovose Substrate (Sulfo-Ed Pathway)
Organism: Pseudomonas putida
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2025-02-12 Classification: OXIDOREDUCTASE Ligands: YZT |
 |
Organism: Btrs-betacov/yn2013
Method: ELECTRON MICROSCOPY Release Date: 2025-02-05 Classification: VIRAL PROTEIN Ligands: NAG, BLR |