Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
 |
Organism: Thermoproteus sp. az2
Method: X-RAY DIFFRACTION Release Date: 2026-01-14 Classification: HYDROLASE Ligands: BGC, GOL, ACT |
 |
Organism: Thermoproteus sp. az2
Method: X-RAY DIFFRACTION Release Date: 2026-01-14 Classification: HYDROLASE |
 |
Organism: Thermoproteus sp. az2
Method: X-RAY DIFFRACTION Release Date: 2026-01-14 Classification: HYDROLASE Ligands: GOL, ACY, ACT |
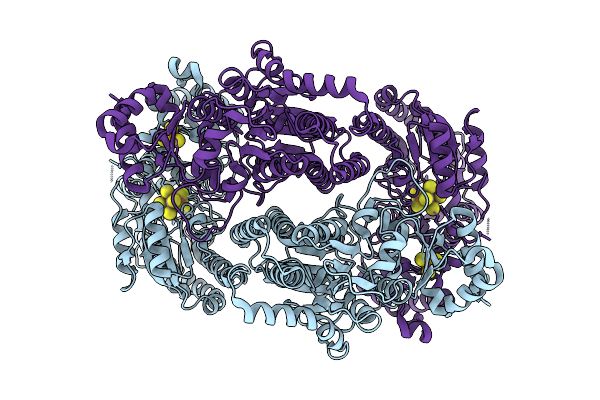 |
Organism: Geobacter metallireducens
Method: ELECTRON MICROSCOPY Release Date: 2025-11-05 Classification: METAL BINDING PROTEIN Ligands: SF4, S5Q |
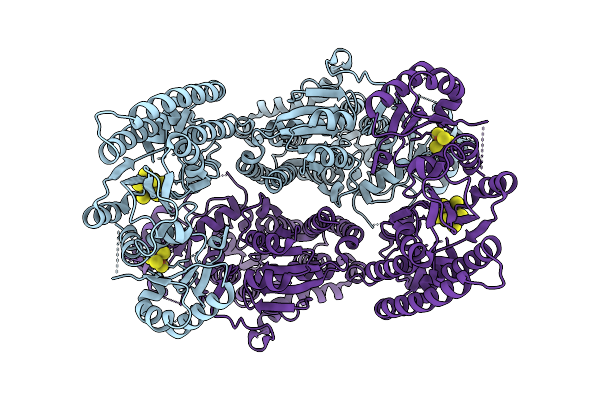 |
Organism: Geobacter metallireducens
Method: ELECTRON MICROSCOPY Release Date: 2025-11-05 Classification: METAL BINDING PROTEIN Ligands: SF4, S5Q |
 |
X-Ray Structure Of Cytochrome P450 Olet From Lacicoccus Alkaliphilus In Complex With Icosanoic Acid
Organism: Lacicoccus alkaliphilus dsm 16010
Method: X-RAY DIFFRACTION Release Date: 2025-08-06 Classification: OXIDOREDUCTASE Ligands: HEM, DCR, GOL |
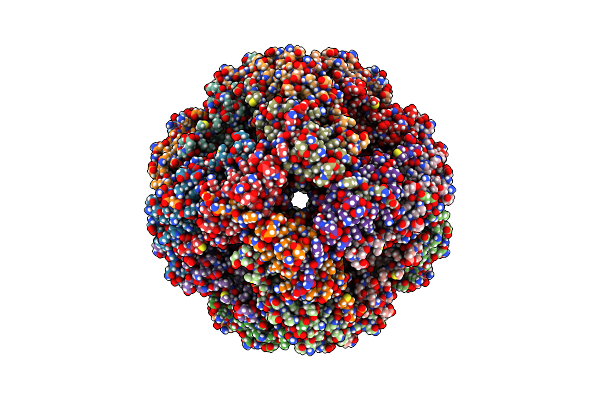 |
Organism: Methanocaldococcus jannaschii dsm 2661
Method: ELECTRON MICROSCOPY Resolution:2.50 Å Release Date: 2025-04-16 Classification: CHAPERONE |
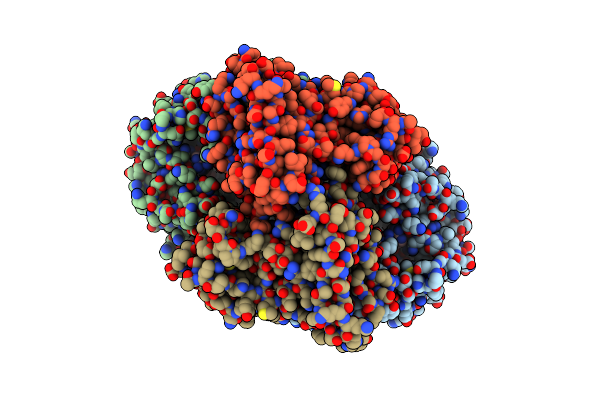 |
Organism: Novosphingopyxis baekryungensis dsm 16222
Method: ELECTRON MICROSCOPY Release Date: 2025-04-02 Classification: RNA BINDING PROTEIN/RNA Ligands: MG |
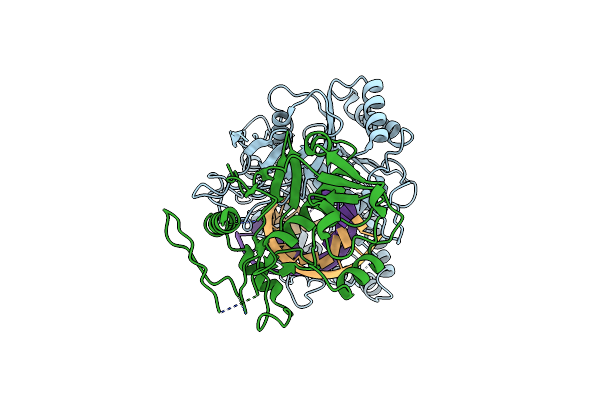 |
Organism: Novosphingopyxis baekryungensis dsm 16222
Method: ELECTRON MICROSCOPY Release Date: 2025-04-02 Classification: RNA BINDING PROTEIN/RNA/DNA Ligands: MG |
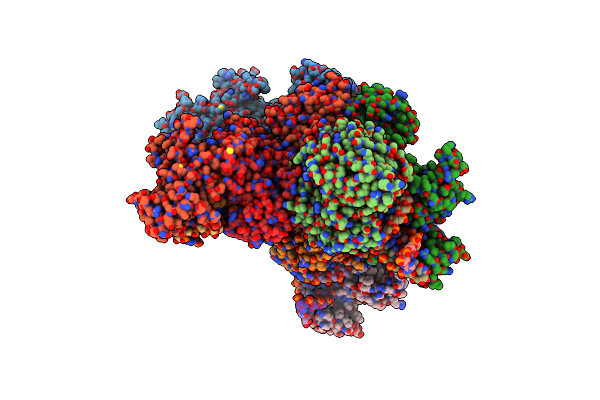 |
Organism: Novosphingopyxis baekryungensis dsm 16222
Method: ELECTRON MICROSCOPY Release Date: 2025-04-02 Classification: RNA BINDING PROTEIN/RNA/DNA Ligands: MG |
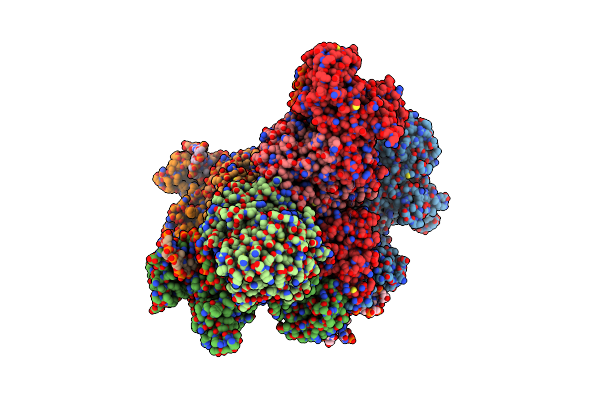 |
Organism: Novosphingopyxis baekryungensis dsm 16222
Method: ELECTRON MICROSCOPY Release Date: 2025-04-02 Classification: RNA BINDING PROTEIN/RNA/DNA Ligands: MG |
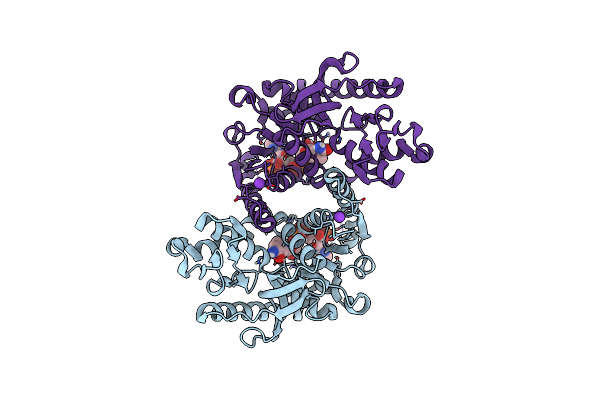 |
Crystal Structure Of Methanopyrus Kandleri Malate Dehydrogenase Mutant 4 At Room Temperature
Organism: Methanopyrus kandleri
Method: X-RAY DIFFRACTION Resolution:2.39 Å Release Date: 2025-03-19 Classification: OXIDOREDUCTASE Ligands: NDP, CL, K |
 |
Crystal Structure Of P. Syringae Phosphinothricin Acetyltransferase Pspto_3321
Organism: Pseudomonas syringae pv. tomato str. dc3000
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2025-03-12 Classification: TRANSFERASE Ligands: CIT, PEG, NA, PGE |
 |
Crystal Structure Of P. Syringae Phosphinothricin Acetyltransferase Pspto_3321 In Complex With L-Phosphinothricin
Organism: Pseudomonas syringae pv. tomato str. dc3000
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2025-03-12 Classification: TRANSFERASE Ligands: SO4, PPQ, ACT, NA, EDO, PEG |
 |
Organism: Pseudomonas syringae pv. tomato str. dc3000
Method: ELECTRON MICROSCOPY Release Date: 2024-08-21 Classification: PLANT PROTEIN Ligands: LBV |
 |
Organism: Pseudomonas syringae pv. tomato str. dc3000
Method: ELECTRON MICROSCOPY Release Date: 2024-08-21 Classification: PLANT PROTEIN Ligands: LBV |
 |
Organism: Pseudomonas syringae pv. tomato str. dc3000
Method: ELECTRON MICROSCOPY Release Date: 2024-08-21 Classification: PLANT PROTEIN Ligands: LBV |
 |
Organism: Pseudomonas syringae pv. tomato str. dc3000
Method: ELECTRON MICROSCOPY Release Date: 2024-08-21 Classification: PLANT PROTEIN Ligands: LBV |
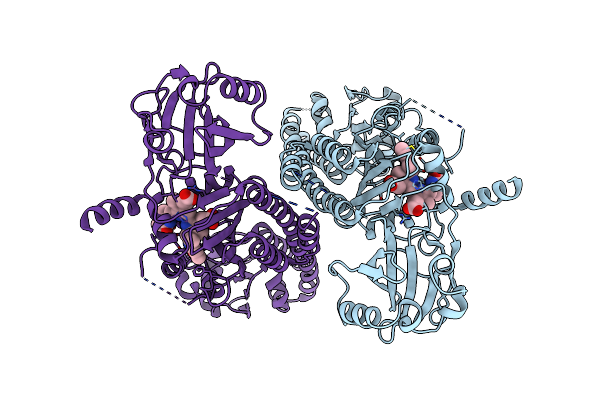 |
Organism: Pseudomonas syringae pv. tomato str. dc3000
Method: ELECTRON MICROSCOPY Release Date: 2024-08-21 Classification: PLANT PROTEIN Ligands: LBV |
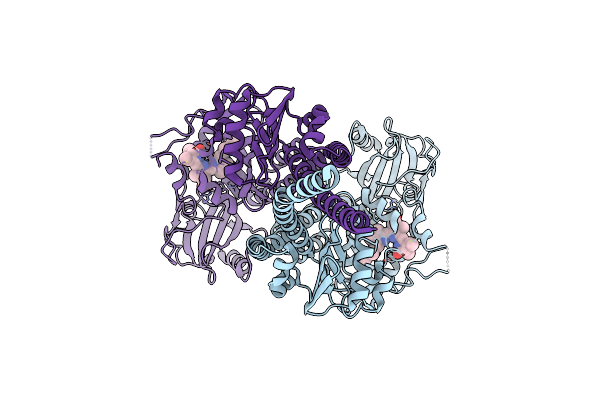 |
Organism: Pseudomonas syringae pv. tomato str. dc3000
Method: ELECTRON MICROSCOPY Release Date: 2024-08-21 Classification: PLANT PROTEIN Ligands: LBV |