Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
 |
Organism: Ruminococcus sp. xpd3002, Cloning vector pgrna-ccdb-1
Method: ELECTRON MICROSCOPY Release Date: 2025-10-08 Classification: RNA BINDING PROTEIN/RNA Ligands: MG |
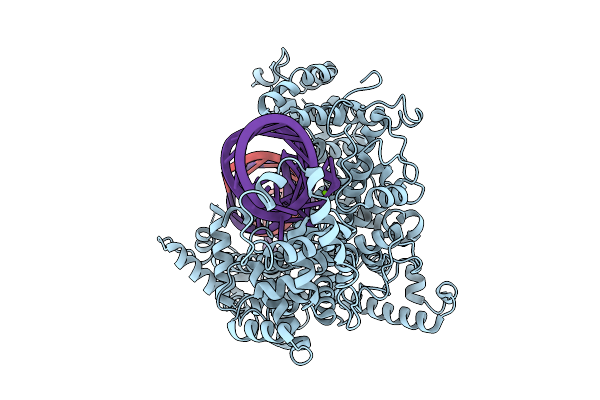 |
Organism: Ruminococcus sp. xpd3002, Cloning vector pgrna-ccdb-1
Method: ELECTRON MICROSCOPY Release Date: 2025-10-08 Classification: RNA BINDING PROTEIN/RNA Ligands: MG |
 |
Organism: Ruminococcus sp. xpd3002
Method: ELECTRON MICROSCOPY Release Date: 2025-09-17 Classification: RNA BINDING PROTEIN/RNA Ligands: MG |
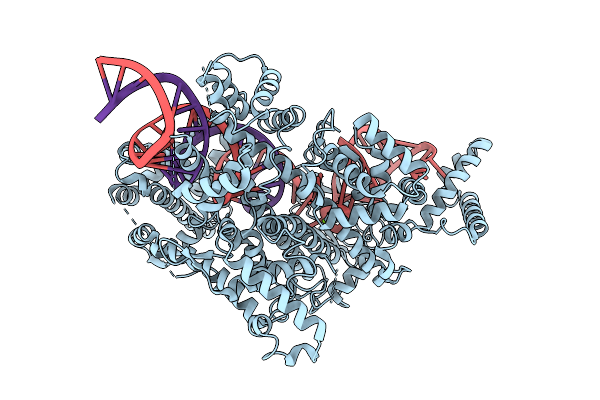 |
Organism: Ruminococcus sp. xpd3002
Method: ELECTRON MICROSCOPY Release Date: 2025-09-17 Classification: RNA BINDING PROTEIN/RNA Ligands: MG |
 |
Three-Dimensional Structure Of Homo-Dimer Of Cystathione Beta Lyase/Plp From Bacillus Cereus(Bcpatb)
Organism: Bacillus cereus atcc 14579
Method: X-RAY DIFFRACTION Resolution:2.28 Å Release Date: 2024-11-20 Classification: BIOSYNTHETIC PROTEIN |
 |
Three-Dimensional Structure Of Homo-Dimer Of Cystathione Beta Lyase/Plp/+L-Alliin Complex From Bacillus Cereus(Bcpatb)
Organism: Bacillus cereus atcc 14579
Method: X-RAY DIFFRACTION Resolution:1.88 Å Release Date: 2024-11-20 Classification: BIOSYNTHETIC PROTEIN Ligands: OO0 |
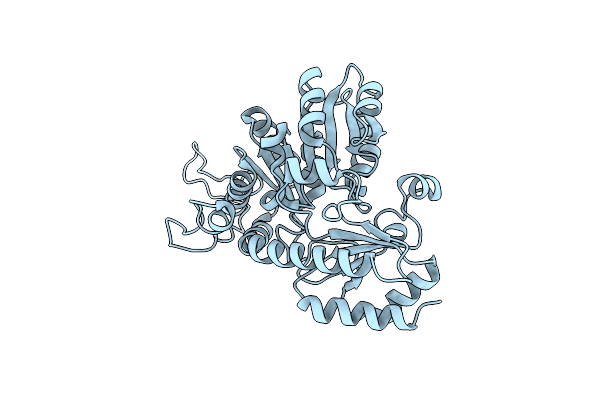 |
Crystal Structures Of Cystathionine Beta Lyase From Bacillus Cereus Atcc 14579
Organism: Bacillus cereus atcc 14579
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2024-07-03 Classification: LYASE Ligands: PLP |
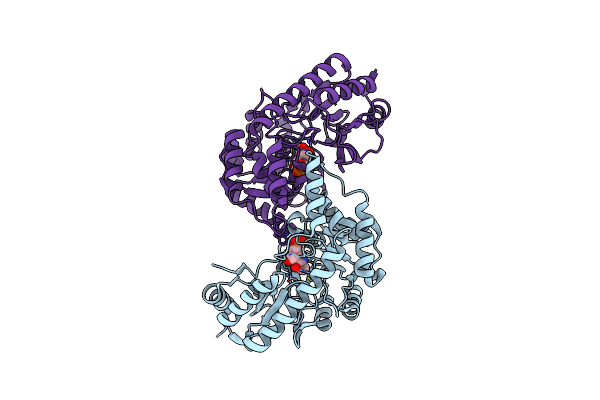 |
Crystal Structures Of Cystathionine Beta Lyase From Bacillus Cereus Atcc 14579
Organism: Bacillus cereus atcc 14579
Method: X-RAY DIFFRACTION Resolution:1.99 Å Release Date: 2024-07-03 Classification: LYASE Ligands: PLP |
 |
Crystal Structure Of A Double Mutant Acetyltransferase From Bacillus Cereus Species.
Organism: Bacillus cereus atcc 14579
Method: X-RAY DIFFRACTION Resolution:1.52 Å Release Date: 2024-03-06 Classification: TRANSFERASE Ligands: EDO, ZN |
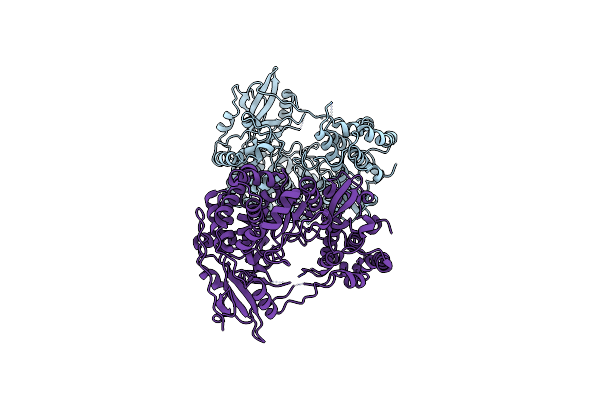 |
Organism: Alongshan virus
Method: X-RAY DIFFRACTION Resolution:3.20 Å Release Date: 2024-02-07 Classification: VIRAL PROTEIN |
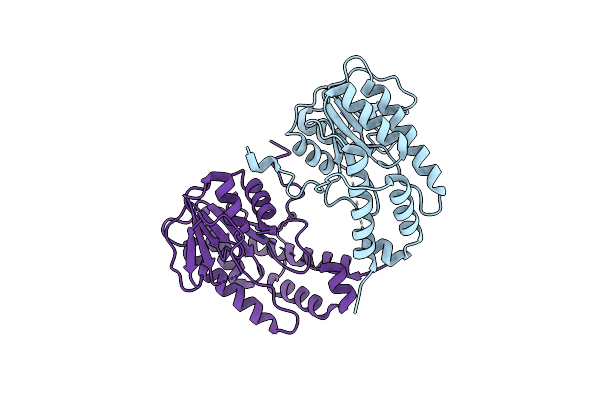 |
Organism: Alongshan virus
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2023-09-27 Classification: VIRAL PROTEIN |
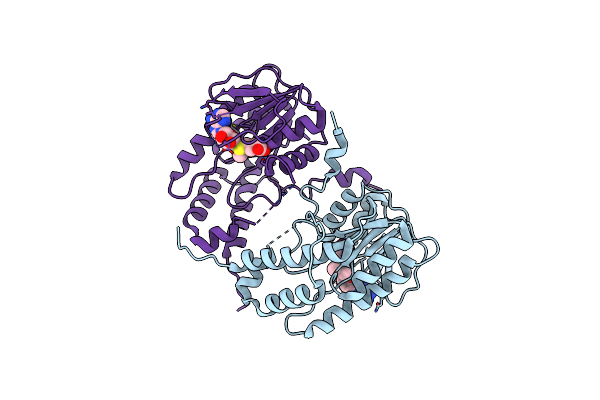 |
Crystal Structure Of Alongshan Virus Methyltransferase Bound To S-Adenosyl-L-Methionine
Organism: Alongshan virus
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2023-09-27 Classification: VIRAL PROTEIN Ligands: SAM |
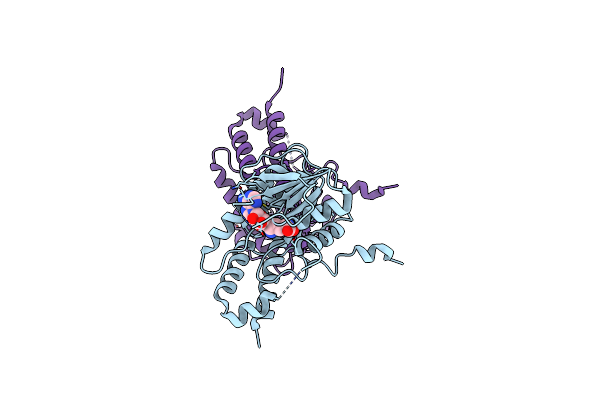 |
Organism: Alongshan virus
Method: X-RAY DIFFRACTION Resolution:2.01 Å Release Date: 2023-09-27 Classification: VIRAL PROTEIN Ligands: SFG |
 |
Crystal Structure Of Alongshan Virus Methyltransferase Bound To S-Adenosyl-L-Homocysteine
Organism: Alongshan virus
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2023-09-27 Classification: VIRAL PROTEIN Ligands: SAH |
 |
Organism: Bacillus cereus atcc 14579
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2023-07-12 Classification: OXIDOREDUCTASE Ligands: PDO |
 |
Organism: Bacillus cereus atcc 14579
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2023-07-12 Classification: OXIDOREDUCTASE Ligands: HEM, PGO, BU1, PDO |
 |
Crystal Structure Of Asymmetric Ferredoxin/Flavodoxin Nadp+ Oxidoreductase 2 (Fnr2) H326V Mutant From Bacillus Cereus
Organism: Bacillus cereus atcc 14579
Method: X-RAY DIFFRACTION Resolution:4.20 Å Release Date: 2023-07-12 Classification: OXIDOREDUCTASE Ligands: FAD |
 |
Crystal Structure Of Ferredoxin/Flavodoxin Nadp+ Oxidoreductase 1 (Fnr1) V329H Mutant From Bacillus Cereus
Organism: Bacillus cereus atcc 14579
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2023-07-12 Classification: OXIDOREDUCTASE Ligands: FAD, OXM, ACT |
 |
Organism: Bacillus cereus atcc 14579
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2023-02-22 Classification: TRANSPORT PROTEIN Ligands: MPD, RB, BA |
 |
Organism: Bacillus cereus atcc 14579
Method: X-RAY DIFFRACTION Resolution:2.75 Å Release Date: 2023-02-22 Classification: TRANSPORT PROTEIN Ligands: RB, CA, MPD, CL |