Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
 |
Organism: Homo sapiens, Metamycoplasma arthritidis
Method: ELECTRON MICROSCOPY Release Date: 2024-10-02 Classification: IMMUNE SYSTEM Ligands: NAG |
 |
Crystal Structure Of The Cofactor-Devoid 1-H-3-Hydroxy-4- Oxoquinaldine 2,4-Dioxygenase (Hod) S101A Variant Complexed With 2-Methyl-Quinolin-4(1H)-One Under Hyperoxyc Conditions
Organism: Paenarthrobacter nitroguajacolicus
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2024-01-17 Classification: OXIDOREDUCTASE Ligands: VFH, OXY, TAR, GOL, NA |
 |
Crystal Structure Of The Cofactor-Devoid 1-H-3-Hydroxy-4- Oxoquinaldine 2,4-Dioxygenase (Hod) S101A Variant Complexed With 2-Methyl-Quinolin-4(1H)-One Under Normoxyc Conditions
Organism: Paenarthrobacter nitroguajacolicus
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2024-01-17 Classification: OXIDOREDUCTASE Ligands: VFH, GOL, SRT |
 |
Crystal Structure Of The Cofactor-Devoid 1-H-3-Hydroxy-4- Oxoquinaldine 2,4-Dioxygenase (Hod) H251A Variant Complexed With N-Acetylanthranilate As Result Of In Crystallo Turnover Of Its Natural Substrate 1-H-3-Hydroxy-4- Oxoquinaldine Under Hyperoxic Conditions
Organism: Paenarthrobacter nitroguajacolicus
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2024-01-17 Classification: OXIDOREDUCTASE Ligands: ZZ8, SRT, GOL, K |
 |
Organism: Blautia producta atcc 27340 = dsm 2950
Method: X-RAY DIFFRACTION Resolution:2.32 Å Release Date: 2023-11-29 Classification: HYDROLASE Ligands: GOL, MES, ACT, CL, SO4, EDO |
 |
Room Temperature Crystal Structure Of The Cofactor-Devoid 1-H-3-Hydroxy-4- Oxoquinaldine 2,4-Dioxygenase (Hod) Under Xenon Pressure (30 Bar)
Organism: Paenarthrobacter nitroguajacolicus
Method: X-RAY DIFFRACTION Resolution:2.90 Å Release Date: 2023-07-05 Classification: OXIDOREDUCTASE Ligands: XE, TAR |
 |
Crystal Structure Of The Cofactor-Devoid 1-H-3-Hydroxy-4- Oxoquinaldine 2,4-Dioxygenase (Hod) Catalytically Inactive H251A Variant Complexed With 2-Methyl-Quinolin-4(1H)-One Under Normoxic Conditions
Organism: Paenarthrobacter nitroguajacolicus
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2022-06-01 Classification: OXIDOREDUCTASE Ligands: VFH, K, GOL, SRT, NA |
 |
Crystal Structure Of The Cofactor-Devoid 1-H-3-Hydroxy-4- Oxoquinaldine 2,4-Dioxygenase (Hod) Catalytically Inactive H251A Variant Complexed With 2-Methyl- Quinolin-4(1H)-One Under Hyperoxic Conditions
Organism: Paenarthrobacter nitroguajacolicus
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2022-06-01 Classification: OXIDOREDUCTASE Ligands: VFH, K, TAR, GOL |
 |
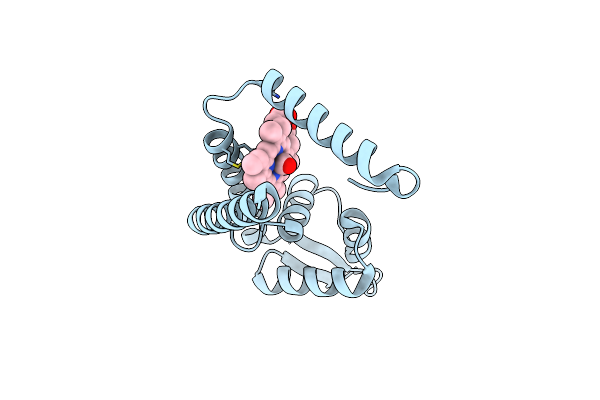
Crystal Structure Of Heme Sensor Protein Pefr From Streptococcus Agalactiae In Complex With Heme
Organism: Streptococcus agalactiae serotype iii (strain nem316)
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2021-09-29 Classification: TRANSCRIPTION Ligands: HEM, CO |
 |
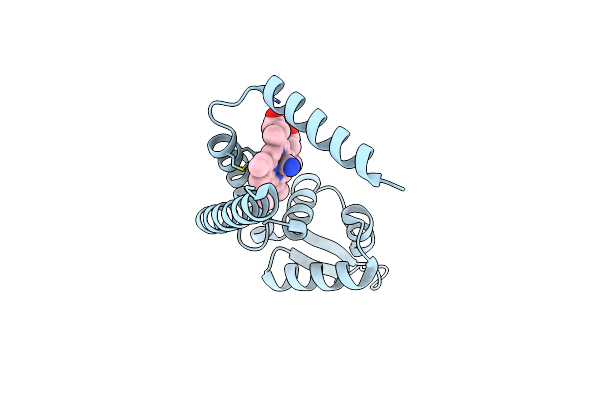
Crystal Structure Of Heme Sensor Protein Pefr In Complex With Heme And Carbon Monoxide
Organism: Streptococcus agalactiae serotype iii (strain nem316)
Method: X-RAY DIFFRACTION Resolution:2.09 Å Release Date: 2021-09-29 Classification: TRANSCRIPTION Ligands: HEM, CMO |
 |
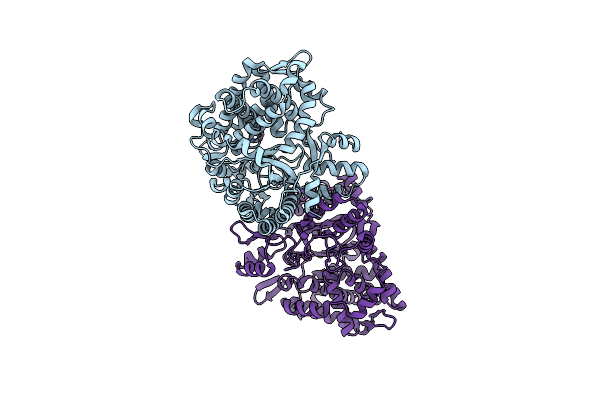
Crystal Structure Of Heme Sensor Protein Pefr In Complex With Heme And Cyanide
Organism: Streptococcus agalactiae serotype iii (strain nem316)
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2021-09-29 Classification: TRANSCRIPTION Ligands: HEM, CYN |
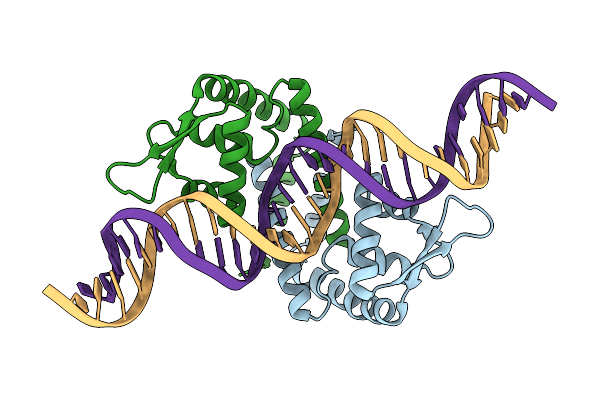 |
Heme Sensor Protein Pefr From Streptococcus Agalactiae Bound To Operator Dna (28-Mer)
Organism: Streptococcus agalactiae serotype iii (strain nem316), Streptococcus agalactiae nem316
Method: X-RAY DIFFRACTION Resolution:2.49 Å Release Date: 2021-09-29 Classification: TRANSCRIPTION |
 |
Organism: Corallococcus sp. egb
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2019-06-12 Classification: HYDROLASE Ligands: ZN |
 |
Organism: Streptococcus agalactiae nem316
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2019-01-02 Classification: CELL ADHESION |
 |
Organism: Streptococcus agalactiae serotype iii (strain nem316)
Method: X-RAY DIFFRACTION Resolution:1.36 Å Release Date: 2018-04-18 Classification: TRANSFERASE Ligands: NAD, SO4 |
 |
Organism: Streptococcus agalactiae serotype iii (strain nem316)
Method: X-RAY DIFFRACTION Resolution:2.41 Å Release Date: 2016-06-22 Classification: CELL ADHESION Ligands: ACT, PEG, EDO |
 |
Organism: Streptococcus agalactiae serotype iii (strain nem316)
Method: X-RAY DIFFRACTION Resolution:1.89 Å Release Date: 2016-06-22 Classification: CELL ADHESION |
 |
Organism: Streptococcus agalactiae serotype iii (strain nem316)
Method: X-RAY DIFFRACTION Resolution:2.19 Å Release Date: 2016-06-22 Classification: CELL ADHESION Ligands: EDO, PEG |
 |
Crystal Structure Of Recombinant Foot-And-Mouth-Disease Virus A22-H2093F Empty Capsid
Organism: Foot-and-mouth disease virus - type a
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2015-09-23 Classification: VIRUS |
 |
Crystal Structure Of Keratin 4 Binding Domain Of Surface Adhesin Srr-1 Of S.Agalactiae
Organism: Streptococcus agalactiae nem316
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2015-02-11 Classification: CELL ADHESION |