Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
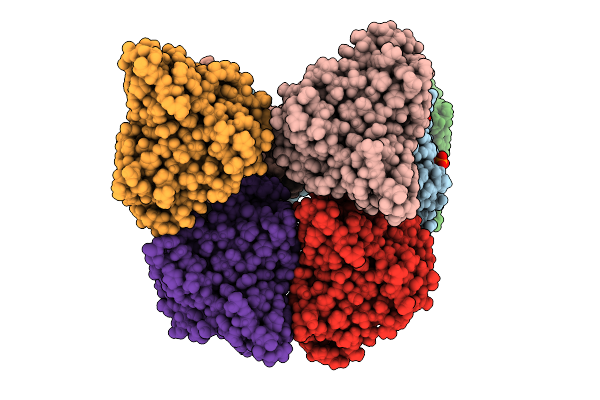 |
Organism: Cellvibrio japonicus
Method: X-RAY DIFFRACTION Release Date: 2025-12-24 Classification: OXIDOREDUCTASE Ligands: CU, PO4 |
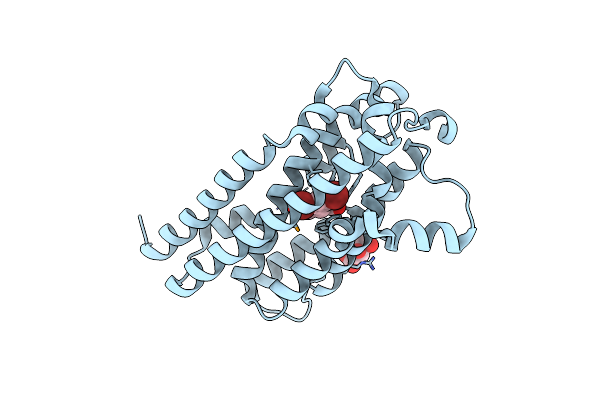 |
Organism: Aetokthonos hydrillicola
Method: X-RAY DIFFRACTION Resolution:2.08 Å Release Date: 2024-08-28 Classification: OXIDOREDUCTASE Ligands: 67I, MLT |
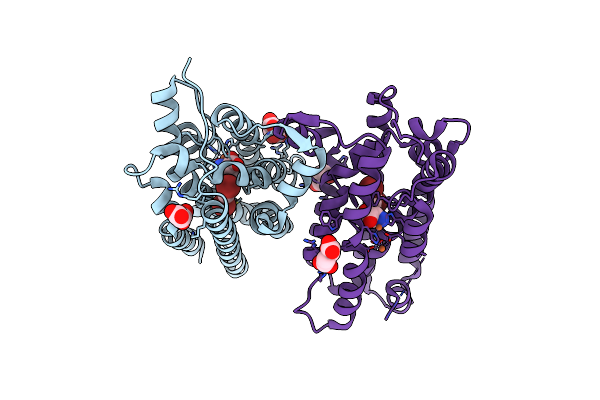 |
Crystal Structure Of Nitrile Synthase Aetd With Substrate Bound And Cofactor Partially Assembled
Organism: Aetokthonos hydrillicola
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2024-08-28 Classification: OXIDOREDUCTASE Ligands: 67I, MLT, SIN, FE, GOL |
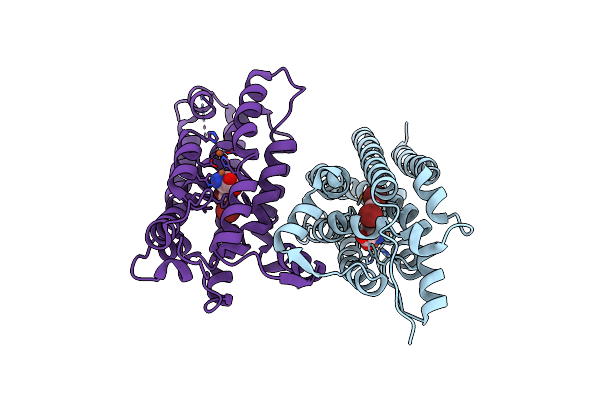 |
Crystal Structure Of Nitrile Synthase Aetd With Substrate Bound And Cofactor Fully Assembled
Organism: Aetokthonos hydrillicola
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2024-08-28 Classification: OXIDOREDUCTASE Ligands: 67I, FE |
 |
Organism: Cellvibrio japonicus
Method: X-RAY DIFFRACTION Resolution:1.67 Å Release Date: 2023-11-29 Classification: HYDROLASE Ligands: RBH, XYS, YLL |
 |
Organism: Cellvibrio japonicus
Method: X-RAY DIFFRACTION Resolution:1.57 Å Release Date: 2023-11-29 Classification: HYDROLASE |
 |
Organism: Cellvibrio japonicus
Method: X-RAY DIFFRACTION Resolution:2.18 Å Release Date: 2023-11-22 Classification: HYDROLASE |
 |
Organism: Bacillus phage ar9
Method: X-RAY DIFFRACTION Resolution:3.30 Å Release Date: 2022-07-06 Classification: TRANSCRIPTION Ligands: ZN |
 |
X-Ray Structure Of The Phage Ar9 Non-Virion Rna Polymerase Holoenzyme In Complex With A Forked Oligonucleotide Containing The P077 Promoter
Organism: Bacillus phage ar9
Method: X-RAY DIFFRACTION Resolution:3.40 Å Release Date: 2022-07-06 Classification: TRANSCRIPTION/DNA Ligands: ZN |
 |
Structure Of The Phage Ar9 Non-Virion Rna Polymerase Holoenzyme In Complex With Two Dna Oligonucleotides Containing The Ar9 P077 Promoter As Determined By Cryo-Em
Organism: Bacillus phage ar9, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2022-07-06 Classification: TRANSCRIPTION Ligands: ZN |
 |
Structure Of Bacteriophage Ar9 Non-Virion Rnap Polymerase Holoenzyme Determined By Cryo-Em
Organism: Bacillus phage ar9
Method: ELECTRON MICROSCOPY Release Date: 2022-07-06 Classification: TRANSCRIPTION Ligands: ZN |
 |
Organism: Cellvibrio japonicus
Method: X-RAY DIFFRACTION Resolution:1.46 Å Release Date: 2020-08-19 Classification: HYDROLASE |
 |
Crystal Structure Of Cjagd31B (Alpha-Transglucosylase From Glycoside Hydrolase Family 31) In Complex With Alpha Cyclophellitol Cyclosulfate Probe Me647
Organism: Cellvibrio japonicus
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2017-08-09 Classification: HYDROLASE Ligands: SO4, PG4, 93Z, OXL |
 |
Crystal Structure Of D412N Nucleophile Mutant Cjagd31B (Alpha-Transglucosylase From Glycoside Hydrolase Family 31) In Complex With Unreacted Alpha Cyclophellitol Cyclosulfate Probe Me647
Organism: Cellvibrio japonicus
Method: X-RAY DIFFRACTION Resolution:1.96 Å Release Date: 2017-08-09 Classification: HYDROLASE Ligands: SO4, EDO, PG4, 94E, OXL |
 |
Crystal Structure Of D412N Nucleophile Mutant Cjagd31B (Alpha-Transglucosylase From Glycoside Hydrolase Family 31) In Complex With Alpha Cyclophellitol Aziridine Probe Cf021
Organism: Cellvibrio japonicus
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2017-08-09 Classification: HYDROLASE Ligands: SO4, PG4, EDO, 94B, OXL |
 |
Structural And Functional Analysis Of A Lytic Polysaccharide Monooxygenase Important For Efficient Utilization Of Chitin In Cellvibrio Japonicus
Organism: Cellvibrio japonicus
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2016-02-17 Classification: SUGAR BINDING PROTEIN Ligands: CU |
 |
Crystal Structure Of Apo Agd31B, Alpha-Transglucosylase In Glycoside Hydrolase Family 31
Organism: Cellvibrio japonicus
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2012-11-14 Classification: HYDROLASE Ligands: SO4, OXL, EDO, PG4 |
 |
Crystal Structure Of Agd31B, Alpha-Transglucosylase, Complexed With Acarbose
Organism: Cellvibrio japonicus
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2012-11-14 Classification: HYDROLASE Ligands: SO4, OXL, PEG, EDO |
 |
Crystal Structure Of Agd31B, Alpha-Transglucosylase, Complexed With 5F-Alpha-Glcf
Organism: Cellvibrio japonicus
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2012-11-14 Classification: HYDROLASE Ligands: ARG, 5GF, EDO, SO4, PGE, PEG |
 |
Crystal Structure Of Alpha-Xylosidase (Gh31) From Cellvibrio Japonicus In Complex With 5-Fluoro-Alpha-D-Xylopyranosyl Fluoride
Organism: Cellvibrio japonicus
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2011-04-13 Classification: HYDROLASE Ligands: CL, NI, FFX |