Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
 |
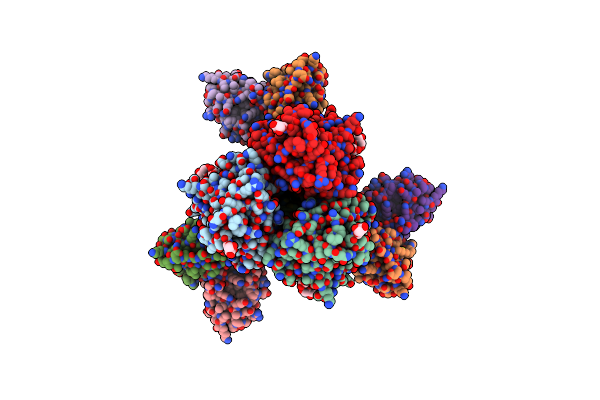
Organism: Clostridium symbiosum
Method: X-RAY DIFFRACTION Release Date: 2025-08-20 Classification: TRANSFERASE Ligands: ANP, SO4 |
 |
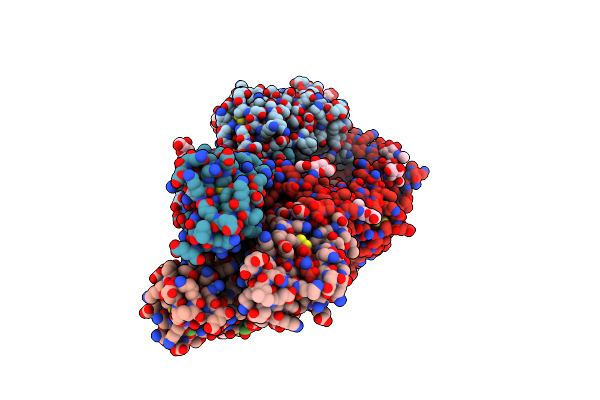
Hemagglutinin H1 New Caledonia 1999 In Complex With Monoclonal Antibody Fab 43_S0008
Organism: Influenza a virus, Mus sp.
Method: ELECTRON MICROSCOPY Release Date: 2024-12-11 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
 |
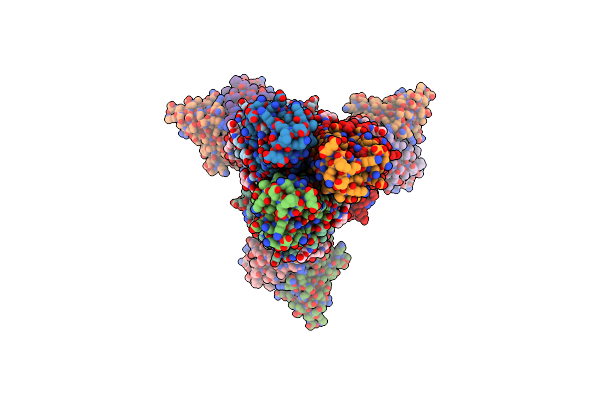
Organism: Influenza a virus (a/new caledonia/20/1999(h1n1)), Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-10-11 Classification: VIRAL PROTEIN/Immune System Ligands: NAG |
 |
Organism: Influenza a virus (a/new caledonia/20/1999(h1n1)), Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-10-11 Classification: VIRAL PROTEIN/Immune System Ligands: NAG |
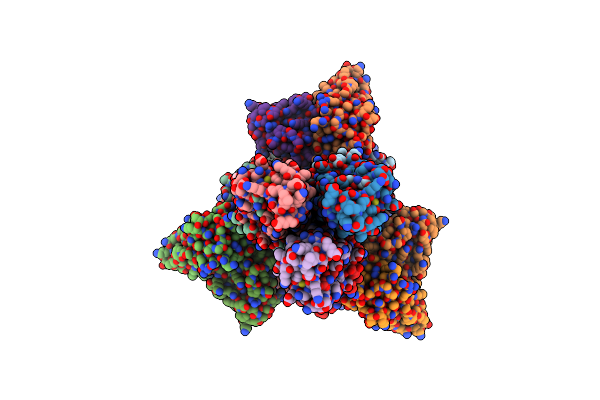 |
H1 Hemagglutinin (Nc99) In Complex With Medial-Junction-Targeting Fab 2-2-1G06
Organism: Influenza a virus (a/new caledonia/20/1999(h1n1)), Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-10-11 Classification: VIRAL PROTEIN/Immune System Ligands: NAG |
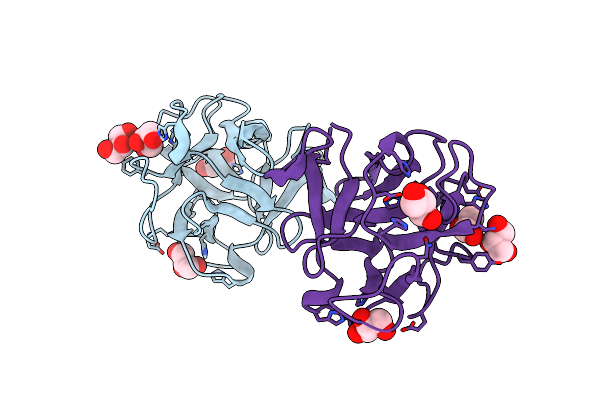 |
Organism: Crenomytilus grayanus
Method: X-RAY DIFFRACTION Resolution:1.08 Å Release Date: 2016-04-06 Classification: SUGAR BINDING PROTEIN Ligands: GOL |
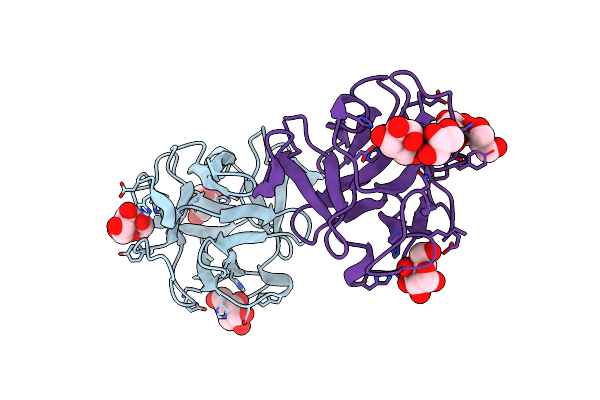 |
Crystal Structure Of A Crenomytilus Grayanus Lectin In Complex With Galactose
Organism: Crenomytilus grayanus
Method: X-RAY DIFFRACTION Resolution:1.56 Å Release Date: 2016-04-06 Classification: SUGAR BINDING PROTEIN Ligands: GAL, GLA, GOL |
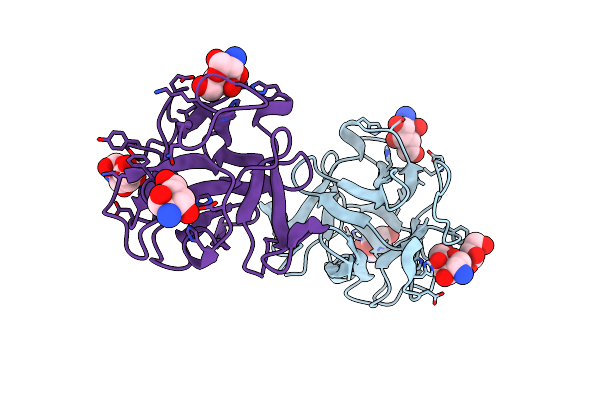 |
Crystal Structure Of A Crenomytilus Grayanus Lectin In Complex With Galactosamine
Organism: Crenomytilus grayanus
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2016-04-06 Classification: SUGAR BINDING PROTEIN Ligands: X6X, GOL |
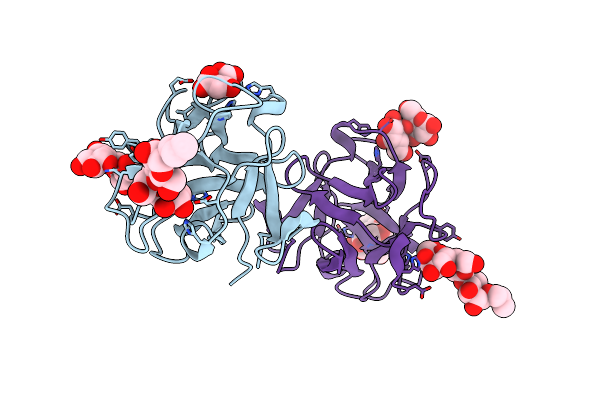 |
Crystal Structure Of A Crenomytilus Grayanus Lectin In Complex With Gb3 Allyl
Organism: Crenomytilus grayanus
Method: X-RAY DIFFRACTION Resolution:1.64 Å Release Date: 2016-04-06 Classification: SUGAR BINDING PROTEIN Ligands: LMR, 5VQ |
 |
Organism: Crenomytilus grayanus
Method: X-RAY DIFFRACTION Resolution:2.12 Å Release Date: 2015-11-11 Classification: SUGAR BINDING PROTEIN Ligands: GOL |
 |
Crystal Structure Of A Generation 3 Influenza Hemagglutinin Stabilized Stem Complexed With The Broadly Neutralizing Antibody C179
Organism: Influenza a virus, human immunodeficiency virus type 1 group m subtype b, enterobacteria phage t4, Mus musculus
Method: X-RAY DIFFRACTION Resolution:3.19 Å Release Date: 2015-09-02 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
 |
Crystal Structure Of A Generation 4 Influenza Hemagglutinin Stabilized Stem In Complex With The Broadly Neutralizing Antibody Cr6261
Organism: Influenza a virus, human immunodeficiency virus type 1 group m subtype b, enterobacteria phage t4, Homo sapiens
Method: X-RAY DIFFRACTION Resolution:4.30 Å Release Date: 2015-09-02 Classification: VIRAL PROTEIN/IMMUNE SYSTEM |
 |
Organism: Escherichia coli, Clostridium difficile cd160
Method: X-RAY DIFFRACTION Resolution:1.72 Å Release Date: 2015-01-14 Classification: CELL ADHESION Ligands: MLI |
 |
Organism: Clostridium symbiosum, Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2012-04-04 Classification: OXIDOREDUCTASE |
 |
Glutaconyl-Coa Decarboxylase A Subunit From Clostridium Symbiosum Co-Crystallized With Glutaconyl-Coa
Organism: Clostridium symbiosum
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2009-07-28 Classification: LYASE Ligands: COO, CL |
 |
Glutaconyl-Coa Decarboxylase A Subunit From Clostridium Symbiosum Apoprotein
Organism: Clostridium symbiosum
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2009-07-28 Classification: LYASE Ligands: SO4 |
 |
Glutaconyl-Coa Decarboxylase A Subunit From Clostridium Symbiosum Co-Crystallized With Crotonyl-Coa
Organism: Clostridium symbiosum
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2009-07-28 Classification: LYASE Ligands: CL, COO |
 |
Glutaconyl-Coa Decarboxylase A Subunit From Clostridium Symbiosum Co-Crystallized With Glutaryl-Coa
Organism: Clostridium symbiosum
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2009-07-28 Classification: LYASE Ligands: GRA |
 |
Pyruvate Phosphate Dikinase (Ppdk) Triple Mutant R219E/E271R/S262D Adapts A Second Conformational State
Organism: Clostridium symbiosum
Method: X-RAY DIFFRACTION Resolution:3.60 Å Release Date: 2008-01-01 Classification: TRANSFERASE Ligands: SO4 |
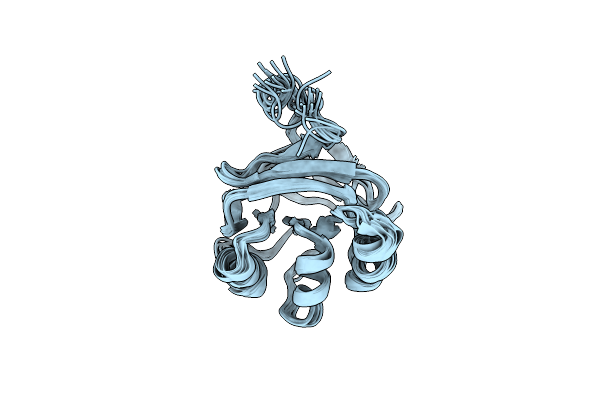 |
Nmr Structure Of The Phosphoryl Carrier Domain Of Pyruvate Phosphate Dikinase
Organism: Clostridium symbiosum
Method: SOLUTION NMR Release Date: 2006-10-03 Classification: TRANSFERASE |