Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
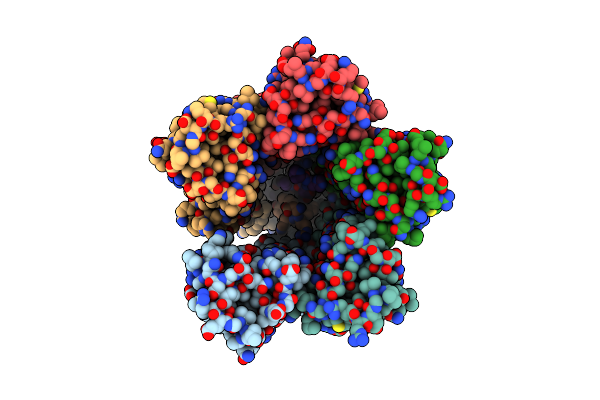 |
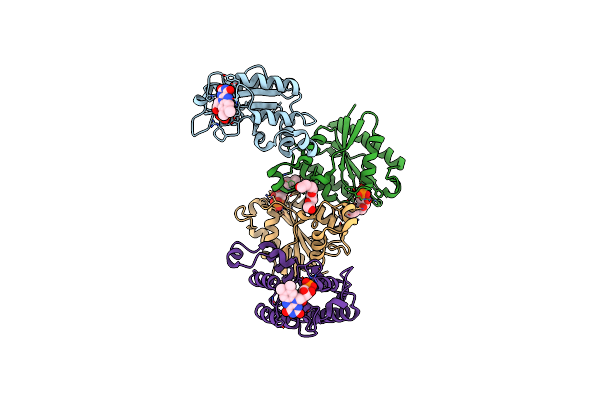
The Chimeric Flagellar Motor Complex Between Mota1B1 From Paenibacillus Sp. Tca20 And Motab From E.Coli, State 1
Organism: Paenibacillus sp. tca20, Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:3.30 Å Release Date: 2025-04-09 Classification: MOTOR PROTEIN Ligands: AV0 |
 |
The Chimeric Flagellar Motor Complex Between Mota1B1 From Paenibacillus Sp. Tca20 And Motab From E.Coli, State 2
Organism: Paenibacillus sp. tca20, Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:3.30 Å Release Date: 2025-04-09 Classification: MOTOR PROTEIN |
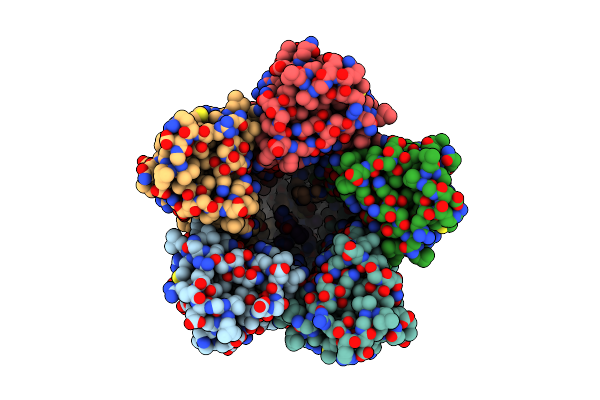 |
The Chimeric Flagellar Motor Complex Between Mota1B1 From Paenibacillus Sp. Tca20 And Motab From E.Coli, State 3
Organism: Paenibacillus sp. tca20
Method: ELECTRON MICROSCOPY Resolution:3.50 Å Release Date: 2025-04-09 Classification: MOTOR PROTEIN |
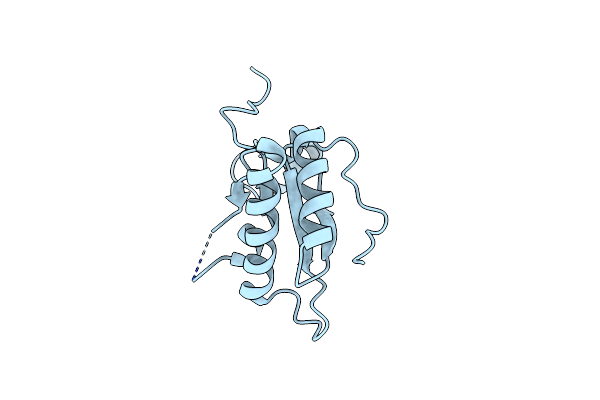 |
Organism: Escherichia phage ecd7
Method: X-RAY DIFFRACTION Resolution:2.49 Å Release Date: 2024-07-31 Classification: ANTIMICROBIAL PROTEIN |
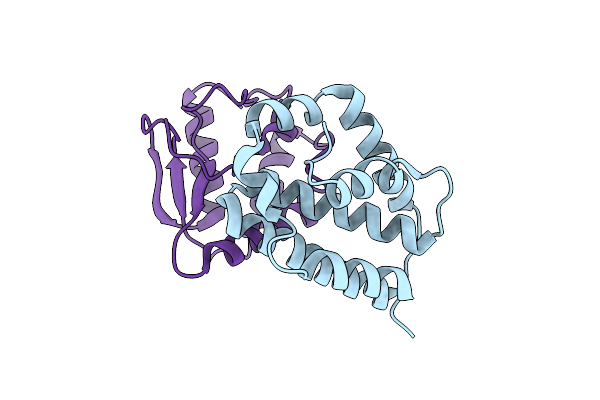 |
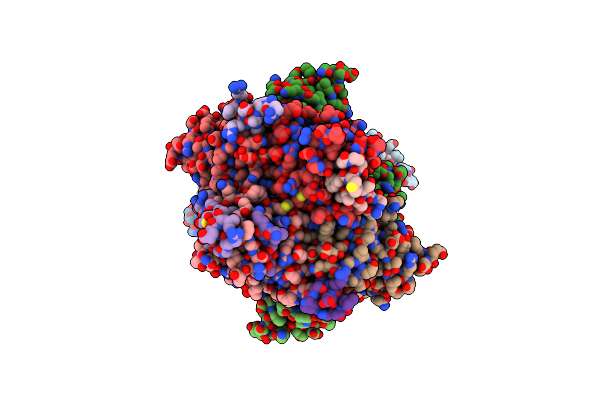
The Crystal Structure Of The Polymorphic Toxin Pt1(Em) H44A Mutant And Its Cognate Immunity Pim1(Em) Complex
Organism: Escherichia marmotae
Method: X-RAY DIFFRACTION Resolution:1.39 Å Release Date: 2024-07-03 Classification: TOXIN |
 |
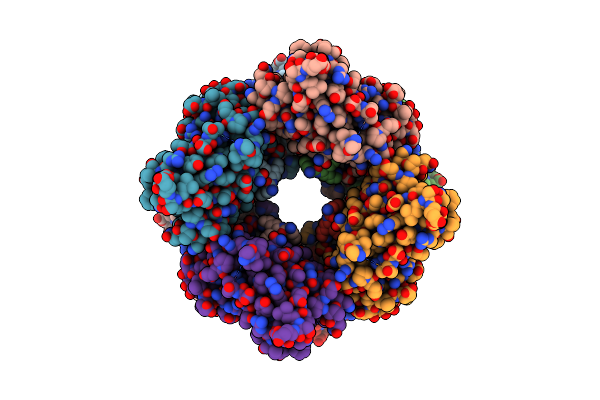
Organism: Komagataeibacter hansenii atcc 23769
Method: ELECTRON MICROSCOPY Release Date: 2022-12-28 Classification: STRUCTURAL PROTEIN |
 |
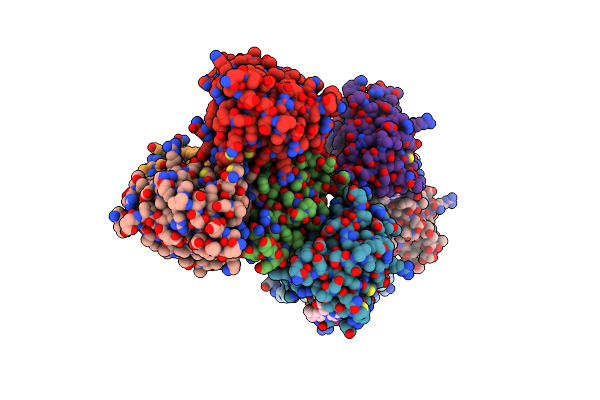
Organism: Komagataeibacter hansenii atcc 23769
Method: ELECTRON MICROSCOPY Release Date: 2022-12-28 Classification: STRUCTURAL PROTEIN |
 |
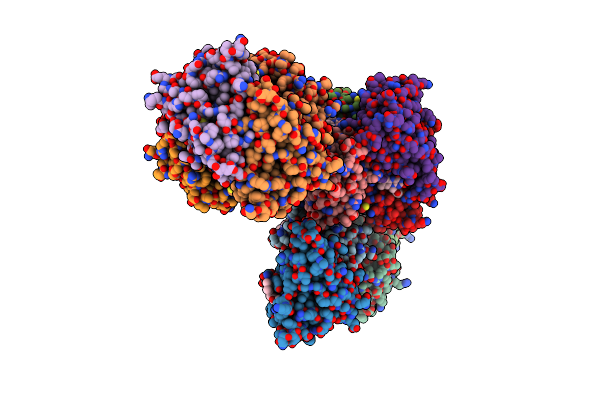
Crystal Structure Of A Putative Chromate Reductase From Gluconacetobacter Hansenii, Gh-Chrr, Containing A Y129N Substitution.
Organism: Gluconacetobacter hansenii
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2012-12-26 Classification: OXIDOREDUCTASE Ligands: FMN |
 |
Crystal Structure Of A Putative Chromate Reductase From Gluconacetobacter Hansenii, Gh-Chrr, Containing A R101A Substitution.
Organism: Gluconacetobacter hansenii
Method: X-RAY DIFFRACTION Resolution:2.56 Å Release Date: 2012-10-10 Classification: OXIDOREDUCTASE Ligands: FMN |
 |
Crystal Structure Of A Chromate/Uranium Reductase From Gluconacetobacter Hansenii
Organism: Gluconacetobacter hansenii
Method: X-RAY DIFFRACTION Resolution:2.24 Å Release Date: 2012-05-30 Classification: OXIDOREDUCTASE Ligands: FMN, CL, PG4 |
 |
Crystal Structure Of Carbohydrate-Binding Module Family 28 From Clostridium Josui Cel5A In Complex With Cellopentaose
Organism: Clostridium josui
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2010-03-31 Classification: HYDROLASE Ligands: CA, PO4 |
 |
Crystal Structure Of Carbohydrate-Binding Module Family 28 From Clostridium Josui Cel5A In A Ligand-Free Form
Organism: Clostridium josui
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2010-03-02 Classification: HYDROLASE Ligands: CA, SO4 |
 |
Crystal Structure Of Carbohydrate-Binding Module Family 28 From Clostridium Josui Cel5A In Complex With Cellobiose
Organism: Clostridium josui
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2010-03-02 Classification: HYDROLASE Ligands: CA, PO4, GOL |
 |
Crystal Structure Of Carbohydrate-Binding Module Family 28 From Clostridium Josui Cel5A In Complex With Cellotetraose
Organism: Clostridium josui
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2010-03-02 Classification: HYDROLASE Ligands: CA, PO4 |
 |
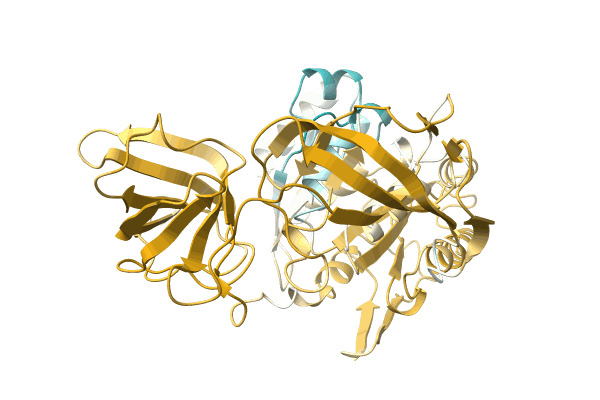 |
 |
 |
 |
 |