Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
 |
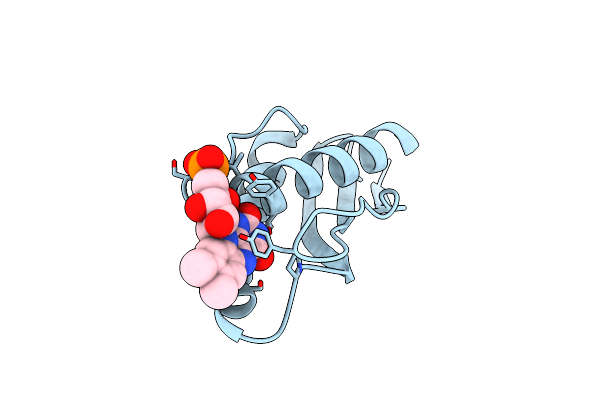
Organism: Brevibacillus laterosporus, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2024-07-10 Classification: RNA BINDING PROTEIN/RNA/DNA Ligands: EDO, SO4, CL |
 |
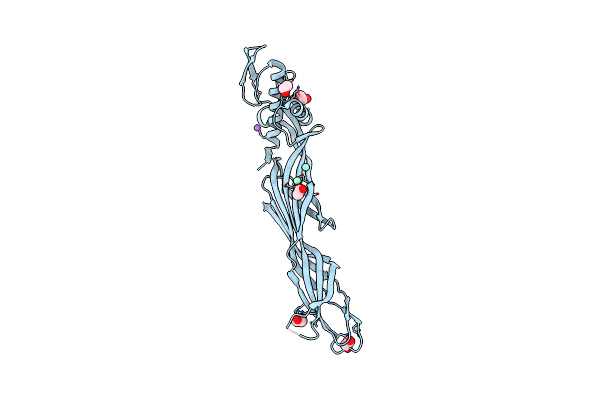
Organism: Streptomyces azureus
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2023-07-19 Classification: FLAVOPROTEIN Ligands: FMN |
 |
Organism: Brevibacillus laterosporus
Method: X-RAY DIFFRACTION Resolution:1.94 Å Release Date: 2021-05-05 Classification: TOXIN Ligands: SM, ACT, EDO, NA |
 |
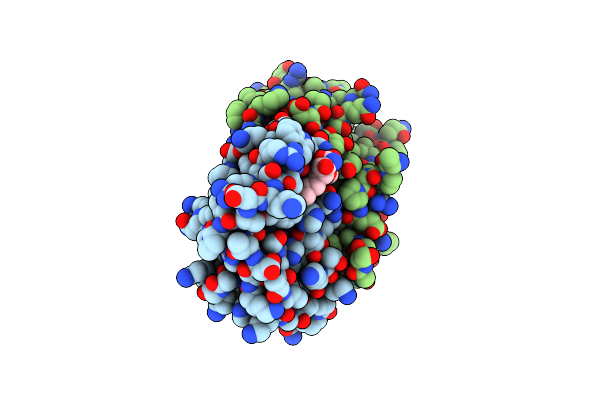
Organism: Streptomyces curacoi
Method: X-RAY DIFFRACTION Resolution:1.63 Å Release Date: 2018-08-29 Classification: BIOSYNTHETIC PROTEIN Ligands: FE2 |
 |
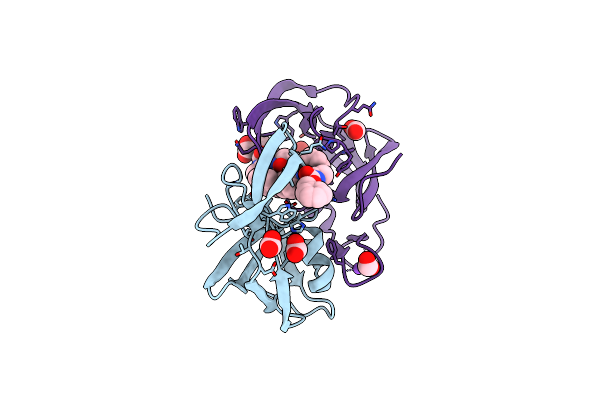
Organism: Dg-75 murine leukemia virus, Streptomyces
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2012-08-01 Classification: HYDROLASE/HYDROLASE INHIBITOR Ligands: FMT |
 |
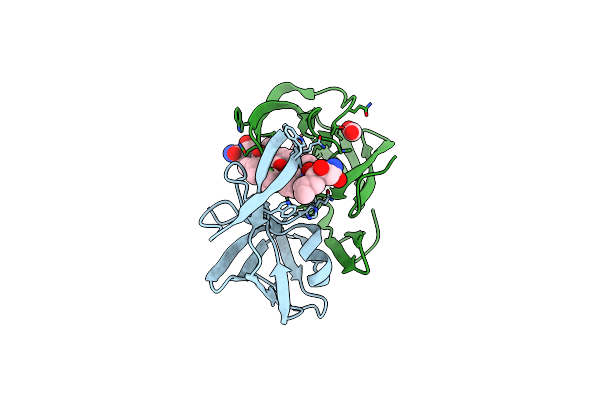
Organism: Dg-75 murine leukemia virus
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2011-10-12 Classification: hydrolase/hydrolase inhibitor Ligands: 3TL, FMT, NA |
 |
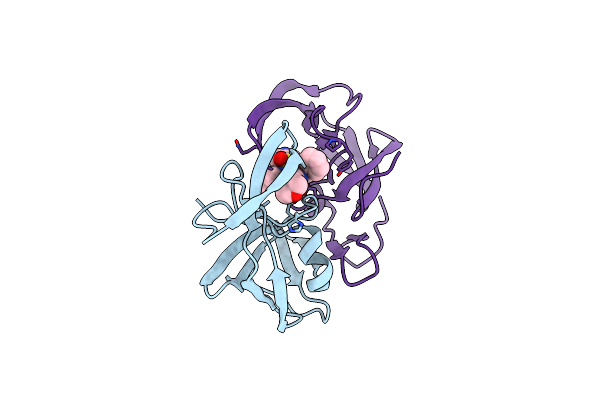
Organism: Dg-75 murine leukemia virus, Streptomyces argenteolus subsp. toyonakensis
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2011-10-12 Classification: hydrolase/hydrolase inhibitor Ligands: FMT |
 |
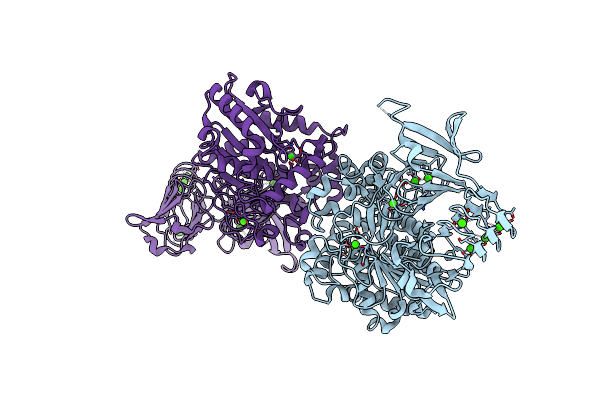
Organism: Dg-75 murine leukemia virus
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2011-10-12 Classification: hydrolase/hydrolase inhibitor Ligands: 478 |
 |
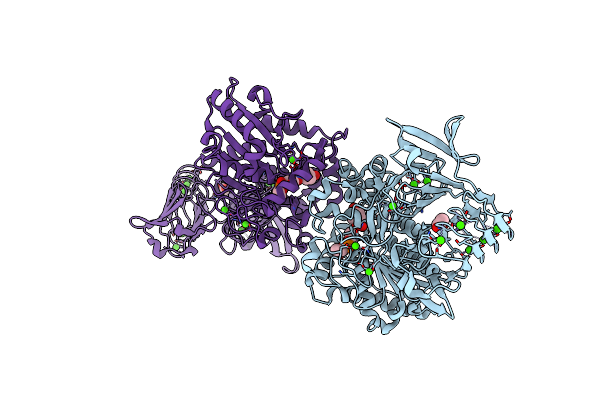
Crystal Structure Of Pseudomonas Sp. Mis38 Lipase (Pml) In The Open Conformation Following Dialysis Against Ca-Free Buffer
Organism: Pseudomonas
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2010-05-26 Classification: HYDROLASE Ligands: CA |
 |
Crystal Structure Of Pseudomonas Sp. Mis38 Lipase In Complex With Diethyl Phosphate
Organism: Pseudomonas
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2010-05-26 Classification: HYDROLASE Ligands: CA, DEP, NPO, ACT |
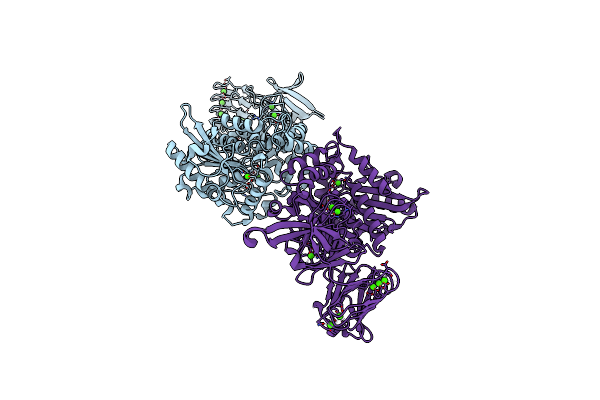 |
Organism: Pseudomonas
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2009-11-10 Classification: HYDROLASE Ligands: CA |
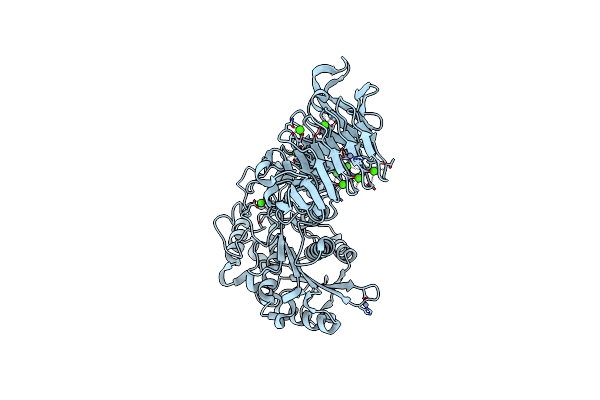 |
Organism: Pseudomonas sp.
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2008-12-02 Classification: HYDROLASE Ligands: CA, ZN |
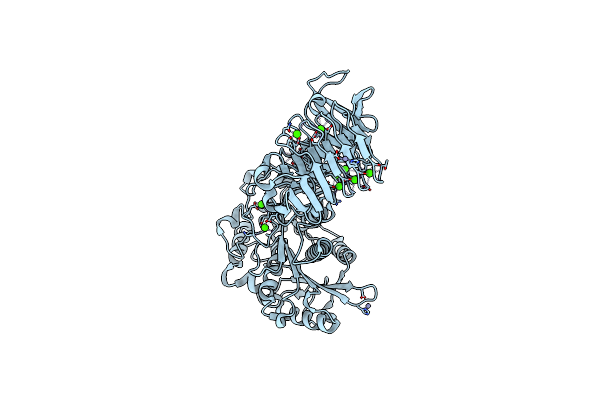 |
Organism: Pseudomonas sp.
Method: X-RAY DIFFRACTION Resolution:2.21 Å Release Date: 2008-12-02 Classification: HYDROLASE Ligands: CA, ZN |
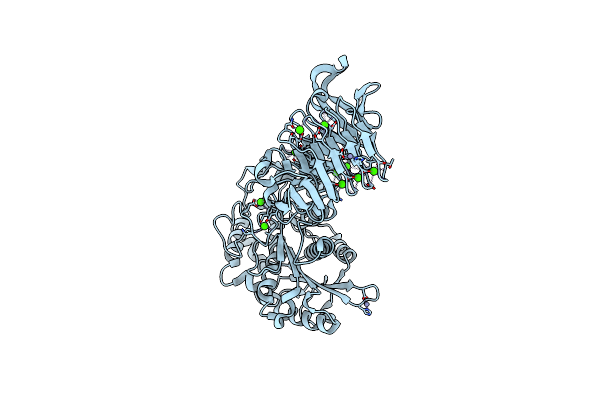 |
Organism: Pseudomonas sp. mis38
Method: X-RAY DIFFRACTION Resolution:1.48 Å Release Date: 2007-10-30 Classification: HYDROLASE Ligands: CA, ZN |
 |
Crystal Structure Of A Platinum-Bound S445C Mutant Of Pseudomonas Sp. Mis38 Lipase
Organism: Pseudomonas sp.
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2007-10-30 Classification: HYDROLASE Ligands: CA, ZN, PT |
 |
 |
 |
 |
NA
|
 |
NA
|