Search Count: 1,150
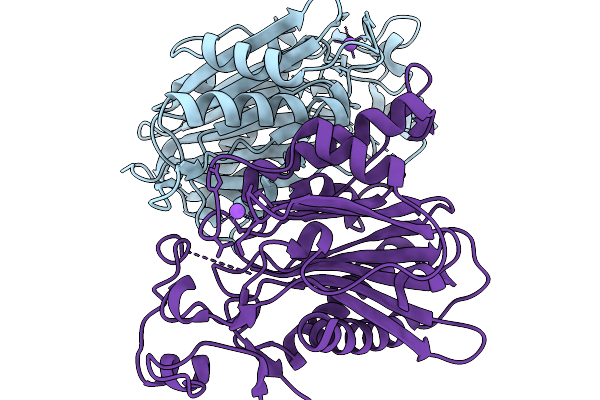 |
Crystal Structure Of The Poly(Hexamethylene Adipamide) (Nylon66) Hydrolase Nyl50 In Its Apo Form At Room Temperature
Organism: Alphaproteobacteria bacterium
Method: X-RAY DIFFRACTION Release Date: 2025-11-19 Classification: HYDROLASE |
 |
X-Ray Crystallographic Structure Of The Poly(Hexamethylene Adipamide) (Nylon66) Hydrolase Nyl50 At Room Temperature
Organism: Alphaproteobacteria bacterium
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2025-01-29 Classification: HYDROLASE Ligands: EDO, NA, PGE |
 |
X-Ray Crystallographic Structure Of The Poly(Hexamethylene Adipamide) (Nylon66) Hydrolase Nyl50 At Room Temperature Bound To Tetraethylene Glycol
Organism: Alphaproteobacteria bacterium
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2025-01-29 Classification: HYDROLASE Ligands: EDO, NA, PG4 |
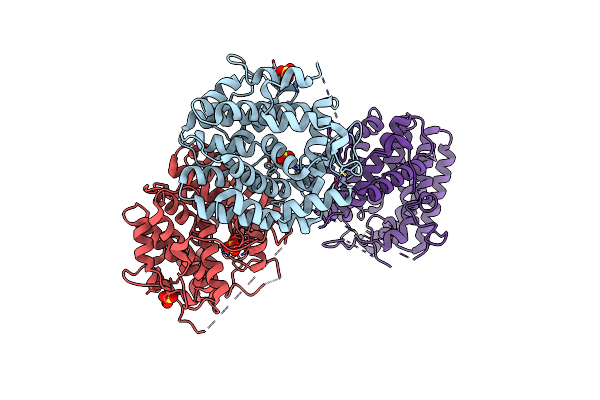 |
Organism: Shimazuella kribbensis
Method: X-RAY DIFFRACTION Resolution:1.96 Å Release Date: 2024-12-11 Classification: LYASE Ligands: ZN, SO4 |
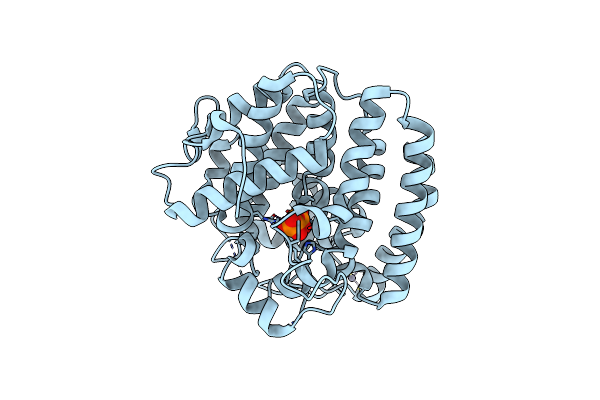 |
Crystal Structure Of Skaba3 From Shimazuella Kribbensis In Complex With Ppi
Organism: Shimazuella kribbensis
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2024-12-11 Classification: LYASE Ligands: ZN, MG, POP |
 |
Crystal Structure Of 2-Hydroxacyl-Coa Lyase/Synthase Apbhacs From Alphaproteobacteria Bacterium In The Complex With Thdp, L-Lactyl-Coa, And Adp
Organism: Alphaproteobacteria bacterium
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2024-10-02 Classification: LYASE Ligands: 8FL, UQ3, ADP, ACE, MG, GOL |
 |
Crystal Structure Of 2-Hydroxacyl-Coa Lyase/Synthase Apbhacs From Alphaproteobacteria Bacterium In The Complex With Thdp, D-Lactyl-Coa, And Adp
Organism: Alphaproteobacteria bacterium
Method: X-RAY DIFFRACTION Resolution:1.98 Å Release Date: 2024-10-02 Classification: LYASE Ligands: MG, 8FL, UQ3, ADP, EDO, ACY, PO4, CL, ACE, PEG |
 |
Crystal Structure Of 2-Hydroxyacyl-Coa Lyase/Synthse Apbhacs From Alphaproteobacteria Bacterium In The Complex With Thdp, Formyl-Coa, And Adp
Organism: Alphaproteobacteria bacterium
Method: X-RAY DIFFRACTION Resolution:1.72 Å Release Date: 2024-10-02 Classification: LYASE Ligands: A1AEK, FYN, MG, ADP, EDO, PO4, PEG |
 |
Crystal Structure Of 2-Hydroxyacyl-Coa Lyase/Synthase Apbhacs From Alphaproteobacteria Bacterium In The Complex With Thdp, Coenzyme A, And Adp
Organism: Alphaproteobacteria bacterium
Method: X-RAY DIFFRACTION Resolution:2.36 Å Release Date: 2024-10-02 Classification: LYASE Ligands: FYN, TPP, MG, ADP, EDO, SO4, CL, GOL |
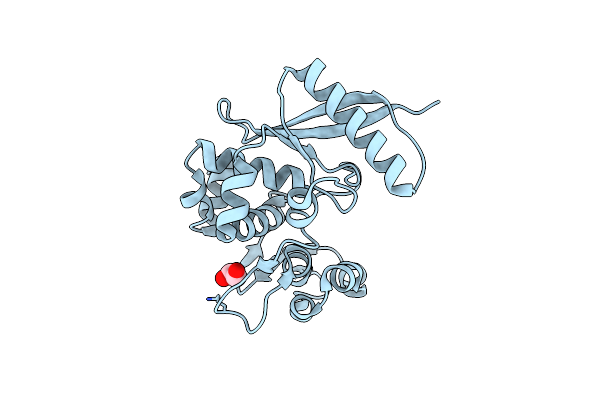 |
Organism: Trichlorobacter lovleyi
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2023-07-12 Classification: HYDROLASE Ligands: EDO, CL |
 |
Organism: Gammaproteobacteria bacterium
Method: X-RAY DIFFRACTION Resolution:2.63 Å Release Date: 2021-11-17 Classification: UNKNOWN FUNCTION Ligands: BTN, GOL |
 |
Organism: Gammaproteobacteria bacterium
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2021-11-17 Classification: UNKNOWN FUNCTION |
 |
Organism: Gammaproteobacteria bacterium
Method: X-RAY DIFFRACTION Resolution:2.18 Å Release Date: 2021-11-17 Classification: UNKNOWN FUNCTION Ligands: 1PE, P6G, PEG, PGE |
 |
Organism: Gammaproteobacteria bacterium
Method: X-RAY DIFFRACTION Resolution:1.67 Å Release Date: 2021-11-17 Classification: UNKNOWN FUNCTION Ligands: BTN, PG4, PGE, EDO, PEG, P33, NA |
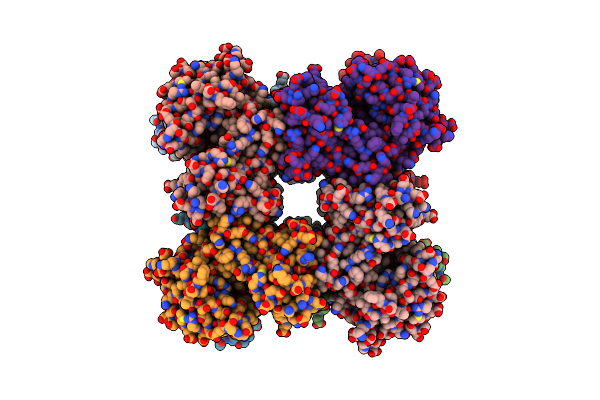 |
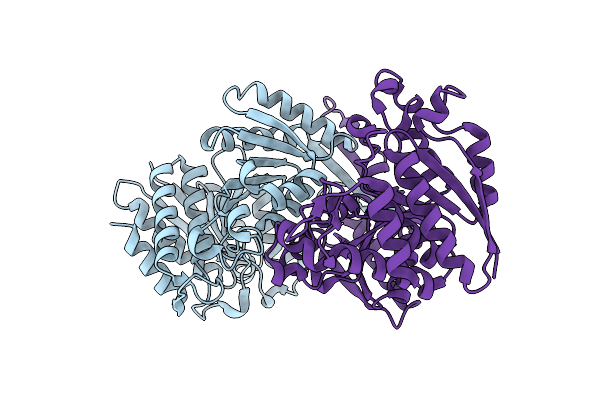
Structure Of A Bacillus Coagulans Polyol Dehydrogenase Double Mutant With An Acquired D-Lactate Dehydrogenase Activity
Organism: Bacillus coagulans
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2019-07-03 Classification: OXIDOREDUCTASE |
 |
The Crystal Structure Of 3-Isopropylmalate Dehydrogenase From Bacillus Coagulans
Organism: Bacillus coagulans
Method: X-RAY DIFFRACTION Resolution:2.85 Å Release Date: 2005-02-15 Classification: OXIDOREDUCTASE |
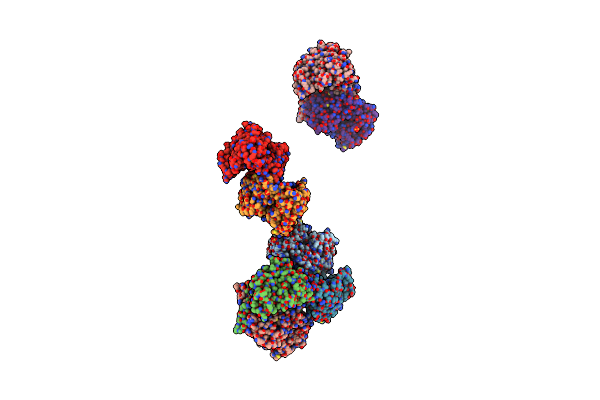 |
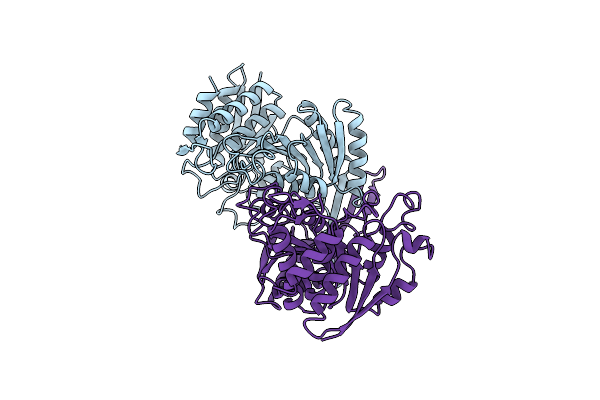
The Structure Of The Mutant, S225A And E251L, Of 3-Isopropylmalate Dehydrogenase From Bacillus Coagulans
Organism: Bacillus coagulans
Method: X-RAY DIFFRACTION Resolution:2.95 Å Release Date: 2005-02-15 Classification: OXIDOREDUCTASE Ligands: SO4 |
 |
3-Isopropylmalate Dehydrogenase From The Moderate Facultative Thermophile, Bacillus Coagulans
Organism: Bacillus coagulans
Method: X-RAY DIFFRACTION Resolution:3.00 Å Release Date: 1998-05-27 Classification: OXIDOREDUCTASE |
 |
Organism: Coleophora sp. BOLD:ABZ5590
Method: Alphafold Release Date: Classification: NA Ligands: NA |
 |

