Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
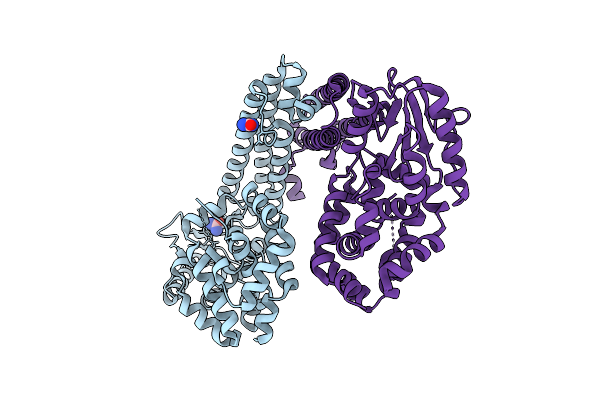 |
Organism: Gammaproteobacteria
Method: X-RAY DIFFRACTION Release Date: 2025-08-06 Classification: DNA BINDING PROTEIN Ligands: URE |
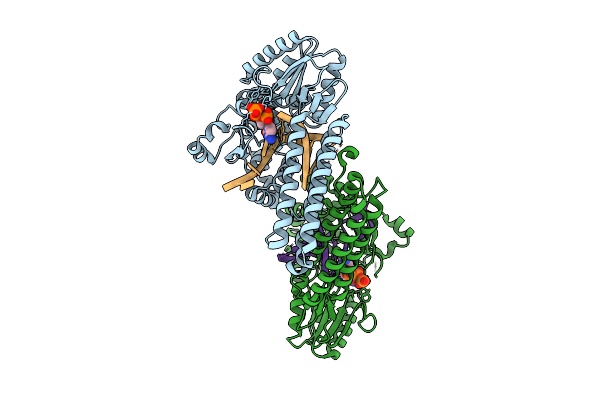 |
Organism: Gammaproteobacteria, Synthetic construct
Method: X-RAY DIFFRACTION Release Date: 2025-07-30 Classification: DNA BINDING PROTEIN Ligands: MN, DCP |
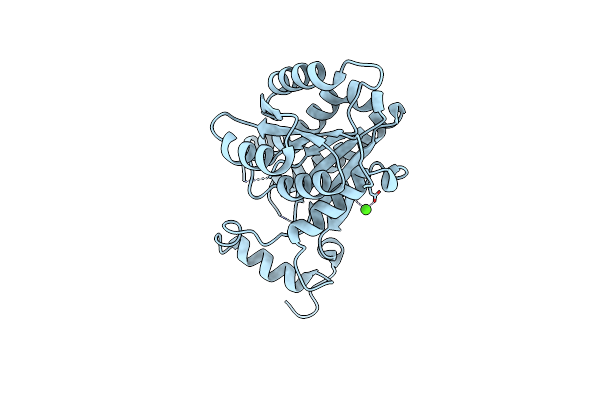 |
Crystal Structure Of The Apo Enoyl-Acp-Reductase (Fabi) From Moraxella Catarrhalis
Organism: Moraxella catarrhalis (strain bbh18)
Method: X-RAY DIFFRACTION Resolution:2.38 Å Release Date: 2022-07-20 Classification: OXIDOREDUCTASE Ligands: CA |
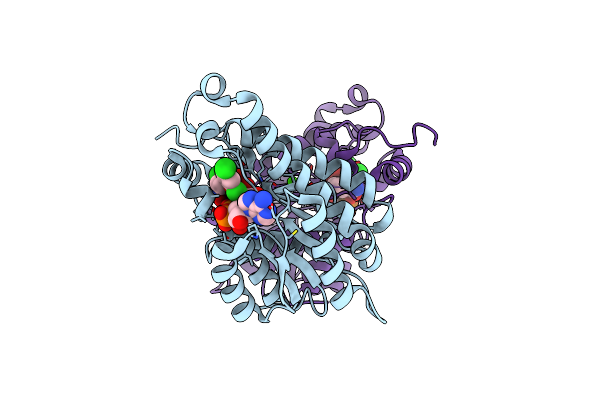 |
Crystal Structure Of Moraxella Catarrhalis Enoyl-Acp-Reductase (Fabi) In Complex With Nad And Triclosan
Organism: Moraxella catarrhalis (strain bbh18)
Method: X-RAY DIFFRACTION Resolution:2.22 Å Release Date: 2022-07-20 Classification: OXIDOREDUCTASE Ligands: NAD, TCL, CA |
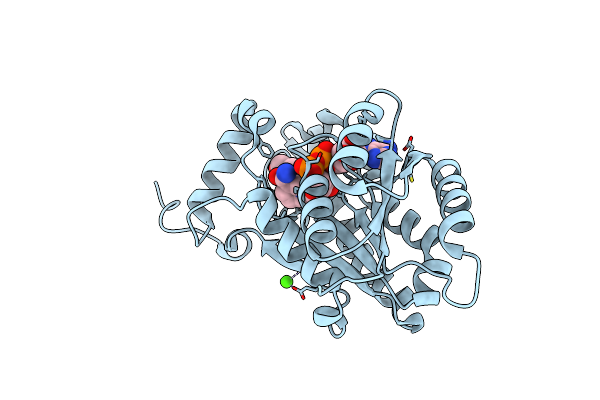 |
Crystal Structure Of Moraxella Catarrhalis Enoyl-Acp-Reductase (Fabi) In Complex With The Cofactor Nad
Organism: Moraxella catarrhalis (strain bbh18)
Method: X-RAY DIFFRACTION Resolution:2.12 Å Release Date: 2022-06-22 Classification: OXIDOREDUCTASE Ligands: NAD, CA, GOL |
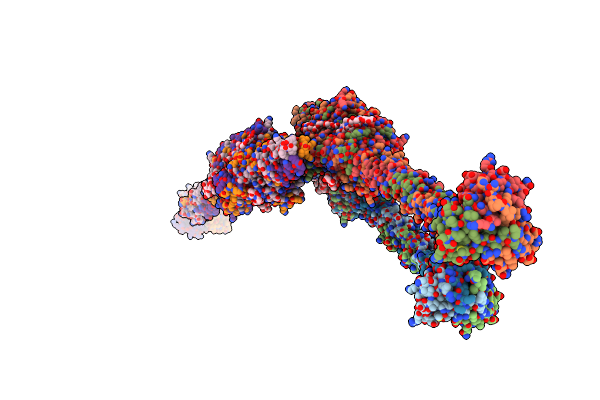 |
Crystal Structure Of Full-Length Streptococcal Bacteriophage Hyaluronidase In Complex With Unsaturated Hyaluronan Octa-Saccharides
Organism: Streptococcus pyogenes phage h4489a
Method: X-RAY DIFFRACTION Resolution:3.58 Å Release Date: 2021-06-09 Classification: LYASE |
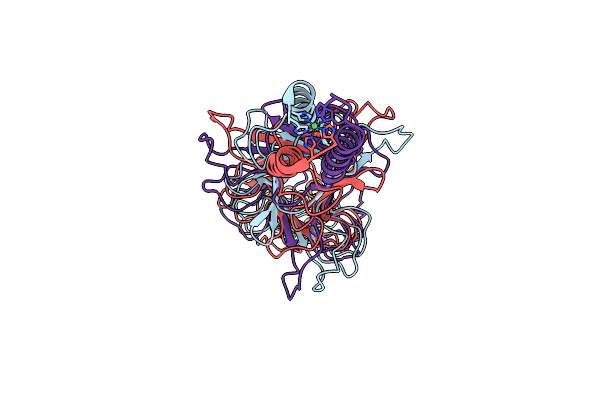 |
Crystal Structure Of Streptococcal Bacteriophage Hyaluronidase: Presence Of A Prokaryotic Collagen And Elucidation Of Catalytic Mechanism
Organism: Streptococcus pyogenes phage h4489a
Method: X-RAY DIFFRACTION Resolution:2.21 Å Release Date: 2021-05-12 Classification: LYASE Ligands: NI |
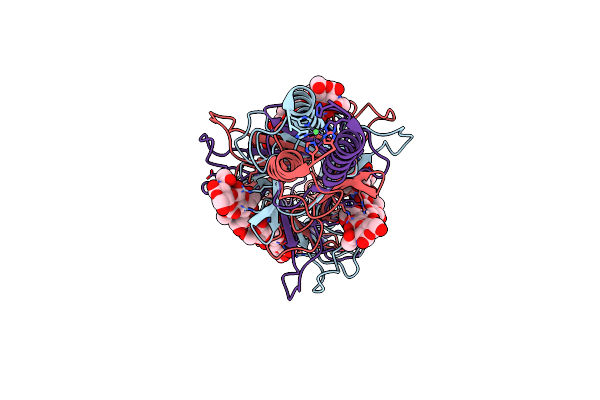 |
Crystal Structure Of Truncated Bacteriophage Hyaluronan Lyase Hylp In Complex With Unsaturated Hyaluronan Tetra-Saccharides
Organism: Streptococcus pyogenes phage h4489a
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2021-05-12 Classification: LYASE Ligands: NI |
 |
Crystal Structure Of Truncated Streptococcal Bacteriophage Hyaluronidase Complexed With Unsaturated Hyaluronan Hexa-Saccharides
Organism: Streptococcus pyogenes phage h4489a
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2021-05-12 Classification: LYASE Ligands: NI |
 |
Adenylation, Ketoreductase, And Pseudo Asub Multidomain Structure Of A Keto Acid-Selecting Nrps Module
Organism: Bacillus stratosphericus lama 585
Method: X-RAY DIFFRACTION Resolution:3.40 Å Release Date: 2020-02-19 Classification: BIOSYNTHETIC PROTEIN Ligands: CA, MG |
 |
Adenylation Domain Of A Keto Acid-Selecting Nrps Module Bound To Keto Acyl Adenylate Space Group P43212
Organism: Bacillus stratosphericus lama 585
Method: X-RAY DIFFRACTION Resolution:2.31 Å Release Date: 2020-02-19 Classification: BIOSYNTHETIC PROTEIN Ligands: QA7, SO4 |
 |
Adenylation Domain Of A Keto Acid-Selecting Nrps Module Bound To Keto Acyl Adenylate Space Group P212121
Organism: Bacillus stratosphericus lama 585
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2020-02-19 Classification: BIOSYNTHETIC PROTEIN Ligands: QA7, SO4, GOL |
 |
A Polyamorous Repressor: Deciphering The Evolutionary Strategy Used By The Phage-Inducible Chromosomal Islands To Spread In Nature.
Organism: Staphylococcus phage phi11, Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:2.52 Å Release Date: 2019-08-28 Classification: STRUCTURAL PROTEIN Ligands: MG, NI, PEG |
 |
Pyrrolysyl-Trna Synthetase From Canditatus Methanomethylophilus Alvus (Mmapylrs)
Organism: Candidatus methanomethylophilus alvus mx1201 1)
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2018-12-12 Classification: LIGASE |
 |
Crystal Structure Of Udp-N-Acetylglucosamine O-Acyltransferase (Lpxa) From Moraxella Catarrhalis Rh4.
Organism: Moraxella catarrhalis (strain bbh18)
Method: X-RAY DIFFRACTION Resolution:3.00 Å Release Date: 2017-06-14 Classification: TRANSFERASE Ligands: GOL, FLC |
 |
Organism: Staphylococcus phage phi11
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2016-05-25 Classification: STRUCTURAL PROTEIN Ligands: FE |
 |
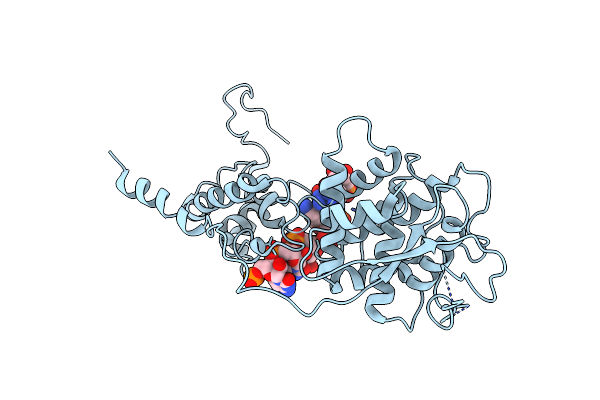
The 3D Structure Of D95N Mutant Dutpase From Phage Phi11 Of S. Aureus Reveals The Molecular Details For The Coordination Of A Structural Mg(Ii) Ion
Organism: Staphylococcus phage phi11
Method: X-RAY DIFFRACTION Resolution:2.90 Å Release Date: 2015-12-23 Classification: HYDROLASE Ligands: DUP, MG |
 |
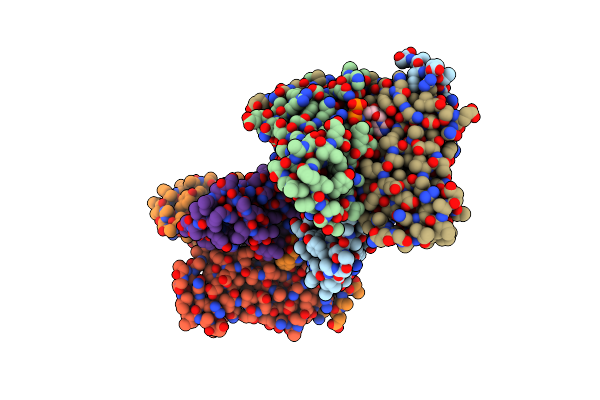
Organism: Measles virus strain halle, Spodoptera frugiperda
Method: ELECTRON MICROSCOPY Resolution:4.30 Å Release Date: 2015-04-29 Classification: RNA BINDING PROTEIN |
 |
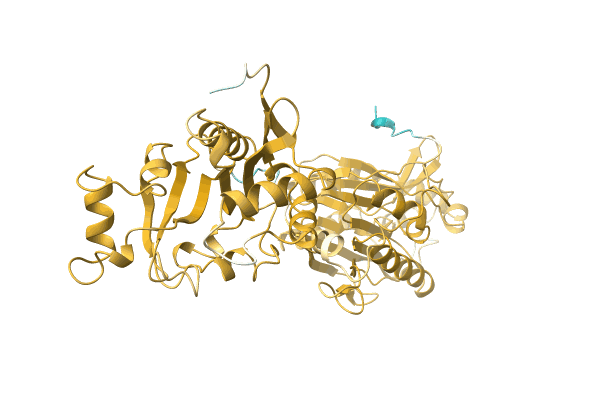
Dutpase From Phage Phi11 Of S.Aureus: Visualization Of The Species-Specific Insert
Organism: Staphylococcus phage 11
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2013-09-11 Classification: HYDROLASE Ligands: DUP, MG |
 |
Organism: Geofilum rubicundum JCM 15548
Method: Alphafold Release Date: Classification: NA Ligands: NA |