Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
 |
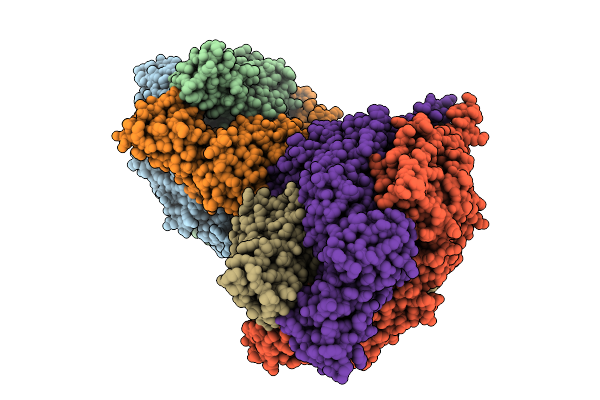
Cyro-Em Structure Of Prefusion Rsv Fusion Glycoprotein In Complex With Ziresovir And Motavizumab Fab
Organism: Human respiratory syncytial virus a2, Tequatrovirus t4, Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-01-14 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: A1EV1 |
 |
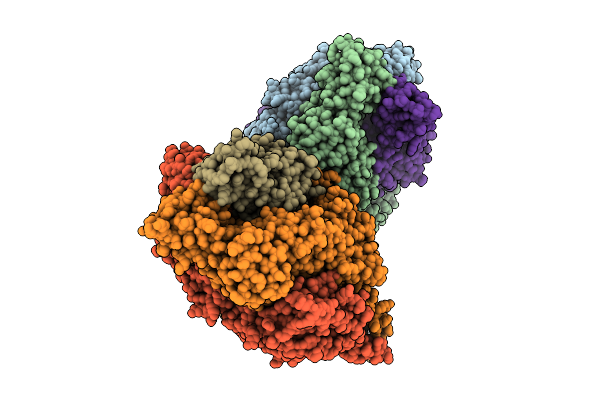
Organism: Human respiratory syncytial virus, Enterobacteria phage t6
Method: X-RAY DIFFRACTION Release Date: 2025-12-17 Classification: VIRAL PROTEIN |
 |
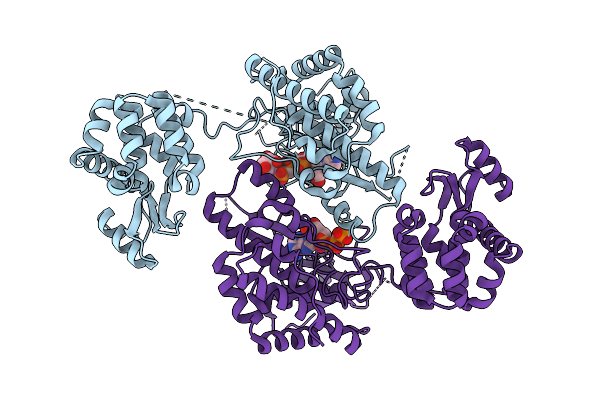
Organism: Respiratory syncytial virus, Enterobacteria phage t6
Method: X-RAY DIFFRACTION Release Date: 2025-12-17 Classification: VIRAL PROTEIN |
 |
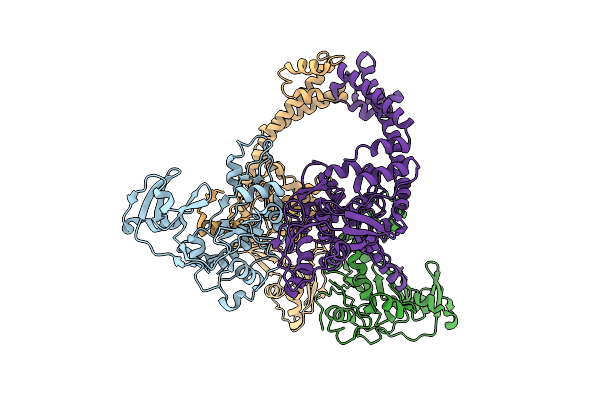
Crystal Structure Of Phosphatidyl Inositol 4-Kinase Ii Beta In Complex With Hh5129
Organism: Homo sapiens, Enterobacteria phage t4
Method: X-RAY DIFFRACTION Release Date: 2025-12-03 Classification: TRANSFERASE Ligands: A1IVA |
 |
Organism: Enterobacteria phage t4
Method: ELECTRON MICROSCOPY Release Date: 2025-11-26 Classification: ISOMERASE |
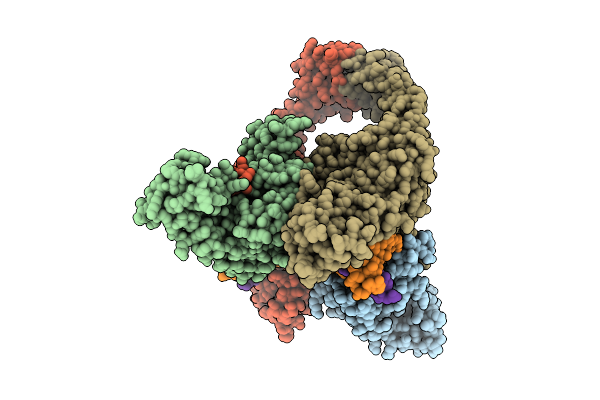 |
Organism: Escherichia phage t4, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-10-22 Classification: ISOMERASE Ligands: MG |
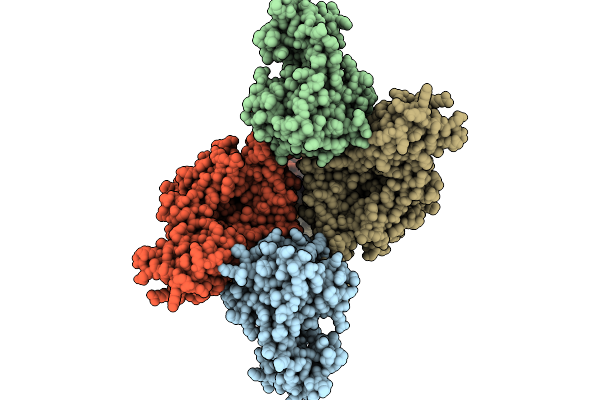 |
Organism: Escherichia phage t4, Dna molecule
Method: ELECTRON MICROSCOPY Release Date: 2025-10-01 Classification: ISOMERASE/DNA |
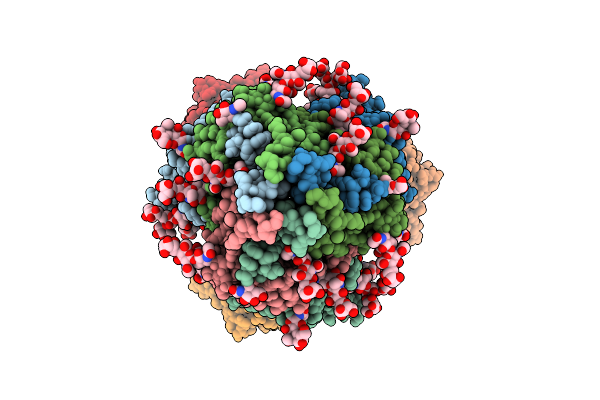 |
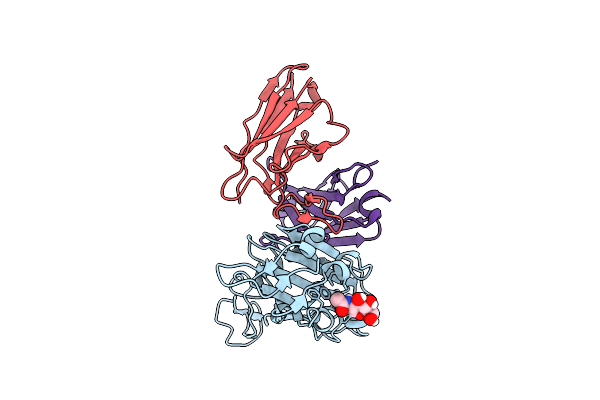
Pre-Fusion Herv-K Envelope Protein Trimer Ectodomain In Complex With Kenv-6 Fab
Organism: Homo sapiens, Enterobacteria phage t4, Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2025-07-30 Classification: VIRAL PROTEIN Ligands: NAG |
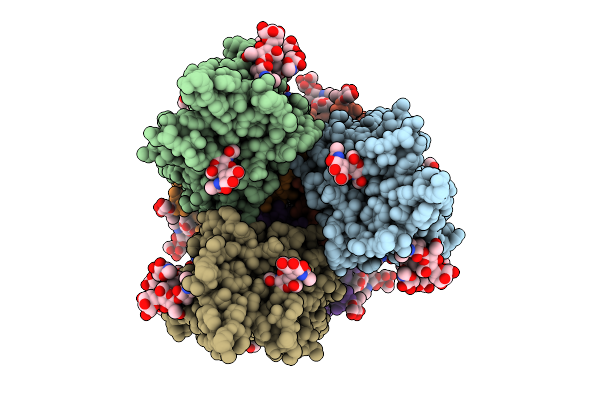 |
Organism: Homo sapiens, Enterobacteria phage t4
Method: ELECTRON MICROSCOPY Release Date: 2025-07-30 Classification: VIRAL PROTEIN Ligands: NAG |
 |
Organism: Severe acute respiratory syndrome coronavirus 2, Tequatrovirus t4, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-07-23 Classification: PROTEIN BINDING Ligands: NAG |
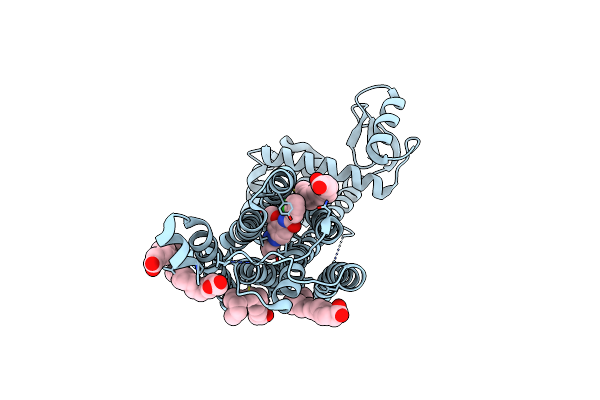 |
Dark Structure Of The Human Metabotropic Glutamate Receptor 5 Transmembrane Domain Bound To Photoswitchable Ligand Alloswitch-1
Organism: Homo sapiens, Enterobacteria phage t4
Method: X-RAY DIFFRACTION Release Date: 2025-06-25 Classification: SIGNALING PROTEIN Ligands: OLA, 4YI |
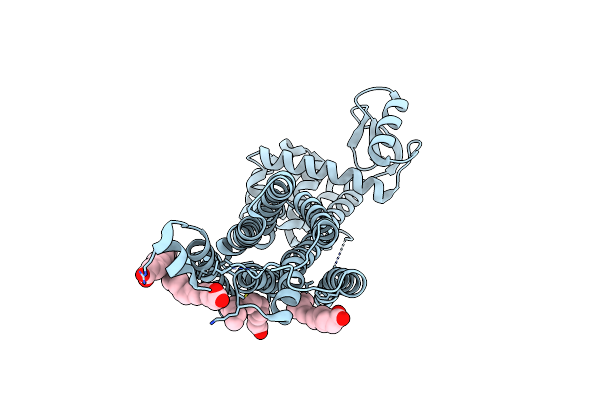 |
Apo-State Structure Of The Human Metabotropic Glutamate Receptor 5 Transmembrane Domain Freeze-Trapped After Light Activation Of Photoswitchable Ligand Alloswitch-1
Organism: Homo sapiens, Enterobacteria phage t4
Method: X-RAY DIFFRACTION Release Date: 2025-06-25 Classification: SIGNALING PROTEIN Ligands: OLA |
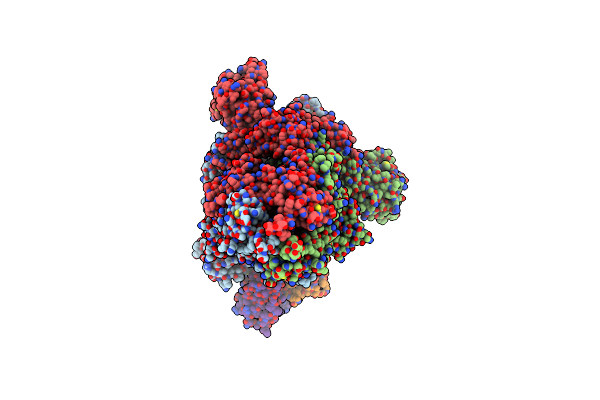 |
Organism: Severe acute respiratory syndrome coronavirus 2, Enterobacteria phage t4, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-06-18 Classification: VIRAL PROTEIN |
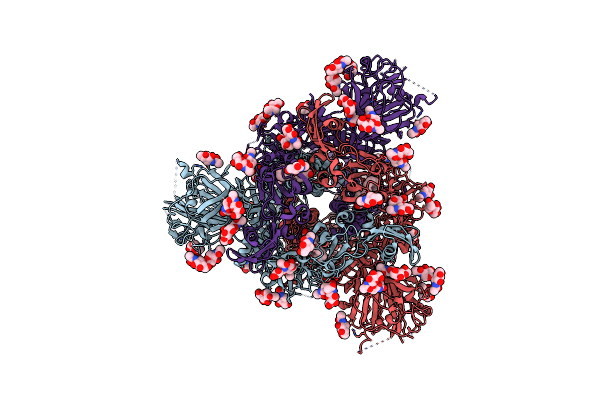 |
Organism: Coronavirus neoromicia/pml-phe1/rsa/2011, Enterobacteria phage t4
Method: ELECTRON MICROSCOPY Release Date: 2025-06-18 Classification: VIRAL PROTEIN Ligands: NAG, EIC |
 |
Cryo-Em Structure Of Eu-Hedgehogcov (Erinaceus/Vmc/Deu/2012) S-Trimer In A Locked-2 Conformation
Organism: Betacoronavirus erinaceus/vmc/deu/2012, Enterobacteria phage t4 (bacteriophage t4)
Method: ELECTRON MICROSCOPY Release Date: 2025-06-18 Classification: VIRAL PROTEIN Ligands: NAG, FOL, EIC |
 |
Cryo-Em Structure Of Hku25-Batcov S-Trimer Stabilized With 2P And X1 Disulfide Bond
Organism: Hypsugo bat coronavirus hku25, Tequatrovirus t4
Method: ELECTRON MICROSCOPY Release Date: 2025-06-18 Classification: VIRAL PROTEIN Ligands: EIC, NAG |
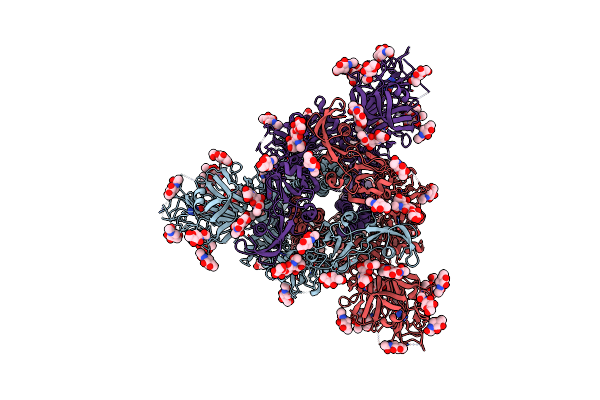 |
Cryo-Em Structure Of Cn-Hedgehogcov (Hku31/Erinaceus Amurensis/China/2014) S-Trimer In A Locked-2 Conformation
Organism: Erinaceus hedgehog coronavirus hku31, Enterobacteria phage t4
Method: ELECTRON MICROSCOPY Release Date: 2025-06-18 Classification: VIRAL PROTEIN Ligands: FOL, NAG |
 |
Cryo-Em Structure Of Cn-Hedgehogcov (Hku31/Erinaceus Amurensis/China/2014) S-Trimer In A Locked-1 Conformation
Organism: Erinaceus hedgehog coronavirus hku31, Tequatrovirus t4
Method: ELECTRON MICROSCOPY Release Date: 2025-06-18 Classification: VIRAL PROTEIN Ligands: NAG, FOL, EIC |
 |
Organism: Bat coronavirus, Tequatrovirus t4
Method: ELECTRON MICROSCOPY Release Date: 2025-06-18 Classification: VIRAL PROTEIN Ligands: NAG, EIC |
 |
Organism: Middle east respiratory syndrome-related coronavirus, Tequatrovirus t4
Method: ELECTRON MICROSCOPY Release Date: 2025-06-18 Classification: VIRAL PROTEIN Ligands: NAG, EIC |