Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
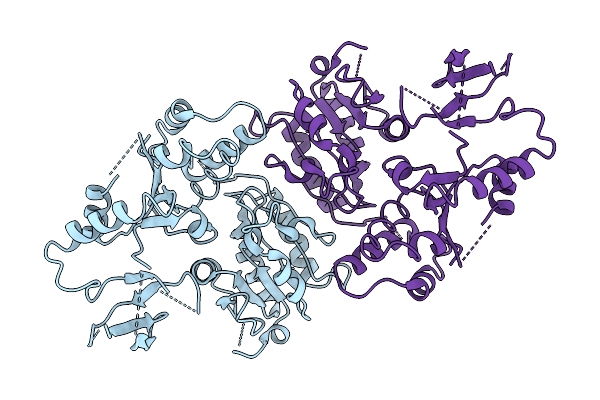 |
Organism: Leptospirillum ferriphilum
Method: ELECTRON MICROSCOPY Release Date: 2025-12-03 Classification: DNA BINDING PROTEIN |
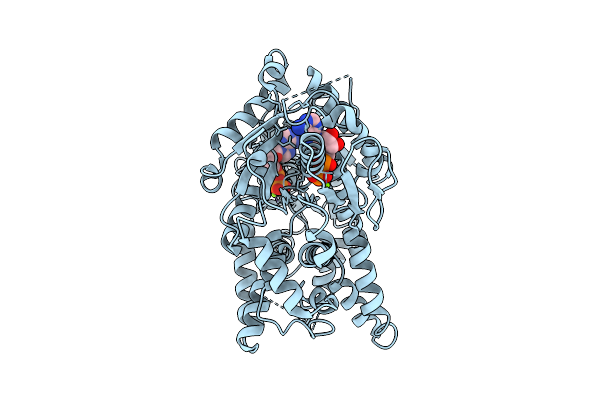 |
Open Conformation Of Arsa From L. Ferriphilum In Complex With Mgadp Determined In The Presence Of Arsenite
Organism: Leptospirillum ferriphilum
Method: ELECTRON MICROSCOPY Release Date: 2025-08-20 Classification: HYDROLASE Ligands: ADP, MG |
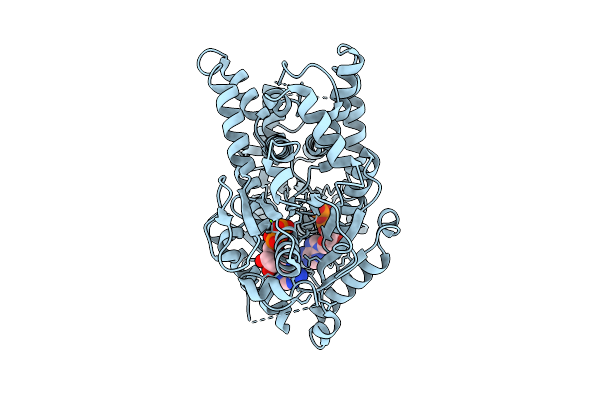 |
Open Conformation Of Arsa From L. Ferriphilum In Complex With Mgadp Determined In Absence Of Arsenite
Organism: Leptospirillum ferriphilum
Method: ELECTRON MICROSCOPY Release Date: 2025-08-20 Classification: HYDROLASE Ligands: ADP, MG |
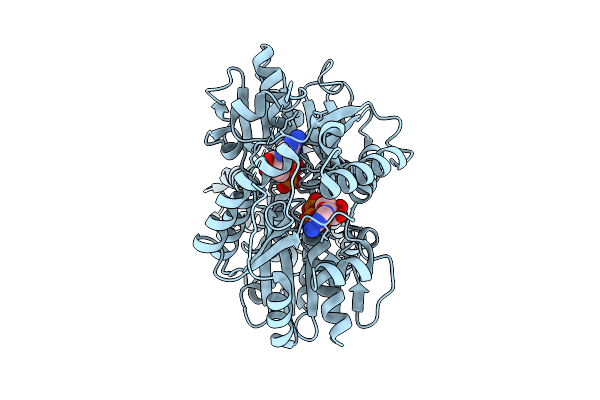 |
Closed Conformation Of Arsa From L. Ferriphilum In Complex With Mgatp And Arsenite
Organism: Leptospirillum ferriphilum
Method: ELECTRON MICROSCOPY Release Date: 2025-08-20 Classification: HYDROLASE Ligands: ATP, MG, ARS |
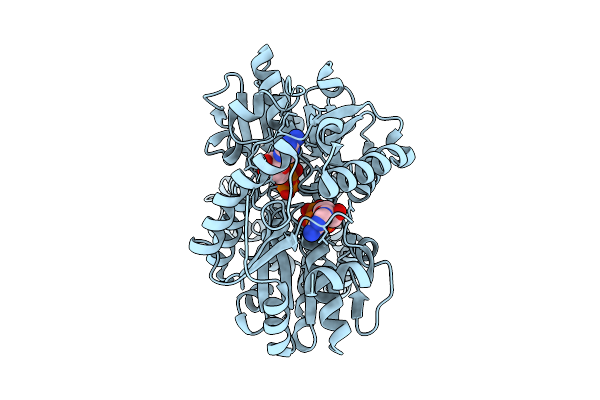 |
Closed Conformation Of Arsa From L. Ferriphilum In Complex With Mgatp And Arsenite At 1.5 Minute Time Point
Organism: Leptospirillum ferriphilum ml-04
Method: ELECTRON MICROSCOPY Release Date: 2025-08-20 Classification: HYDROLASE Ligands: ATP, MG, ARS |
 |
Organism: Spirochaeta thermophila dsm 6578, Corynebacterium pollutisoli
Method: X-RAY DIFFRACTION Resolution:3.34 Å Release Date: 2024-03-27 Classification: TRANSPORT PROTEIN Ligands: THJ, OLC |
 |
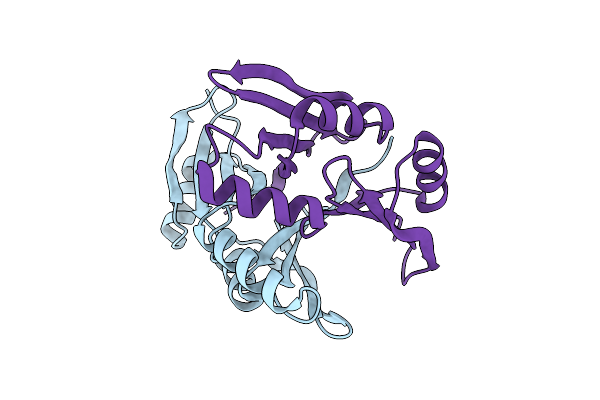
Organism: Spirochaeta thermophila dsm 6578, Corynebacterium pollutisoli
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2024-03-27 Classification: TRANSPORT PROTEIN Ligands: OLC |
 |
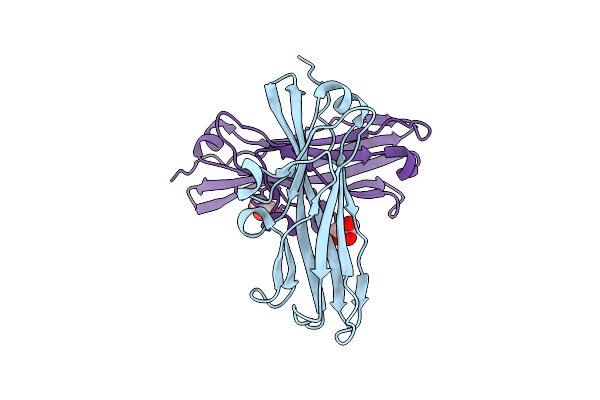
Organism: Streptomyces uncialis
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2019-03-06 Classification: ANTITOXIN |
 |
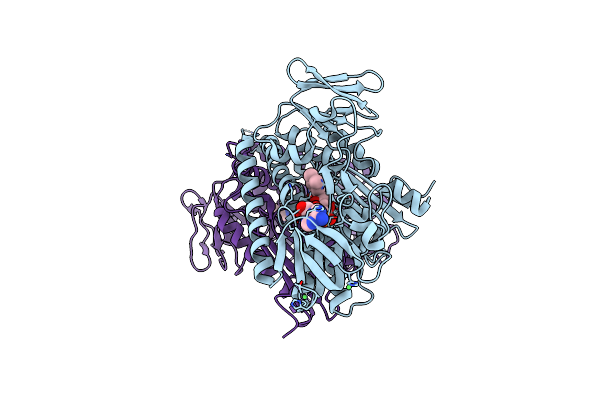
Crystal Structure Of Agga, The Major Subunit Of Aggregative Adherence Fimbriae Type I (Aaf/I) From The Escherichia Coli O4H104
Organism: Escherichia coli o104:h4 str. c227-11
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2014-10-01 Classification: CELL ADHESION Ligands: GOL |
 |
Organism: Marichromatium gracile
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2008-02-19 Classification: OXIDOREDUCTASE Ligands: FAD, CL, NI |
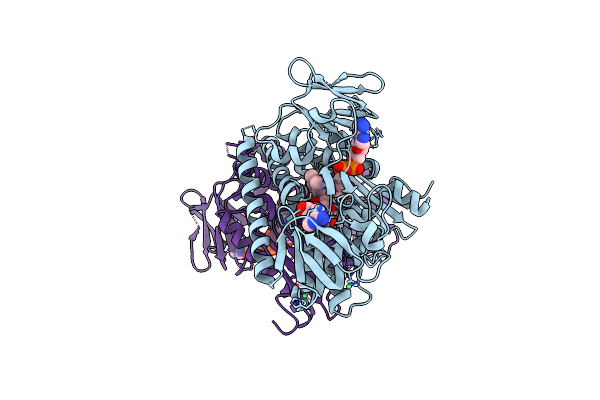 |
Structure Of Glutathione Amide Reductase From Chromatium Gracile In Complex With Nad
Organism: Marichromatium gracile
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2008-02-19 Classification: OXIDOREDUCTASE Ligands: FAD, NAD, CL, NI |
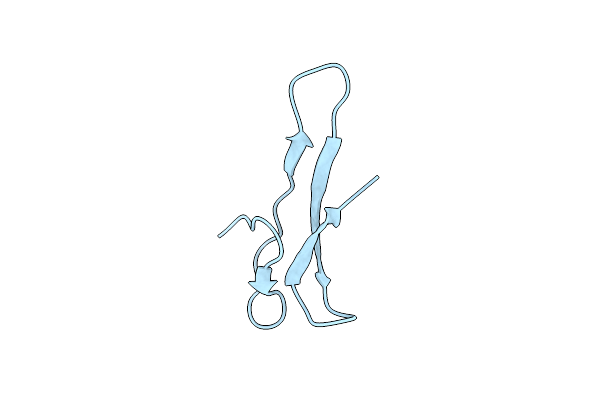 |
Nmr Structure Of The Carboxyl-Terminal Cysteine Domain Of The Vhv1.1 Polydnaviral Gene Product
Organism: Campoletis sonorensis ichnovirus
Method: SOLUTION NMR Release Date: 2004-10-05 Classification: VIRAL PROTEIN |
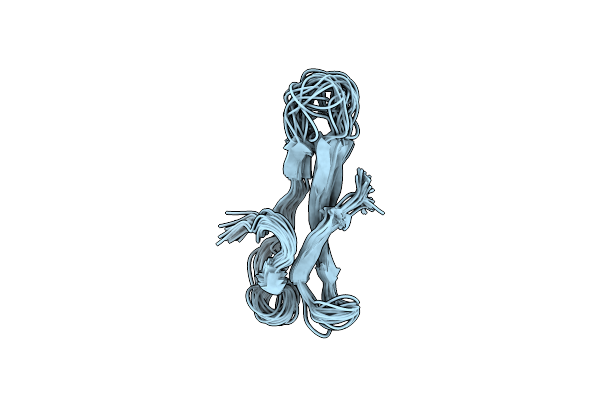 |
3D Solution Structure Of The C-Terminal Cysteine-Rich Domain Of The Vhv1.1 Polydnaviral Gene Product
Organism: Campoletis sonorensis ichnovirus
Method: SOLUTION NMR Release Date: 2004-10-05 Classification: VIRAL PROTEIN |
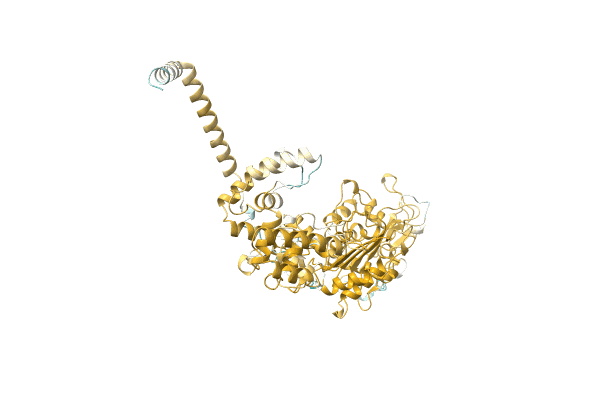 |
 |
 |
 |
 |
 |
 |