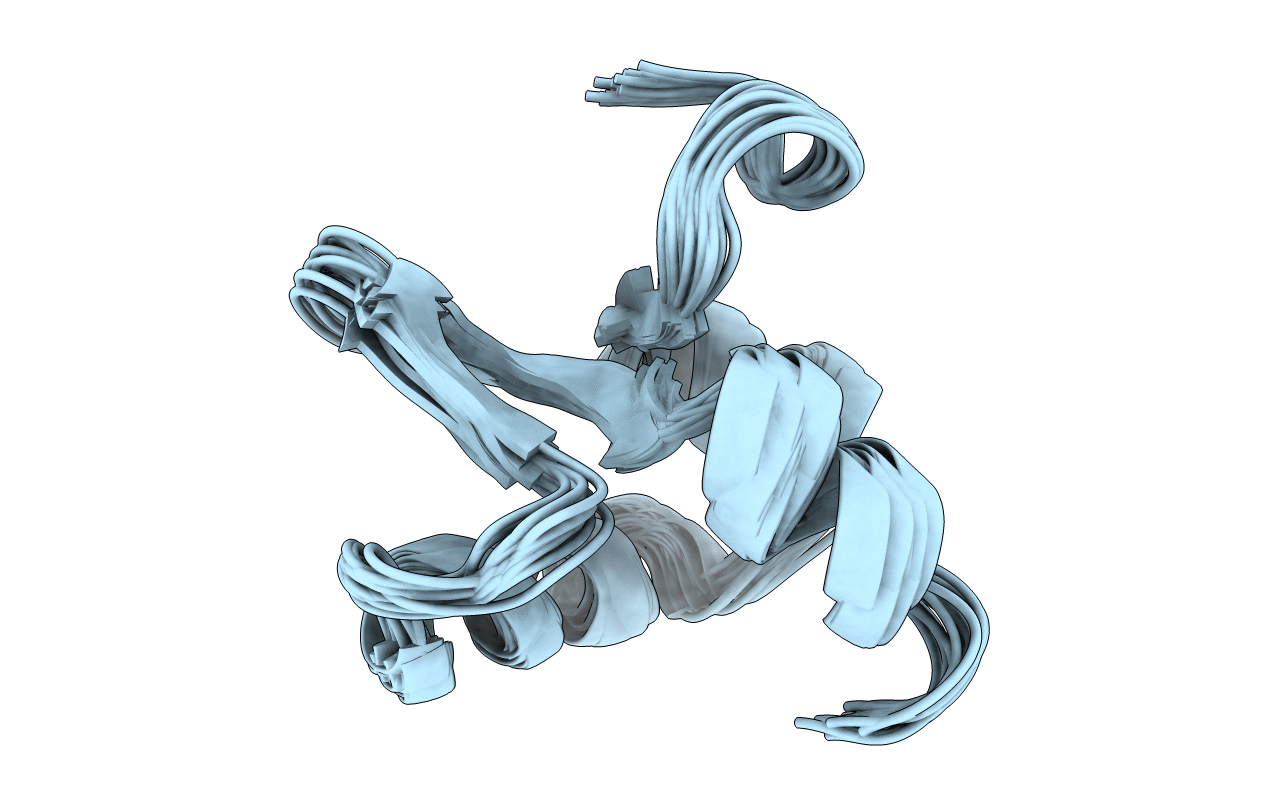
Deposition Date
2004-12-07
Release Date
2005-03-15
Last Version Date
2024-05-22
Entry Detail
PDB ID:
1Y6U
Keywords:
Title:
The Structure of the Excisionase (Xis) Protein from Conjugative Transposon Tn916 Provides Insights into the Regulation of Heterobivalent Tyrosine Recombinases
Biological Source:
Source Organism:
Enterococcus faecalis (Taxon ID: 1351)
Host Organism:
Method Details:
Experimental Method:
Conformers Calculated:
200
Conformers Submitted:
20
Selection Criteria:
structures with the lowest energy