Search Count: 18
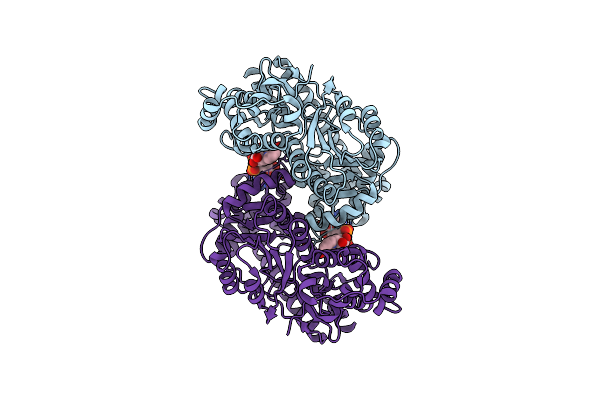 |
Organism: Pseudomonas fluorescens
Method: ELECTRON MICROSCOPY Release Date: 2024-07-17 Classification: LYASE Ligands: NAD |
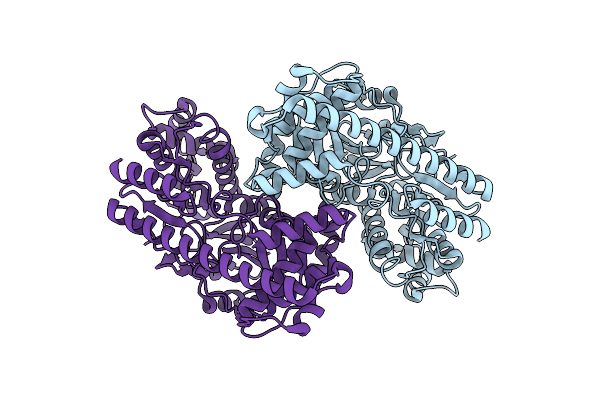 |
Organism: Pseudomonas fluorescens
Method: ELECTRON MICROSCOPY Release Date: 2024-07-17 Classification: LYASE |
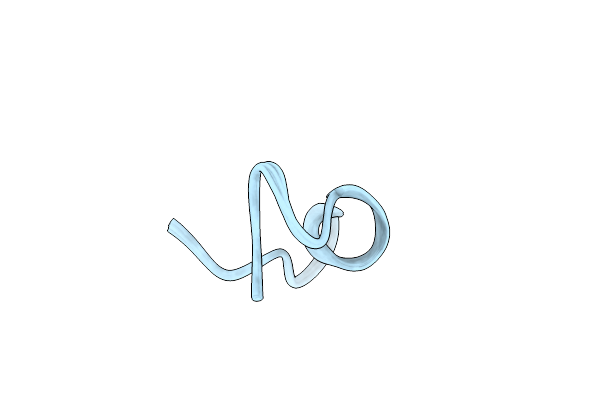 |
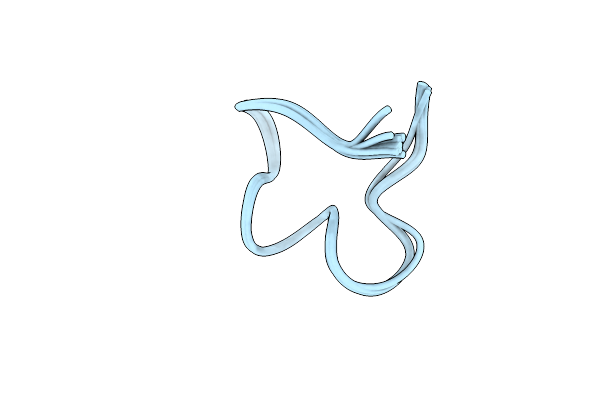
The Nmr Structure Of Noursin, A Tricyclic Ribosomal Peptide Containing A Histidine-To-Butyrine Crosslink
Organism: Streptomyces noursei atcc 11455
Method: SOLUTION NMR Release Date: 2023-05-31 Classification: UNKNOWN FUNCTION |
 |
Organism: Streptomyces noursei atcc 11455
Method: SOLUTION NMR Release Date: 2023-05-31 Classification: UNKNOWN FUNCTION |
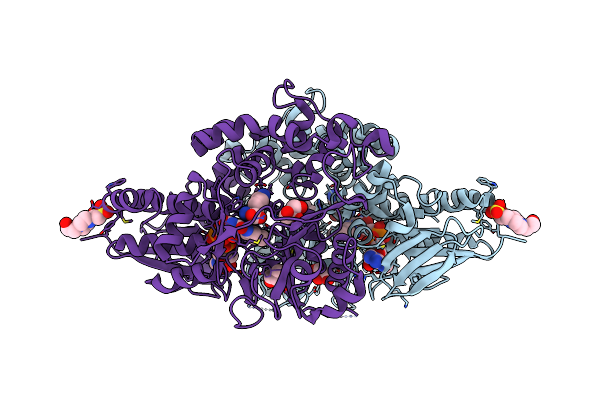 |
Organism: Hypocrea rufa
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2020-09-23 Classification: OXIDOREDUCTASE Ligands: FAD, LYS, GOL, EPE |
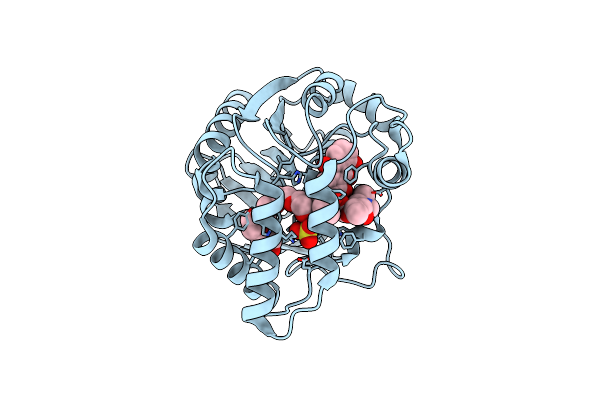 |
Crystal Structure Of Aclacinomycin Methylesterase (Rdmc) With Bound Product Analogue, 10-Decarboxymethylaclacinomycin T (Dcmat)
Organism: Streptomyces purpurascens
Method: X-RAY DIFFRACTION Resolution:1.45 Å Release Date: 2003-11-25 Classification: HYDROLASE Ligands: AKT, SO4, 1PE |
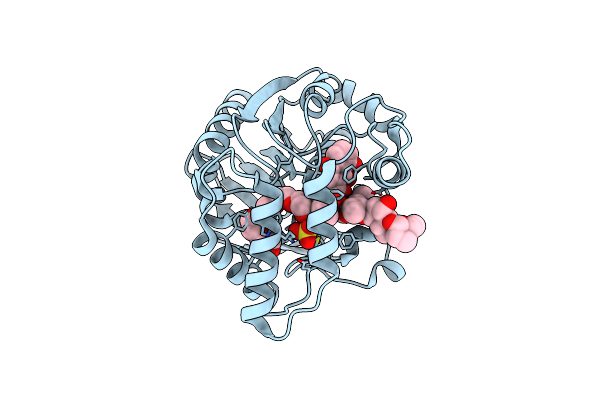 |
Crystal Structure Of Aclacinomycin Methylesterase (Rdmc) With Bound Product Analogue, 10-Decarboxymethylaclacinomycin A (Dcma)
Organism: Streptomyces purpurascens
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2003-11-25 Classification: HYDROLASE Ligands: AKA, SO4, 1PE |
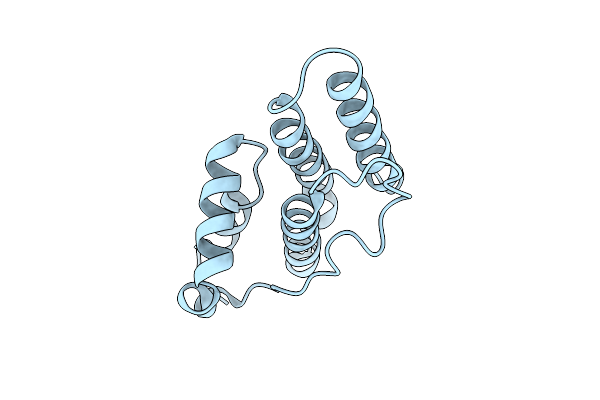 |
Organism: Streptomyces violaceoruber
Method: X-RAY DIFFRACTION Resolution:1.05 Å Release Date: 2003-06-10 Classification: HYDROLASE |
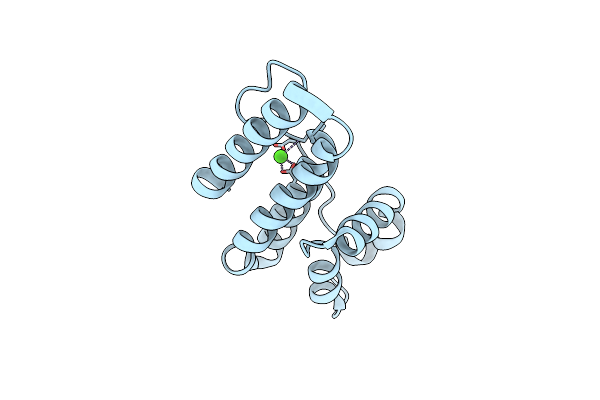 |
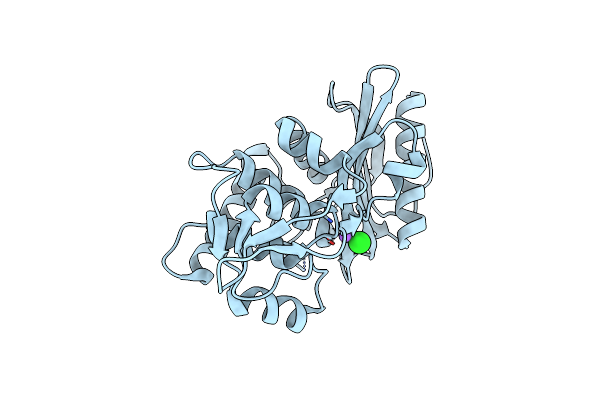
Organism: Streptomyces violaceoruber
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2002-09-04 Classification: HYDROLASE Ligands: CA |
 |
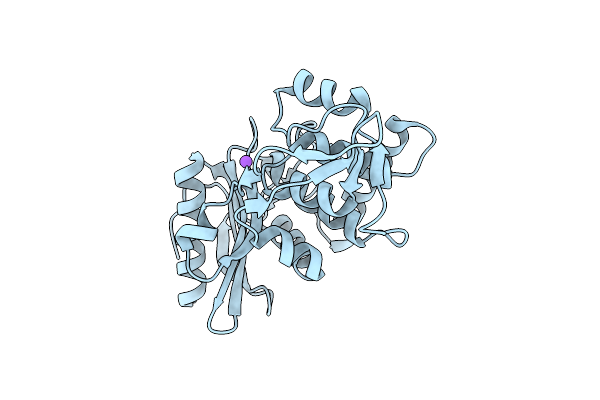
Organism: Streptomyces sp.
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2001-06-13 Classification: HYDROLASE |
 |
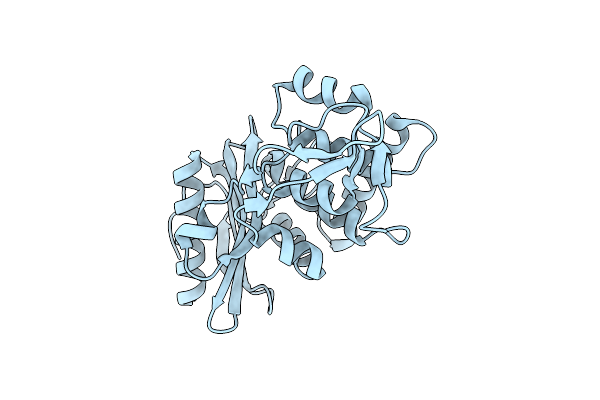
Organism: Streptomyces sp.
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2000-05-03 Classification: HYDROLASE |
 |
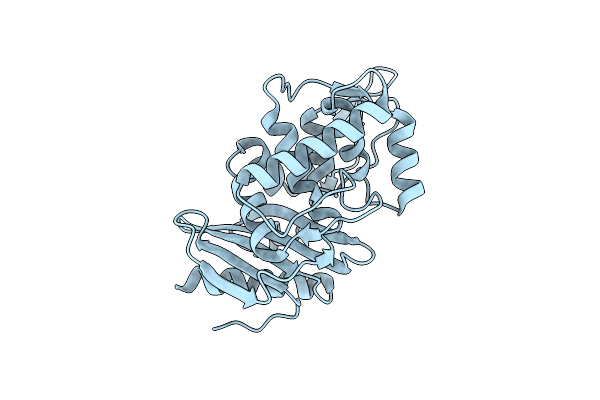
Organism: Streptomyces sp.
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2000-05-03 Classification: HYDROLASE |
 |
Organism: Streptomyces sp.
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2000-05-03 Classification: HYDROLASE |
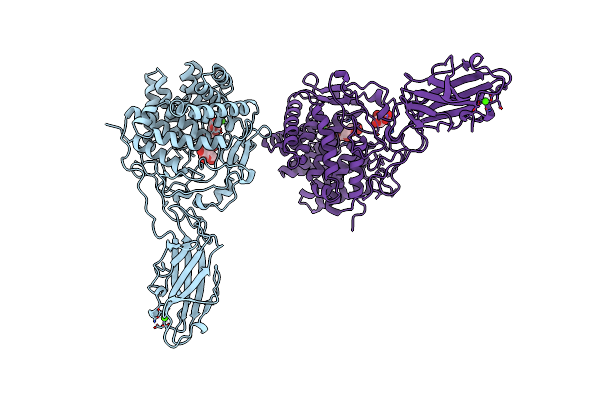 |
Organism: Thermobifida fusca
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 1997-09-17 Classification: GLYCOSYL HYDROLASE Ligands: CA, BGC |
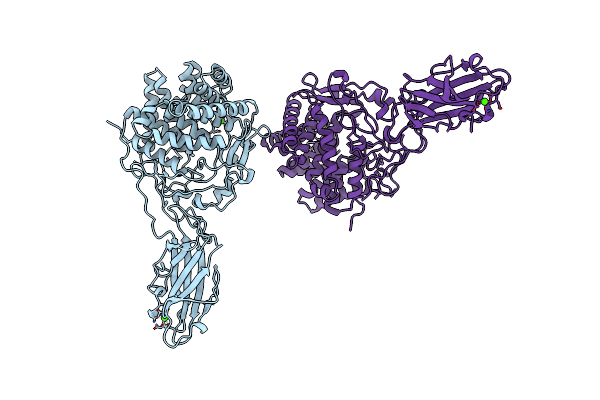 |
Organism: Thermobifida fusca
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 1997-09-04 Classification: GLYCOSYL HYDROLASE Ligands: CA |
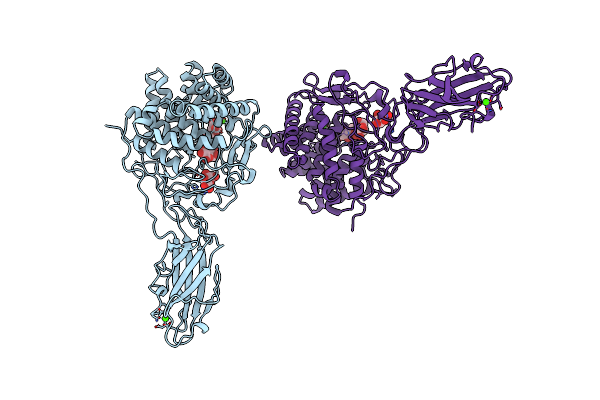 |
Organism: Thermobifida fusca
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 1997-09-04 Classification: GLYCOSYL HYDROLASE Ligands: BGC, CA |
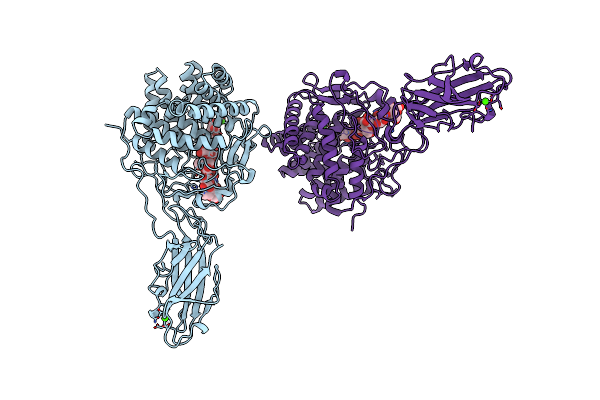 |
Organism: Thermobifida fusca
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 1997-09-04 Classification: GLYCOSYL HYDROLASE Ligands: CA |
 |
Acidothermus Cellulolyticus Endocellulase E1 Catalytic Domain In Complex With A Cellotetraose
Organism: Acidothermus cellulolyticus
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 1996-10-14 Classification: GLYCOSYL HYDROLASE |

