Search Count: 33
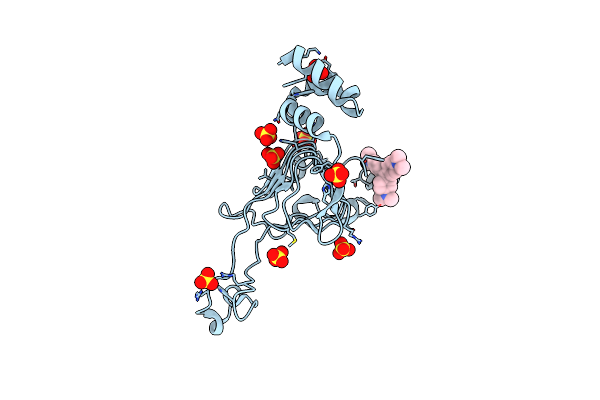 |
Crystal Structure Of The Type B Chloramphenicol Acetyltransferase From Vibrio Cholerae In The Complex With Crystal Violet
Organism: Vibrio cholerae serotype o1 (strain atcc 39315 / el tor inaba n16961)
Method: X-RAY DIFFRACTION Resolution:2.43 Å Release Date: 2019-09-25 Classification: TRANSFERASE Ligands: CVI, SO4, CL |
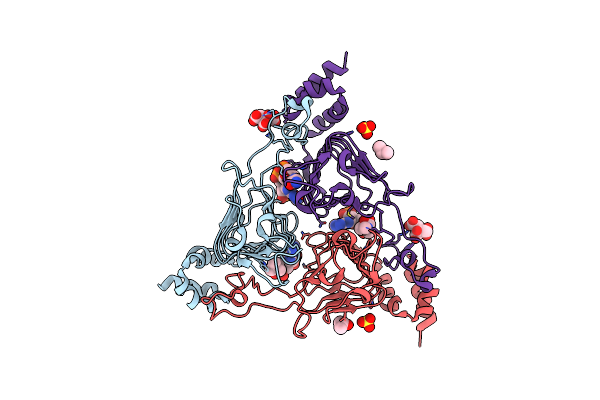 |
The 2.2 A Crystal Structure Of The Type B Chloramphenicol Acetyltransferase From Vibrio Cholerae In The Complex With Acetyl Coa
Organism: Vibrio cholerae serotype o1 (strain atcc 39315 / el tor inaba n16961)
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2019-09-18 Classification: TRANSFERASE Ligands: CIT, CL, ACO, SO4, IPA |
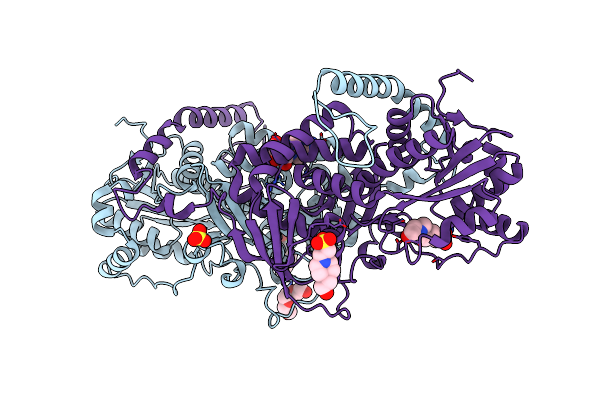 |
2.15 Angstrom Resolution Crystal Structure Of Argininosuccinate Synthase From Bordetella Pertussis
Organism: Bordetella pertussis (strain tohama i / atcc baa-589 / nctc 13251)
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2017-03-01 Classification: HYDROLASE,OXIDOREDUCTASE Ligands: EPE, PGE, ADN, SO4 |
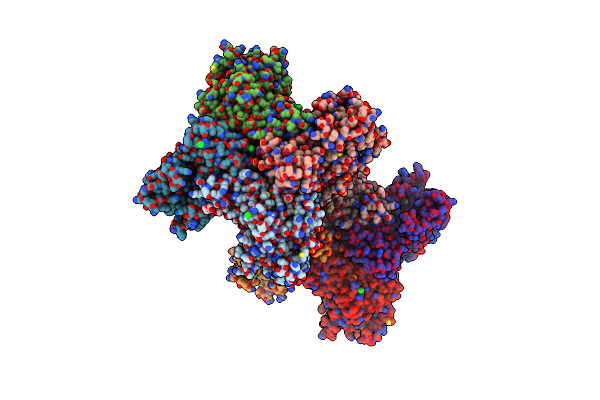 |
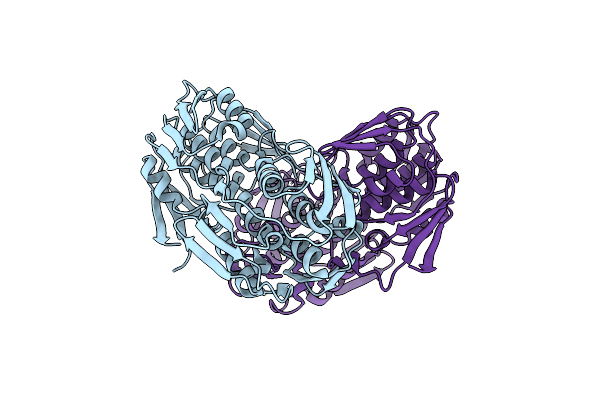
2.65 Angstrom Resolution Crystal Structure Of S-Adenosylhomocysteinase From Cryptosporidium Parvum In Complex With Sah And Nad
Organism: Cryptosporidium parvum (strain iowa ii)
Method: X-RAY DIFFRACTION Resolution:2.65 Å Release Date: 2017-03-01 Classification: HYDROLASE,OXIDOREDUCTASE Ligands: NAD, SAH, CL, SO4, P33, P6G, ADN, PEG |
 |
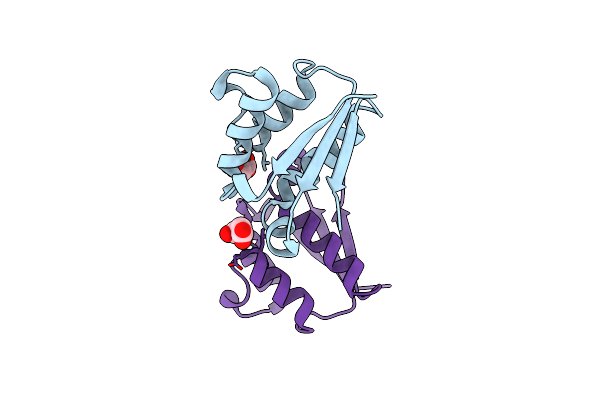
2.45 Angstrom Resolution Crystal Structure Of Udp-N-Acetylglucosamine 1-Carboxyvinyltransferase From Campylobacter Jejuni.
Organism: Campylobacter jejuni subsp. jejuni serotype o:2 (strain atcc 700819 / nctc 11168)
Method: X-RAY DIFFRACTION Resolution:2.46 Å Release Date: 2017-02-01 Classification: HYDROLASE,OXIDOREDUCTASE Ligands: CL |
 |
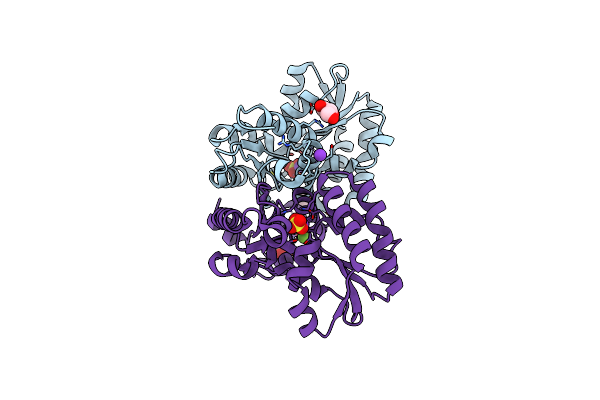
Crystal Structure Of Ribonuclease Inhibitor Barstar From Salmonella Typhimurium
Organism: Salmonella typhimurium (strain lt2 / sgsc1412 / atcc 700720)
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2015-12-23 Classification: HYDROLASE INHIBITOR Ligands: MLI |
 |
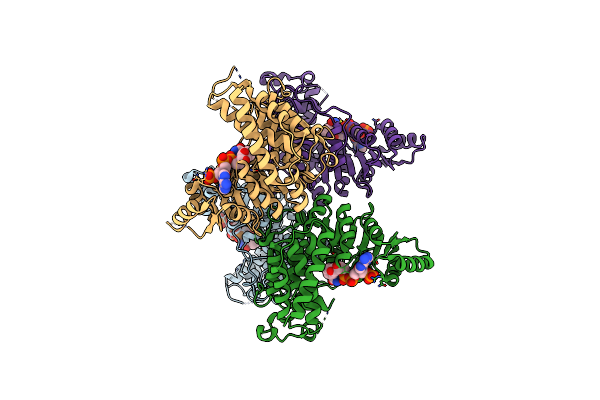
Crystal Structure Of Putative Aspartate Racemase From Salmonella Typhimurium Complexed With Sulfate And Potassium
Organism: Salmonella typhimurium
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2015-12-02 Classification: ISOMERASE Ligands: K, SO4, GOL, FMT, CL, F |
 |
1-Deoxy-D-Xylulose 5-Phosphate Reductoisomerase From Yersinia Pestis In Complex With Nadph
Organism: Yersinia pestis
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2015-09-30 Classification: OXIDOREDUCTASE Ligands: NDP |
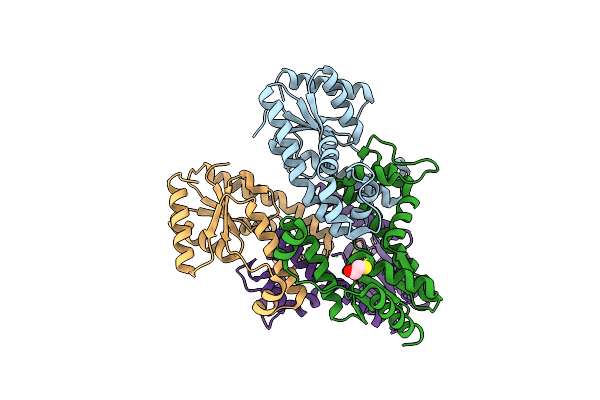 |
Organism: Campylobacter jejuni subsp. jejuni serotype o:2 (strain nctc 11168)
Method: X-RAY DIFFRACTION Resolution:2.57 Å Release Date: 2015-05-13 Classification: TRANSFERASE Ligands: BME |
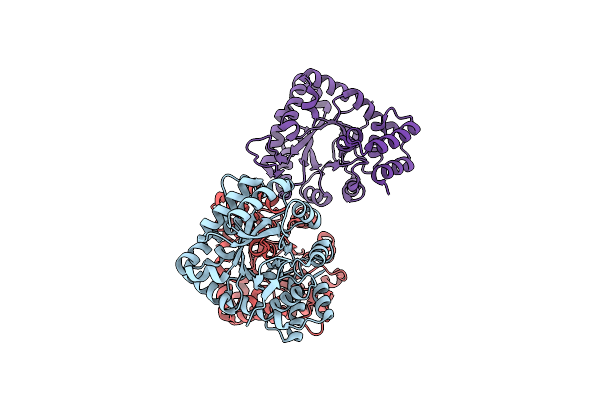 |
Organism: Salmonella enterica subsp. enterica serovar typhimurium
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2014-09-17 Classification: OXIDOREDUCTASE |
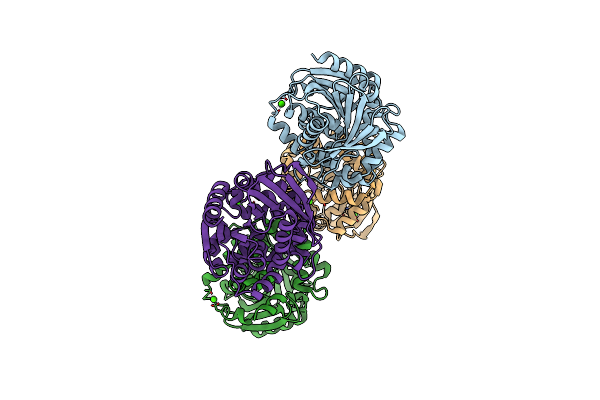 |
2.22 Angstrom Resolution Crystal Structure Of A Putative Acyltransferase From Salmonella Enterica
Organism: Salmonella enterica
Method: X-RAY DIFFRACTION Resolution:2.22 Å Release Date: 2013-12-18 Classification: TRANSFERASE Ligands: CA, CL |
 |
Crystal Structure Of Anabolic Ornithine Carbamoyltransferase From Bacillus Anthracis In Complex With Carbamoyl Phosphate And L-Norvaline
Organism: Bacillus anthracis
Method: X-RAY DIFFRACTION Resolution:1.74 Å Release Date: 2013-11-13 Classification: TRANSFERASE Ligands: CP, NVA, CL |
 |
Crystal Structure Of C103A Mutant Of Dj-1 Superfamily Protein Stm1931 From Salmonella Typhimurium
Organism: Salmonella enterica subsp. enterica serovar typhimurium
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2013-04-24 Classification: UNKNOWN FUNCTION Ligands: ZN |
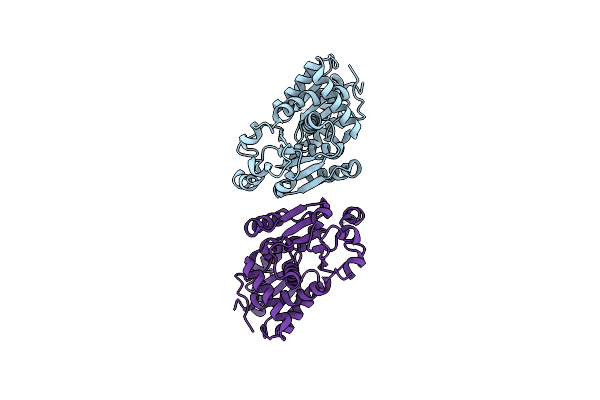 |
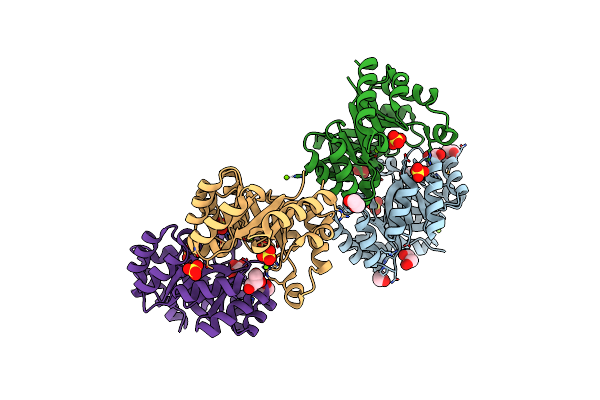
Crystal Structure Of Anabolic Ornithine Carbamoyltransferase From Bacillus Anthracis
Organism: Bacillus anthracis
Method: X-RAY DIFFRACTION Resolution:3.25 Å Release Date: 2012-04-25 Classification: TRANSFERASE |
 |
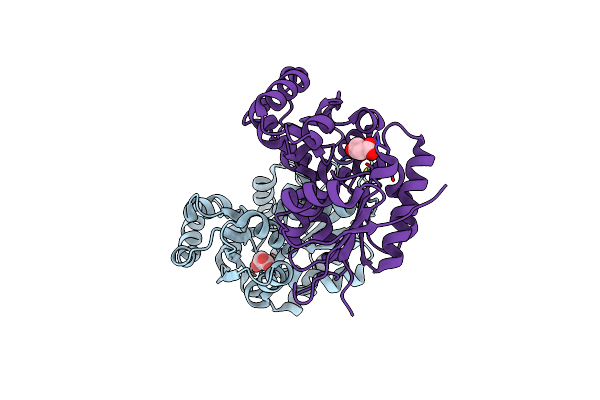
X-Ray Crystal Structure Of The Mg-Bound 3-Keto-L-Gulonate-6-Phosphate Decarboxylase From Vibrio Cholerae O1 Biovar El Tor Str. N16961
Organism: Vibrio cholerae
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2009-10-13 Classification: BIOSYNTHETIC PROTEIN Ligands: EDO, GOL, SO4, MG |
 |
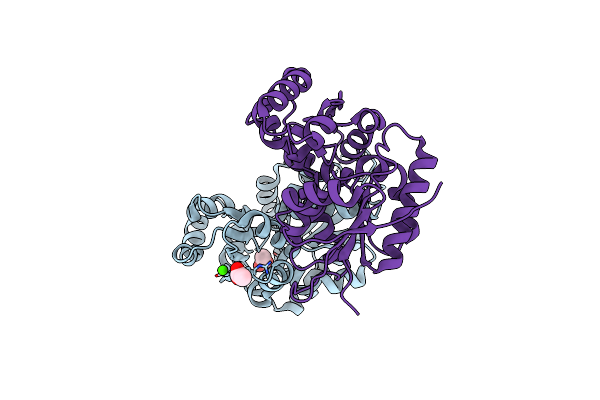
Crystal Structure Of Glutamate Racemase From Listeria Monocytogenes In Complex With Succinic Acid
Organism: Listeria monocytogenes
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2009-09-22 Classification: ISOMERASE Ligands: SIN, CL |
 |
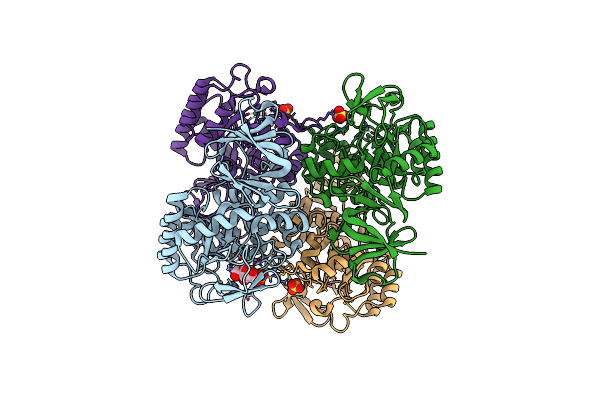
Crystal Structure Of Glutamate Racemase From Listeria Monocytogenes In Complex With Acetate Ion
Organism: Listeria monocytogenes
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2009-09-22 Classification: ISOMERASE Ligands: ACT, CA, CL |
 |
N-Acetylglucosamine-6-Phosphate Deacetylase From Vibrio Cholerae Complexed With Fructose 6-Phosphate
Organism: Vibrio cholerae
Method: X-RAY DIFFRACTION Resolution:2.53 Å Release Date: 2009-09-08 Classification: HYDROLASE Ligands: F6P, NI, SO4 |
 |
Crystal Structure Of 3-Keto-L-Gulonate-6-Phosphate Decarboxylase From Vibrio Cholerae O1 Biovar El Tor Str. N16961
Organism: Vibrio cholerae
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2009-08-18 Classification: BIOSYNTHETIC PROTEIN Ligands: SO4, GOL |
 |
Organism: Yersinia pseudotuberculosis ypiii
Method: X-RAY DIFFRACTION Resolution:2.21 Å Release Date: 2009-08-11 Classification: OXIDOREDUCTASE Ligands: MG, EDO |

