Search Count: 152
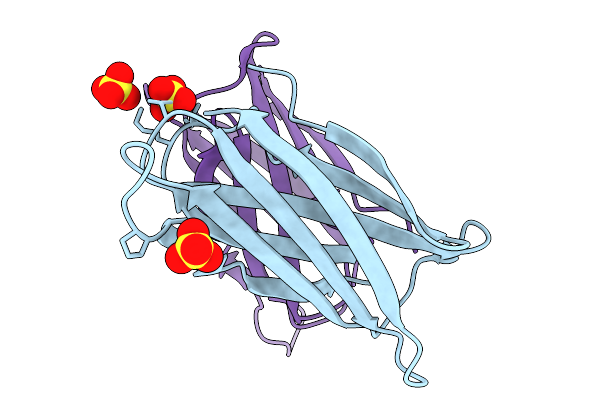 |
Crystal Structure Of The Helicobacter Pylori Copper Resistance Determinant Crda In The Apo Form
Organism: Helicobacter pylori
Method: X-RAY DIFFRACTION Release Date: 2025-11-19 Classification: METAL BINDING PROTEIN Ligands: SO4 |
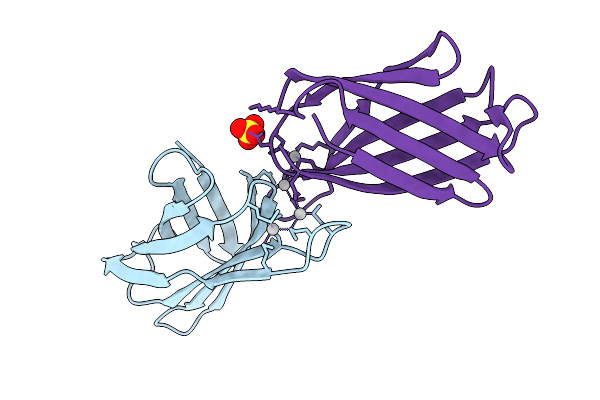 |
Crystal Structure Of The Helicobacter Pylori Copper Resistance Determinant Crda In Complex With Silver Ions In Space Group P21
Organism: Helicobacter pylori
Method: X-RAY DIFFRACTION Release Date: 2025-11-19 Classification: METAL BINDING PROTEIN Ligands: AG, SO4 |
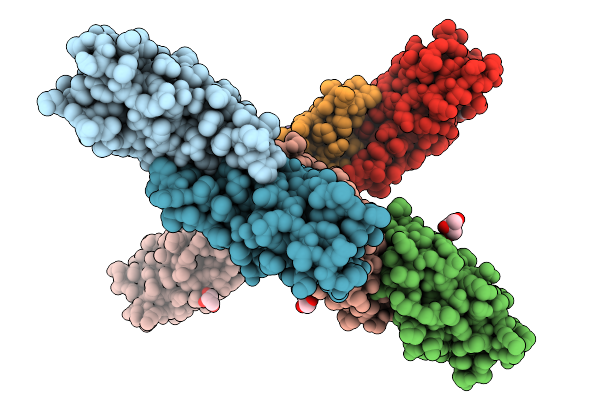 |
Crystal Structure Of The Helicobacter Pylori Copper Resistance Determinant Crda In Complex With Silver Ions In Space Group P1
Organism: Helicobacter pylori
Method: X-RAY DIFFRACTION Release Date: 2025-11-19 Classification: METAL BINDING PROTEIN Ligands: AG, GOL |
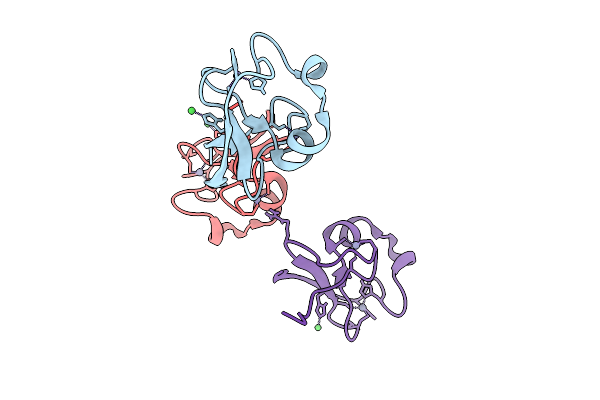 |
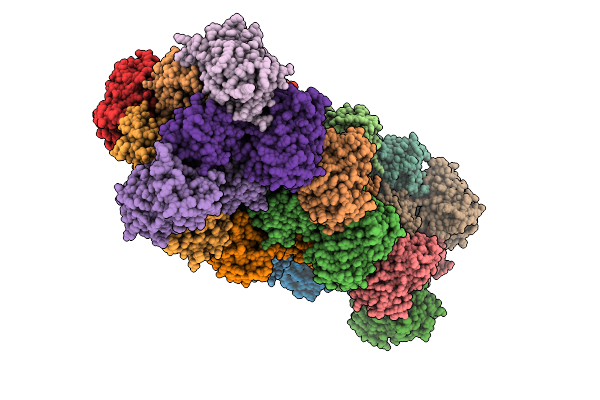
R-Degron Fused Zz-Domain Of The Arabidopsis Thaliana E3 Ubiquitin-Protein Ligase Big
Organism: Arabidopsis thaliana
Method: X-RAY DIFFRACTION Release Date: 2025-09-17 Classification: LIGASE Ligands: ZN, NI |
 |
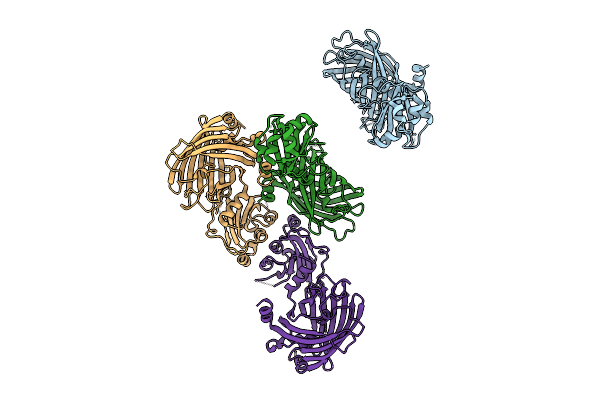
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Release Date: 2025-09-17 Classification: FLUORESCENT PROTEIN Ligands: NO3, SO4 |
 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Release Date: 2025-09-17 Classification: FLUORESCENT PROTEIN |
 |
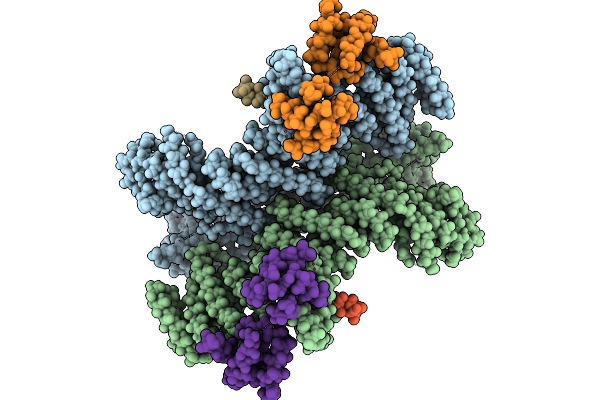
Cryo-Em Structure Of Formyl Peptide Receptor 2/C1R Receptor In Complex With Gi
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-09-10 Classification: STRUCTURAL PROTEIN Ligands: A1L1D |
 |
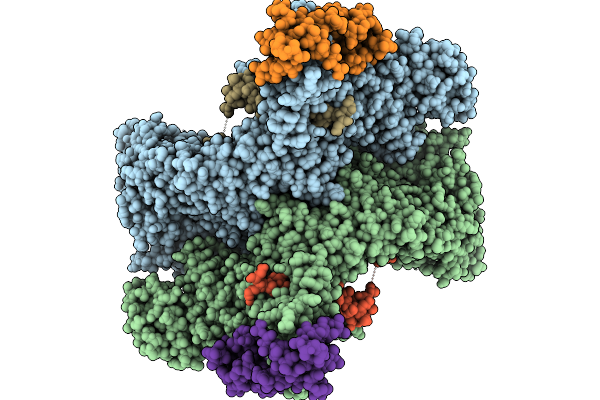
Cryo-Em Structure Of The Core Of The Arabidopsis Thaliana Ubr4/Di19/Calm1 Complex
Organism: Arabidopsis thaliana
Method: ELECTRON MICROSCOPY Release Date: 2025-09-03 Classification: LIGASE |
 |
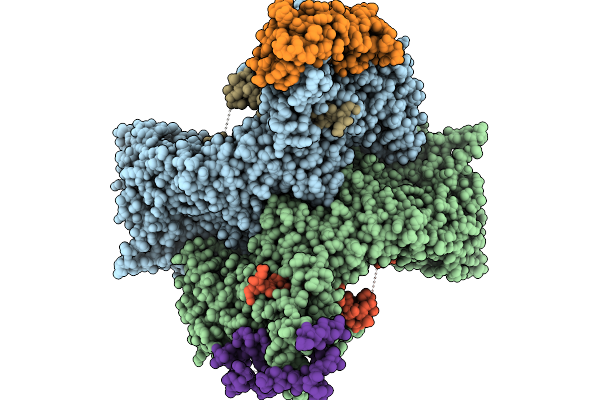
Cryo-Em Structure Of The Human Ubr4/Kcmf1/Calm1 Complex (C-Term Dimer Interface Focused Refinement)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-09-03 Classification: LIGASE Ligands: ZN |
 |
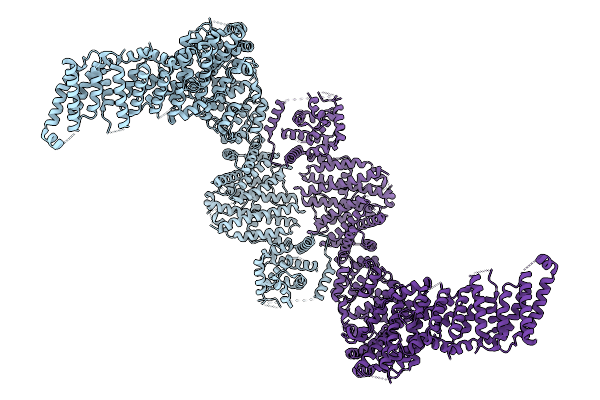
Cryo-Em Structure Of The Human Ubr4/Kcmf1/Calm1 Complex (Calm1 Focused Refinement)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-09-03 Classification: LIGASE Ligands: ZN |
 |
Cryo-Em Structure Of The Human Ubr4/Kcmf1/Calm1 Complex (N-Term Focused Refinement)
|
 |
Cryo-Em Structure Of The Human Ubr4/Kcmf1/Calm1 Complex (Bp Focused Refinement)
|
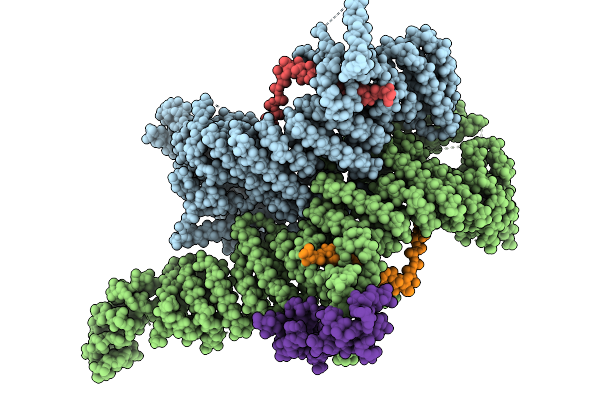 |
Cryo-Em Structure Of The Human Ubr4/Kcmf1/Calm1 Complex (C-Term Focused Refinement)
|
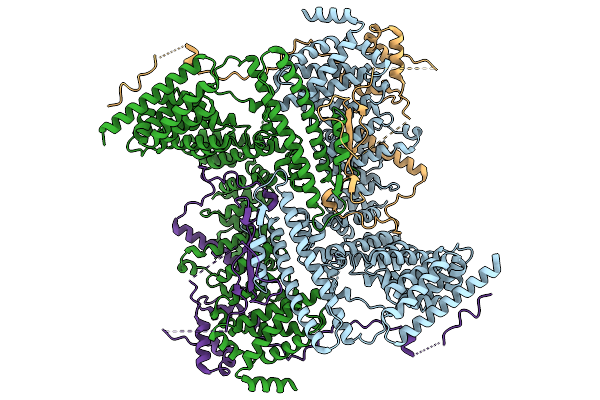 |
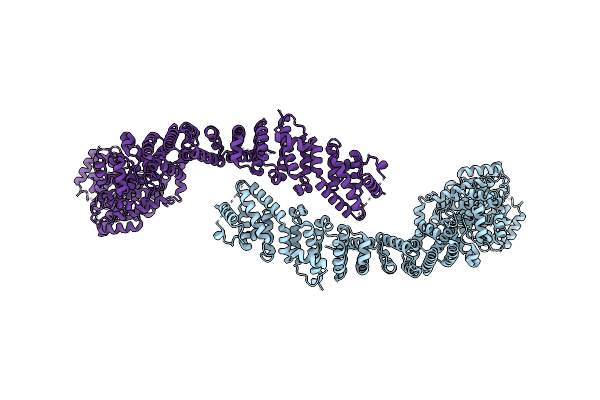
Cryo-Em Structure Of The C. Elegans Ubr4/Kcmf1 Complex (C-Term Dimer Interface Focused Refinement)
Organism: Caenorhabditis elegans
Method: ELECTRON MICROSCOPY Release Date: 2025-09-03 Classification: LIGASE Ligands: ZN |
 |
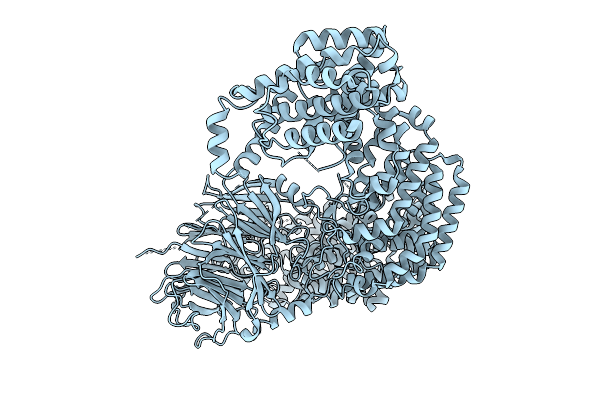
Cryo-Em Structure Of The C. Elegans Ubr4/Kcmf1 Complex (N-Term Focused Refinement)
Organism: Caenorhabditis elegans
Method: ELECTRON MICROSCOPY Release Date: 2025-09-03 Classification: LIGASE |
 |
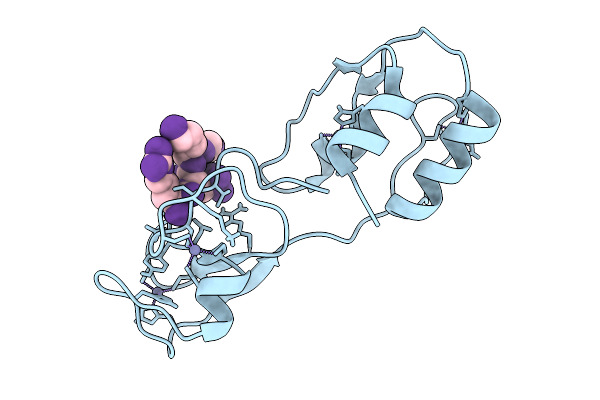
Cryo-Em Structure Of The C. Elegans Ubr4/Kcmf1 Complex (Side Focused Refinement)
Organism: Caenorhabditis elegans
Method: ELECTRON MICROSCOPY Release Date: 2025-09-03 Classification: LIGASE |
 |
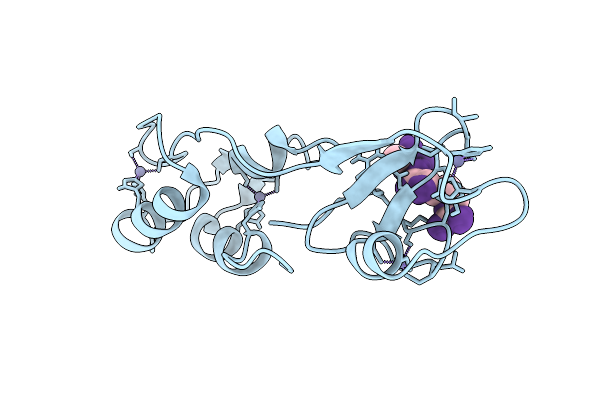
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Release Date: 2025-08-27 Classification: PEPTIDE BINDING PROTEIN Ligands: ZN |
 |
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Release Date: 2025-08-27 Classification: LIGASE Ligands: ZN |
 |
Organism: Arabidopsis thaliana
Method: X-RAY DIFFRACTION Resolution:1.74 Å Release Date: 2025-05-14 Classification: LIGASE Ligands: ZN |
 |
Y-Degron Fused Zz-Domain Of The Arabidopsis Thaliana E3 Ubiquitin-Protein Ligase Prt1
Organism: Arabidopsis thaliana
Method: X-RAY DIFFRACTION Resolution:1.67 Å Release Date: 2025-05-14 Classification: LIGASE Ligands: ZN, MG |

