Search Count: 14
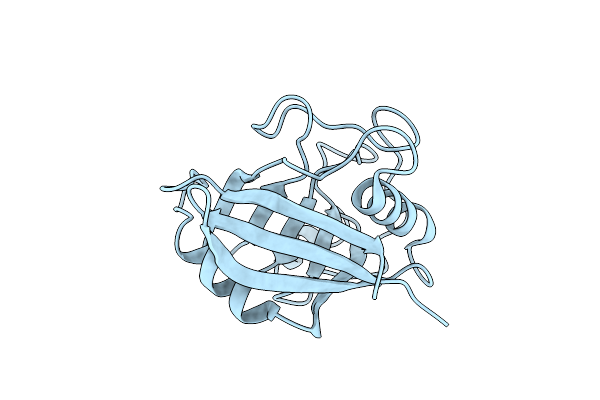 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.62 Å Release Date: 2020-01-29 Classification: ISOMERASE |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2020-01-29 Classification: ISOMERASE |
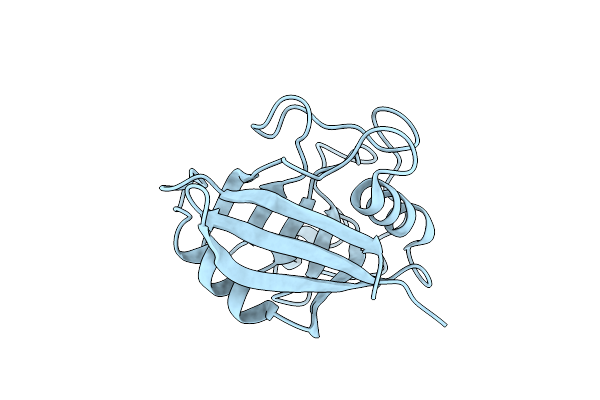 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.56 Å Release Date: 2020-01-29 Classification: ISOMERASE |
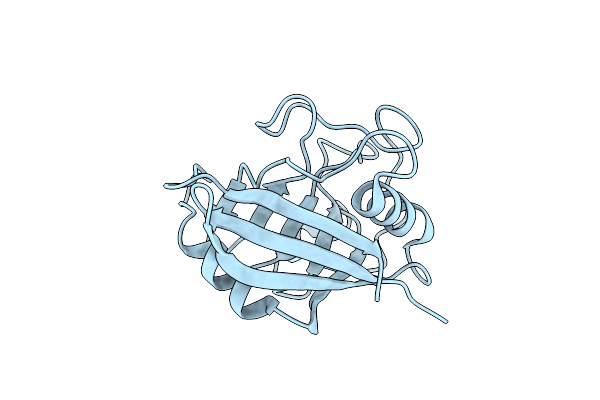 |
Organism: Homo sapiens
Method: ELECTRON CRYSTALLOGRAPHY Resolution:2.50 Å Release Date: 2020-01-29 Classification: ISOMERASE |
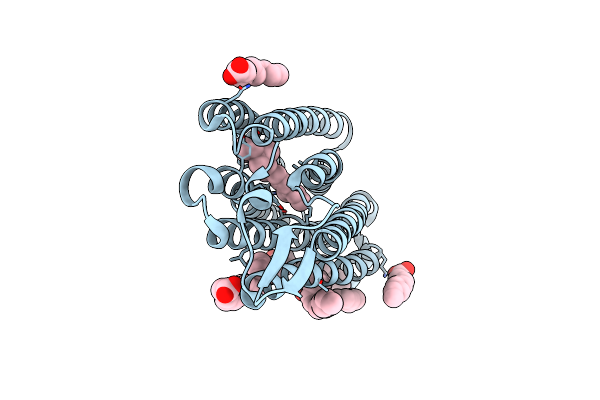 |
Organism: Nonlabens marinus s1-08
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2018-12-05 Classification: SIGNALING PROTEIN Ligands: CL, RET, OLA |
 |
Organism: Nonlabens marinus
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2018-12-05 Classification: TRANSPORT PROTEIN Ligands: CL, RET, OLA |
 |
Organism: Deinococcus radiodurans
Method: X-RAY DIFFRACTION Resolution:1.35 Å Release Date: 2016-10-26 Classification: TRANSFERASE Ligands: ACT, MPD, LBV |
 |
Wild-Type Pas-Gaf Fragment From Deinococcus Radiodurans Bphp Collected At Lcls
Organism: Deinococcus radiodurans
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2016-10-26 Classification: TRANSFERASE Ligands: LBV, ACT |
 |
Wild-Type Pas-Gaf Fragment From Deinococcus Radiodurans Bphp Collected At Sacla
Organism: Deinococcus radiodurans
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2016-10-26 Classification: TRANSFERASE Ligands: LBV, ACT |
 |
Femtosecond Structural Dynamics Drives The Trans/Cis Isomerization In Photoactive Yellow Protein: 200 Ns Time Delay Photo-Activated (Light) Structure
Organism: Halorhodospira halophila
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2016-05-25 Classification: SIGNALING PROTEIN |
 |
Femtosecond Structural Dynamics Drives The Trans/Cis Isomerization In Photoactive Yellow Protein: 100 Fs To 400 Fs Structure
Organism: Halorhodospira halophila
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2016-05-25 Classification: SIGNALING PROTEIN |
 |
Femtosecond Structural Dynamics Drives The Trans/Cis Isomerization In Photoactive Yellow Protein: 800 Fs To 1200 Fs Structure
Organism: Halorhodospira halophila
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2016-05-25 Classification: SIGNALING PROTEIN |
 |
Femtosecond Structural Dynamics Drives The Trans/Cis Isomerization In Photoactive Yellow Protein: 3 Ps Structure
Organism: Halorhodospira halophila
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2016-05-25 Classification: SIGNALING PROTEIN |
 |
Femtosecond Structural Dynamics Drives The Trans/Cis Isomerization In Photoactive Yellow Protein: Dark Structure Of Photoactive Yellow Protein
Organism: Halorhodospira halophila
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2016-05-18 Classification: SIGNALING PROTEIN |

