Search Count: 9
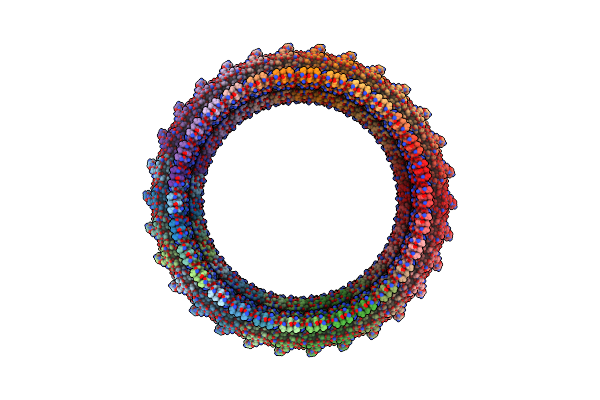 |
Organism: Elizabethkingia anophelis ag1
Method: ELECTRON MICROSCOPY Release Date: 2025-04-09 Classification: TOXIN Ligands: CA |
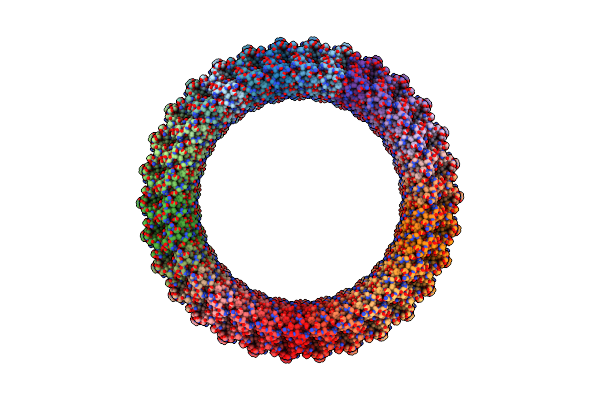 |
Organism: Elizabethkingia anophelis ag1
Method: ELECTRON MICROSCOPY Release Date: 2025-04-09 Classification: TOXIN Ligands: CA |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.24 Å Release Date: 2024-06-05 Classification: TRANSFERASE Ligands: ZN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2024-06-05 Classification: TRANSFERASE Ligands: ZN, EDO |
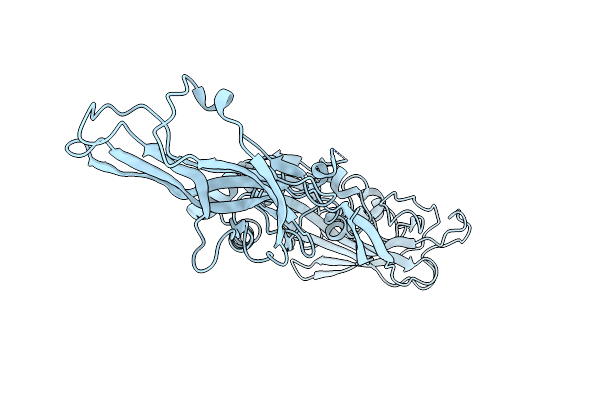 |
Organism: Elizabethkingia anophelis ag1
Method: X-RAY DIFFRACTION Resolution:2.49 Å Release Date: 2024-02-07 Classification: TOXIN |
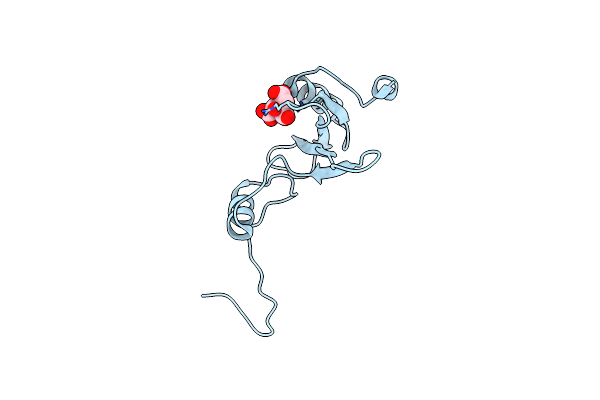 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.35 Å Release Date: 2012-02-08 Classification: TRANSFERASE Ligands: CIT |
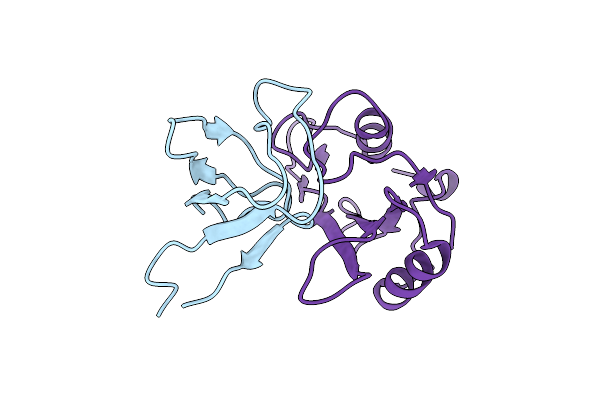 |
Solution Structure Of The Binary Complex Between The Sh3 And Sh2 Domain Of Interleukin-2 Tyrosine Kinase
|
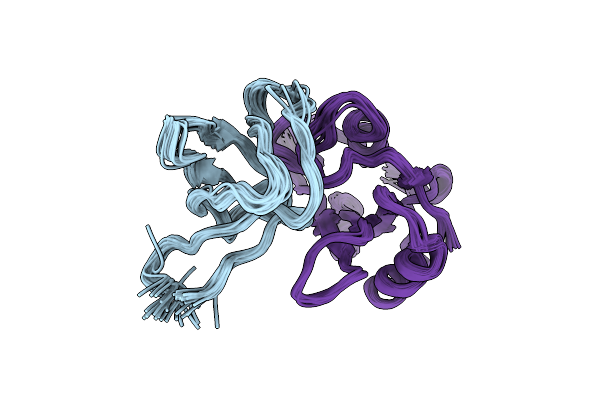 |
Ensemble Structures Of The Binary Complex Between The Sh3 And Sh2 Domain Of Interleukin-2 Tyrosine Kinase.
|
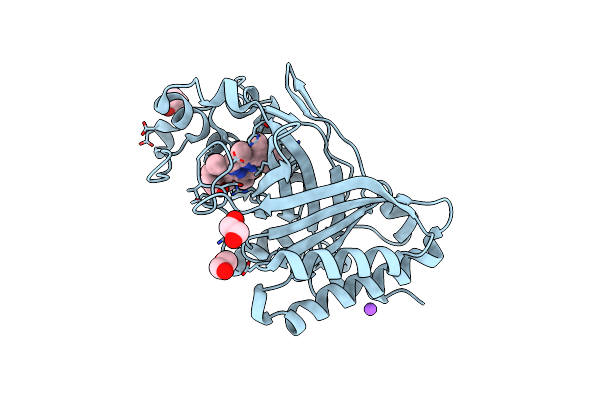 |
Crystal Structure Of Putative Melanin Biosynthesis Protein Tyra With Bound Heme (Np_716371.1) From Shewanella Oneidensis At 2.30 A Resolution
Organism: Shewanella oneidensis
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2006-11-14 Classification: Structural Genomics/Unknown function |

